1CQP
 
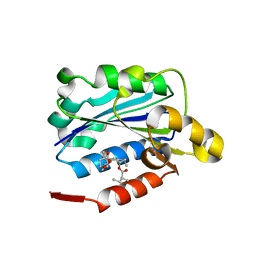 | | CRYSTAL STRUCTURE ANALYSIS OF THE COMPLEX LFA-1 (CD11A) I-DOMAIN / LOVASTATIN AT 2.6 A RESOLUTION | | Descriptor: | ANTIGEN CD11A (P180), LOVASTATIN, MAGNESIUM ION | | Authors: | Kallen, J, Welzenbach, K, Ramage, P, Geyl, D, Kriwacki, R, Legge, G, Cottens, S, Weitz-Schmidt, G, Hommel, U. | | Deposit date: | 1999-08-10 | | Release date: | 2000-08-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for LFA-1 inhibition upon lovastatin binding to the CD11a I-domain.
J.Mol.Biol., 292, 1999
|
|
1XDG
 
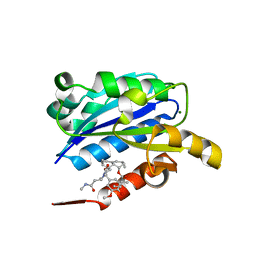 | | X-ray structure of LFA-1 I-domain in complex with LFA878 at 2.1A resolution | | Descriptor: | (1S,3R,8AS)-8-(2-{(4S,6S)-3-(4-HYDROXY-3-METHOXYBENZYL)-4-[2-(METHYLAMINO)-2-OXOETHYL]-2-OXO-1,3-OXAZINAN-6-YL}ETHYL)-3 ,7-DIMETHYL-1,2,3,7,8,8A-HEXAHYDRONAPHTHALEN-1-YL (2R)-2-METHYLBUTANOATE, Integrin alpha-L, MAGNESIUM ION | | Authors: | Weitz-Schmidt, G, Welzenbach, K, Dawson, J, Kallen, J. | | Deposit date: | 2004-09-06 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Improved lymphocyte function-associated antigen-1 (LFA-1) inhibition by statin derivatives: molecular basis determined by x-ray analysis and monitoring of LFA-1 conformational changes in vitro and ex vivo
J.Biol.Chem., 279, 2004
|
|
1XDD
 
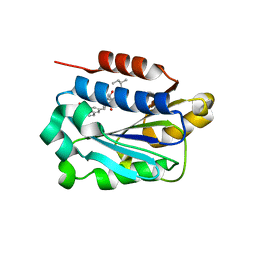 | | X-ray structure of LFA-1 I-domain in complex with LFA703 at 2.2A resolution | | Descriptor: | 8-[2-((2S)-4-HYDROXY-1-{[5-(HYDROXYMETHYL)-6-METHOXY-2-NAPHTHYL]METHYL}-6-OXOPIPERIDIN-2-YL)ETHYL]-3,7-DIMETHYL-1,2,3,7,8,8A-HEXAHYDRONAPHTHALEN-1-YL 2-METHYLBUTANOATE, Integrin alpha-L, MAGNESIUM ION | | Authors: | Weitz-Schmidt, G, Welzenbach, K, Dawson, J, Kallen, J. | | Deposit date: | 2004-09-06 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Improved lymphocyte function-associated antigen-1 (LFA-1) inhibition by statin derivatives: molecular basis determined by x-ray analysis and monitoring of LFA-1 conformational changes in vitro and ex vivo
J.Biol.Chem., 279, 2004
|
|
1CPC
 
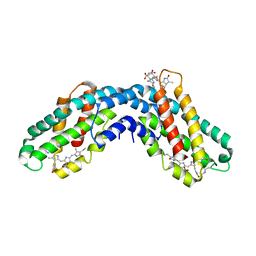 | | ISOLATION, CRYSTALLIZATION, CRYSTAL STRUCTURE ANALYSIS AND REFINEMENT OF CONSTITUTIVE C-PHYCOCYANIN FROM THE CHROMATICALLY ADAPTING CYANOBACTERIUM FREMYELLA DIPLOSIPHON AT 1.66 ANGSTROMS RESOLUTION | | Descriptor: | C-PHYCOCYANIN (ALPHA SUBUNIT), C-PHYCOCYANIN (BETA SUBUNIT), PHYCOCYANOBILIN | | Authors: | Duerring, M, Schmidt, G.B, Huber, R. | | Deposit date: | 1990-10-11 | | Release date: | 1993-01-15 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Isolation, crystallization, crystal structure analysis and refinement of constitutive C-phycocyanin from the chromatically adapting cyanobacterium Fremyella diplosiphon at 1.66 A resolution.
J.Mol.Biol., 217, 1991
|
|
1QPY
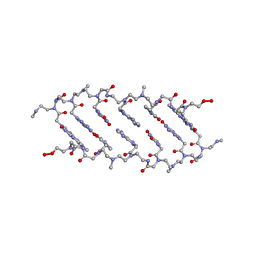 
 | | CRYSTAL STRUCTURE OF BACKBONE MODIFIED PNA HEXAMER | | Descriptor: | PEPTIDE NUCLEIC ACID 5'-(*CP1*GPN*TP1*APN*CP1*GPN*LYS)-3' | | Authors: | Haima, G, Rasmussen, H, Schmidt, G, Jensen, D.K, Kastrup, J.S, Stafshede, P.W, Norden, B, Buchardt, O, Nielsen, P.E. | | Deposit date: | 1999-05-14 | | Release date: | 2001-02-21 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Peptide Nucleic Acids (PNA) derived from N-(N-methylaminoethyl)glycine. Synthesis, hybridization and structural properties
New J.Chem., 23, 1999
|
|
1R4T
 
 | | Solution structure of exoenzyme S | | Descriptor: | exoenzyme S | | Authors: | Langdon, G.M, Leitner, D, Labudde, D, Kuhne, R, Schmieder, P, Aktories, K, Oschkinat, H.O, Schmidt, G. | | Deposit date: | 2003-10-08 | | Release date: | 2005-04-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the N-terminal GTPase activating domain of Pseudomonas aeruginosa exoenzyme S
To be Published
|
|
6GF1
 
 | | The structure of the ubiquitin-like modifier FAT10 reveals a novel targeting mechanism for degradation by the 26S proteasome | | Descriptor: | SULFATE ION, Ubiquitin D | | Authors: | Aichem, A, Anders, S, Catone, N, Roessler, P, Stotz, S, Berg, A, Schwab, R, Scheuermann, S, Bialas, J, Schmidtke, G, Peter, C, Groettrup, M, Wiesner, S. | | Deposit date: | 2018-04-28 | | Release date: | 2018-08-29 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.925 Å) | | Cite: | The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation.
Nat Commun, 9, 2018
|
|
6GF2
 
 | | The structure of the ubiquitin-like modifier FAT10 reveals a novel targeting mechanism for degradation by the 26S proteasome | | Descriptor: | Ubiquitin D | | Authors: | Aichem, A, Anders, S, Catone, N, Roessler, P, Stotz, S, Berg, A, Schwab, R, Scheuermann, S, Bialas, J, Schmidtke, G, Peter, C, Groettrup, M, Wiesner, S. | | Deposit date: | 2018-04-29 | | Release date: | 2018-08-08 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation.
Nat Commun, 9, 2018
|
|
