6ZYI
 
 | |
6ZYJ
 
 | | Crystal structure of Hsc70 ATPase domain in complex with ADP and calcium | | Descriptor: | ACETATE ION, ADENOSINE-5'-DIPHOSPHATE, CALCIUM ION, ... | | Authors: | Yan, Y, Preissler, S, Ron, D. | | Deposit date: | 2020-08-01 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Calcium depletion challenges endoplasmic reticulum proteostasis by destabilising BiP-substrate complexes.
Elife, 9, 2020
|
|
7A4V
 
 | |
6ZYH
 
 | |
7A4U
 
 | |
1RZ9
 
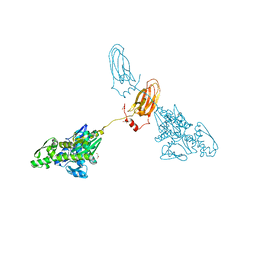
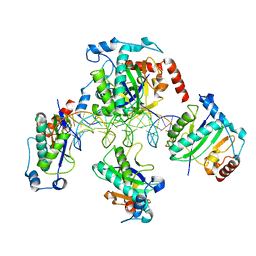 | | Crystal Structure of AAV Rep complexed with the Rep-binding sequence | | Descriptor: | 26-MER, Rep protein | | Authors: | Hickman, A.B, Ronning, D.R, Perez, Z.N, Kotin, R.M, Dyda, F. | | Deposit date: | 2003-12-24 | | Release date: | 2004-02-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The nuclease domain of adeno-associated virus rep coordinates replication initiation using two distinct DNA recognition interfaces.
Mol.Cell, 13, 2004
|
|
5KWI
 
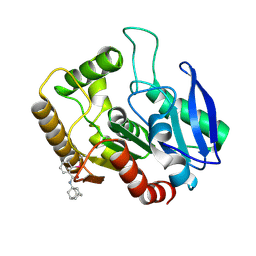 | | M.tb Ag85C modified at C209 by adamantyl-ebselen | | Descriptor: | Diacylglycerol acyltransferase/mycolyltransferase Ag85C, ~{N}-(1-adamantyl)-2-selanyl-benzamide | | Authors: | Goins, C.M, Ronning, D.R. | | Deposit date: | 2016-07-18 | | Release date: | 2017-03-29 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Exploring Covalent Allosteric Inhibition of Antigen 85C from Mycobacterium tuberculosis by Ebselen Derivatives.
ACS Infect Dis, 3, 2017
|
|
8S9D
 
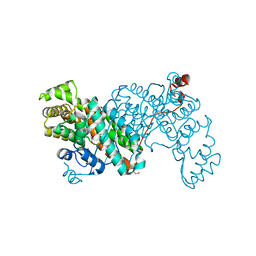 | |
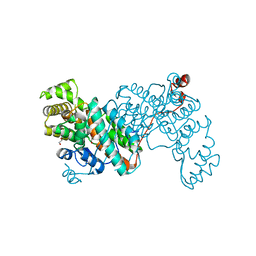
8S97
 
 | |
8FX9
 
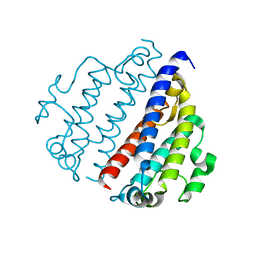 | |
2BZF
 
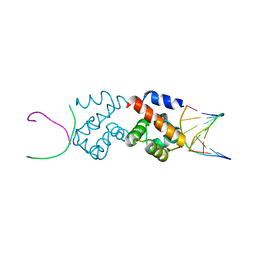 | | Structural basis for DNA bridging by barrier-to-autointegration factor (BAF) | | Descriptor: | 5'-D(*CP*CP*TP*CP*CP*AP*CP)-3', 5'-D(*GP*TP*GP*GP*AP*GP*GP)-3', BARRIER-TO-AUTOINTEGRATION FACTOR | | Authors: | Bradley, C.M, Ronning, D.R, Ghirlando, R, Craigie, R, Dyda, F. | | Deposit date: | 2005-08-16 | | Release date: | 2005-09-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Structural Basis for DNA Bridging by Barrier-to-Autointegration Factor.
Nat.Struct.Mol.Biol., 12, 2005
|
|
7MEL
 
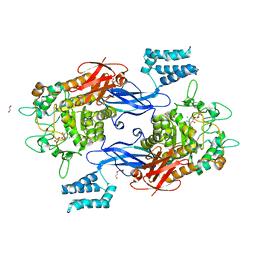 | | Sco GlgEI-V279S in complex with 4-alpha-glucoside of validamine | | Descriptor: | (1R,2R,3S,4S,6R)-4-amino-2,3-dihydroxy-6-(hydroxymethyl)cyclohexyl alpha-D-glucopyranoside, 1,2-ETHANEDIOL, Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase 1, ... | | Authors: | Jayasinghe, T.D, Ronning, D.R. | | Deposit date: | 2021-04-06 | | Release date: | 2021-07-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Stereoselective synthesis of a 4-alpha-glucoside of valienamine and its X-ray structure in complex with Streptomyces coelicolor GlgE1-V279S.
Sci Rep, 11, 2021
|
|
6EZO
 
 | | Eukaryotic initiation factor EIF2B in complex with ISRIB | | Descriptor: | 2-(4-chloranylphenoxy)-~{N}-[4-[2-(4-chloranylphenoxy)ethanoylamino]cyclohexyl]ethanamide, Human eukaryotic initiation factor EIF2B epsilon subunits, Translation initiation factor eIF-2B subunit alpha, ... | | Authors: | Faille, A, Weis, F, Zyryanova, A, Warren, A.J, Ron, D. | | Deposit date: | 2017-11-16 | | Release date: | 2018-03-28 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Binding of ISRIB reveals a regulatory site in the nucleotide exchange factor eIF2B.
Science, 359, 2018
|
|
7MGY
 
 | | Sco GlgEI-V279S in complex with 4-alpha-glucoside of valienamine | | Descriptor: | (1R,4S,5S,6R)-4-amino-5,6-dihydroxy-2-(hydroxymethyl)cyclohex-2-en-1-yl alpha-D-glucopyranoside, Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase 1, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Jayasinghe, T.D, Ronning, D.R. | | Deposit date: | 2021-04-13 | | Release date: | 2021-07-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Stereoselective synthesis of a 4-⍺-glucoside of valienamine and its X-ray structure in complex with Streptomyces coelicolor GlgE1-V279S.
Sci Rep, 11, 2021
|
|
5O4P
 
 | | Crystal structure of AMPylated GRP78 | | Descriptor: | 78 kDa glucose-regulated protein, ADENOSINE MONOPHOSPHATE, SULFATE ION | | Authors: | Yan, Y, Chen, R, Ron, D, Read, R. | | Deposit date: | 2017-05-30 | | Release date: | 2017-11-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | AMPylation targets the rate-limiting step of BiP's ATPase cycle for its functional inactivation.
Elife, 6, 2017
|
|
6EOF
 
 | | Crystal structure of AMPylated GRP78 in ADP state | | Descriptor: | 78 kDa glucose-regulated protein, ADENOSINE MONOPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Yan, Y, Preissler, S, Read, R.J, Ron, D. | | Deposit date: | 2017-10-09 | | Release date: | 2017-11-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | AMPylation targets the rate-limiting step of BiP's ATPase cycle for its functional inactivation.
Elife, 6, 2017
|
|
6I7L
 
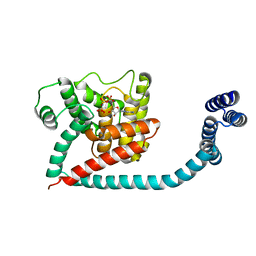 | | Crystal structure of monomeric FICD mutant L258D complexed with MgAMP-PNP | | Descriptor: | Adenosine monophosphate-protein transferase FICD, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
6I7G
 
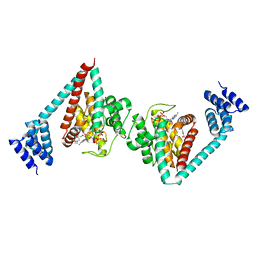 | | Crystal structure of dimeric wild type FICD complexed with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Adenosine monophosphate-protein transferase FICD, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
6I7I
 
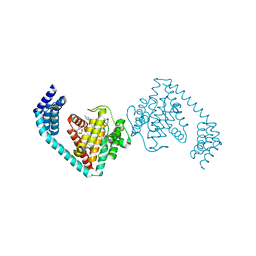 | | Crystal structure of dimeric FICD mutant K256A complexed with MgATP | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, ADENOSINE-5'-TRIPHOSPHATE, Adenosine monophosphate-protein transferase FICD, ... | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
6HA7
 
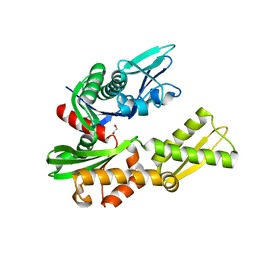 | | Crystal structure of the BiP NBD and MANF complex | | Descriptor: | 1,2-ETHANEDIOL, Endoplasmic reticulum chaperone BiP, Mesencephalic astrocyte-derived neurotrophic factor | | Authors: | Yan, Y, Ron, D. | | Deposit date: | 2018-08-07 | | Release date: | 2019-02-06 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | MANF antagonizes nucleotide exchange by the endoplasmic reticulum chaperone BiP.
Nat Commun, 10, 2019
|
|
5VSJ
 
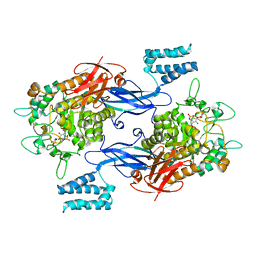 | | Sco GlgEI-V279S in complex with a pyrolidene-based ethyl-phosphonate compound | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase 1, {2-[(2R,3R,4R,5R)-3-(alpha-D-glucopyranosyloxy)-4-hydroxy-2,5-bis(hydroxymethyl)pyrrolidin-1-yl]ethyl}phosphonic acid | | Authors: | Petit, C, Ronning, D.R. | | Deposit date: | 2017-05-11 | | Release date: | 2017-05-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.456 Å) | | Cite: | Zwitterionic pyrrolidene-phosphonates: inhibitors of the glycoside hydrolase-like phosphorylase Streptomyces coelicolor GlgEI-V279S.
Org. Biomol. Chem., 15, 2017
|
|
6I7H
 
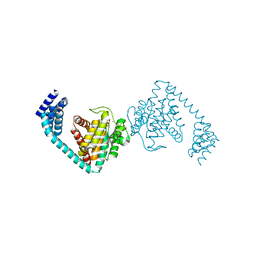 | | Crystal structure of dimeric FICD mutant K256S | | Descriptor: | Adenosine monophosphate-protein transferase FICD, MAGNESIUM ION | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
4MQL
 
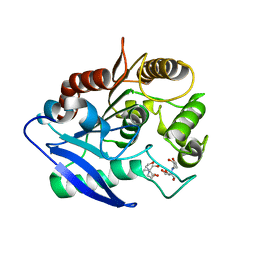 | | Crystal structure of Antigen 85C-C209S mutant | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Diacylglycerol acyltransferase/mycolyltransferase Ag85C | | Authors: | Favrot, L, Grzegorzewicz, A.E, Lajiness, D.H, Marvin, R.K, Boucau, J, Isailovic, D, Jackson, M, Ronning, D.R. | | Deposit date: | 2013-09-16 | | Release date: | 2013-11-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Mechanism of inhibition of Mycobacterium tuberculosis antigen 85 by ebselen.
Nat Commun, 4, 2013
|
|
4MQM
 
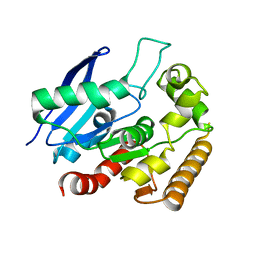 | | Crystal structure of Antigen 85C in presence of Ebselen | | Descriptor: | Diacylglycerol acyltransferase/mycolyltransferase Ag85C | | Authors: | Favrot, L, Grzegorzewicz, A.E, Lajiness, D.H, Marvin, R.K, Boucau, J, Isailovic, D, Jackson, M, Ronning, D.R. | | Deposit date: | 2013-09-16 | | Release date: | 2013-11-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.346 Å) | | Cite: | Mechanism of inhibition of Mycobacterium tuberculosis antigen 85 by ebselen.
Nat Commun, 4, 2013
|
|
6EOE
 
 | | Crystal structure of AMPylated GRP78 with nucleotide | | Descriptor: | 78 kDa glucose-regulated protein, ADENOSINE-5'-DIPHOSPHATE, CITRATE ANION, ... | | Authors: | Yan, Y, Preissler, S, Ron, D, Read, R.J. | | Deposit date: | 2017-10-09 | | Release date: | 2017-11-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | AMPylation targets the rate-limiting step of BiP's ATPase cycle for its functional inactivation.
Elife, 6, 2017
|
|
