3EY8
 
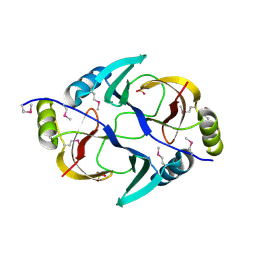 | | Structure from the mobile metagenome of V. Pseudocholerae. VPC_CASS1 | | Descriptor: | Biphenyl-2,3-diol 1,2-dioxygenase III-related protein, SULFATE ION | | Authors: | Harrop, S.J, Deshpande, C.N, Sureshan, V, Boucher, Y, Xu, X, Cui, H, Chang, C, Edwards, A, Joachimiak, A, Savchenko, A, Curmi, P.M.G, Mabbutt, B.C, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2008-10-20 | | Release date: | 2009-01-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure from the mobile metagenome of V. Pseudocholerae. VPC_CASS1
To be Published
|
|
3EYH
 
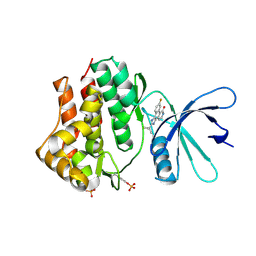 | | Crystal structures of JAK1 and JAK2 inhibitor complexes | | Descriptor: | 2-TERT-BUTYL-9-FLUORO-3,6-DIHYDRO-7H-BENZ[H]-IMIDAZ[4,5-F]ISOQUINOLINE-7-ONE, Tyrosine-protein kinase | | Authors: | Williams, N.K, Bamert, R.S, Patell, O, Wang, C, Walden, P.M, Fantino, E. | | Deposit date: | 2008-10-20 | | Release date: | 2009-02-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Dissecting specificity in the Janus kinases: the structures of JAK-specific inhibitors complexed to the JAK1 and JAK2 protein tyrosine kinase domains.
J.Mol.Biol., 387, 2009
|
|
8FFC
 
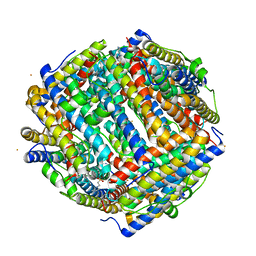 | | Crystal structure of iron bound Dps protein (PA0962) from Pseudomonas aeruginosa (cubic form) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, FE (II) ION, Probable dna-binding stress protein | | Authors: | Lovell, S, Kashipathy, M.M, Battaile, K.P, Rivera, M. | | Deposit date: | 2022-12-08 | | Release date: | 2023-03-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Pseudomonas aeruginosa Dps (PA0962) Functions in H 2 O 2 Mediated Oxidative Stress Defense and Exhibits In Vitro DNA Cleaving Activity.
Int J Mol Sci, 24, 2023
|
|
1U92
 
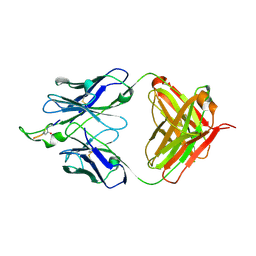 | | Crystal structure of the HIV-1 Cross Neutralizing Monoclonal Antibody 2F5 in complex with gp41 Peptide Analog E-[Dap]-DKWQS (cyclic) | | Descriptor: | ANTIBODY 2F5 (HEAVY CHAIN), ANTIBODY 2F5 (LIGHT CHAIN), GP41 PEPTIDE ANALOG | | Authors: | Bryson, S, Julien, J.-P, Hynes, R.C, Pai, E.F. | | Deposit date: | 2004-08-09 | | Release date: | 2004-10-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Crystallographic definition of the epitope promiscuity of the broadly neutralizing anti-human immunodeficiency virus type 1 antibody 2F5: vaccine design implications
J.Virol., 83, 2009
|
|
7QT6
 
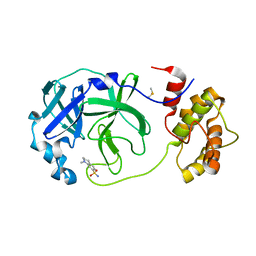 | | Room temperature In-situ SARS-CoV-2 MPRO with bound Z1367324110 | | Descriptor: | 1-methyl-3,4-dihydro-2~{H}-quinoline-7-sulfonamide, 3C-like proteinase nsp5, DIMETHYL SULFOXIDE | | Authors: | Horrell, S, Gildae, R.J, Axford, D, Owen, C.D, Lukacik, P, Strain-Damerell, C, Owen, R.L, Walsh, M.A. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | xia2.multiplex: a multi-crystal data-analysis pipeline.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
7QT5
 
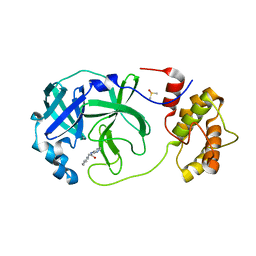 | | Room temperature In-situ SARS-CoV-2 MPRO with bound Z31792168 | | Descriptor: | 2-cyclohexyl-~{N}-pyridin-3-yl-ethanamide, 3C-like proteinase nsp5, DIMETHYL SULFOXIDE | | Authors: | Horrell, S, Gildae, R.J, Axford, D, Owen, C.D, Lukacik, P, Strain-Damerell, C, Owen, R.L, Walsh, M.A. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | xia2.multiplex: a multi-crystal data-analysis pipeline.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
4EIE
 
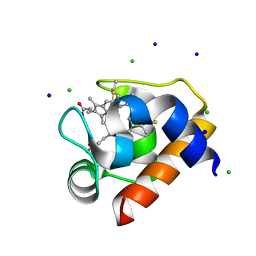 | | Crystal structure of cytochrome c6C from Synechococcus sp. PCC 7002 | | Descriptor: | CHLORIDE ION, Cytochrome c6, HEME C, ... | | Authors: | Krzywda, S, Bialek, W, Zatwarnicki, P, Jaskolski, M, Szczepaniak, A. | | Deposit date: | 2012-04-05 | | Release date: | 2013-04-10 | | Last modified: | 2021-03-10 | | Method: | X-RAY DIFFRACTION (1.03 Å) | | Cite: | Cytochrome c6 and c6C from Synechococcus sp. PCC 7002 - structure and function.
To be Published
|
|
1UA5
 
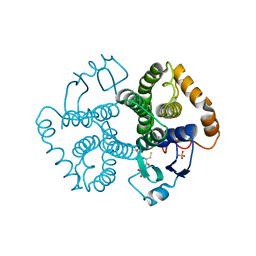 | |
7QT7
 
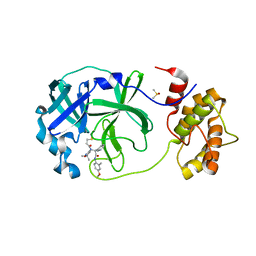 | | Room temperature In-situ SARS-CoV-2 MPRO with bound Z4439011520 | | Descriptor: | 3C-like proteinase nsp5, DIMETHYL SULFOXIDE, N-(5-tert-butyl-1,2-oxazol-3-yl)-N-[(1R)-2-[(4-methoxy-2-methylphenyl)amino]-2-oxo-1-(pyridin-3-yl)ethyl]propanamide | | Authors: | Horrell, S, Gildae, R.J, Axford, D, Owen, C.D, Lukacik, P, Strain-Damerell, C, Owen, R.L, Walsh, M.A. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | xia2.multiplex: a multi-crystal data-analysis pipeline.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
8FT6
 
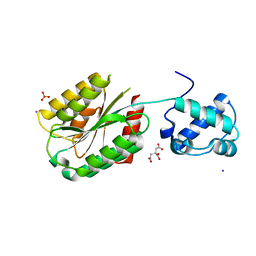 | | The von Willebrand factor A domain of human capillary morphogenesis gene II, flexibly fused to the 1TEL crystallization chaperone, Ala-Ala linker variant, SUMO tag-free preparation. | | Descriptor: | CITRIC ACID, IODIDE ION, SULFATE ION, ... | | Authors: | Gajjar, P.L, Litchfield, C.M, Callahan, M, Redd, N, Doukov, T, Moody, J.D. | | Deposit date: | 2023-01-11 | | Release date: | 2023-03-22 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | Increasing the bulk of the 1TEL-target linker and retaining the 10×His tag in a 1TEL-CMG2-vWa construct improves crystal order and diffraction limits.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
3F2G
 
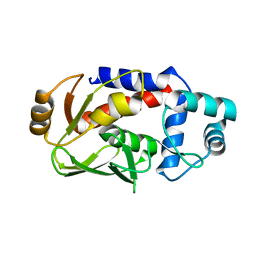 | | Crystal structure of MerB mutant C160S, the Organomercurial Lyase involved in a bacterial mercury resistance system | | Descriptor: | Alkylmercury lyase | | Authors: | Lafrance-Vanasse, J, Lefebvre, M, Di Lello, P, Sygusch, J, Omichinski, J.G. | | Deposit date: | 2008-10-29 | | Release date: | 2008-11-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.781 Å) | | Cite: | Crystal Structures of the Organomercurial Lyase MerB in Its Free and Mercury-bound Forms: INSIGHTS INTO THE MECHANISM OF METHYLMERCURY DEGRADATION
J.Biol.Chem., 284, 2009
|
|
2IGZ
 
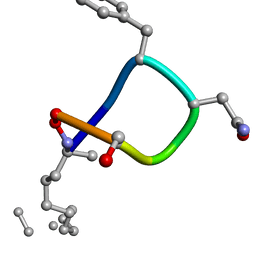 | |
7Q8C
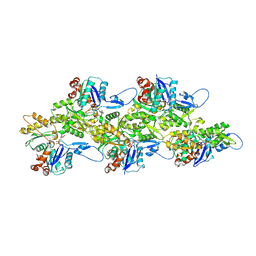 
 | | Leishmania major actin filament in ADP-state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, MAGNESIUM ION | | Authors: | Kotila, T, Muniyandi, S, Lappalainen, P, Huiskonen, J.T. | | Deposit date: | 2021-11-11 | | Release date: | 2022-05-18 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (2.72 Å) | | Cite: | Structural basis of rapid actin dynamics in the evolutionarily divergent Leishmania parasite.
Nat Commun, 13, 2022
|
|
7Q8S
 
 | | Leishmania major ADP-actin filament decorated with Leishmania major cofilin | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADF/Cofilin, Actin, ... | | Authors: | Kotila, T, Muniyandi, S, Lappalainen, P, Huiskonen, J.T. | | Deposit date: | 2021-11-11 | | Release date: | 2022-05-18 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural basis of rapid actin dynamics in the evolutionarily divergent Leishmania parasite.
Nat Commun, 13, 2022
|
|
5T25
 
 | | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli. | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase, LYSINE, SODIUM ION | | Authors: | Chooback, L, Thomas, L.M, Karsten, W.E, Fleming, C.D, Seabourn, P. | | Deposit date: | 2016-08-23 | | Release date: | 2017-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.991 Å) | | Cite: | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli.
To Be Published
|
|
4ER5
 
 | | Crystal structure of human DOT1L in complex with 2 molecules of EPZ004777 | | Descriptor: | 7-{5-[(3-{[(4-tert-butylphenyl)carbamoyl]amino}propyl)(propan-2-yl)amino]-5-deoxy-beta-D-ribofuranosyl}-7H-pyrrolo[2,3-d]pyrimidin-4-amine, Histone-lysine N-methyltransferase, H3 lysine-79 specific, ... | | Authors: | Wernimont, A.K, Tempel, W, Yu, W, Scopton, A, Li, Y, Nguyen, K.T, Federation, A, Marineau, J, Qi, J, Vedadi, M, Bradner, J.E, Schapira, M, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Brown, P.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2012-04-19 | | Release date: | 2012-05-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Catalytic site remodelling of the DOT1L methyltransferase by selective inhibitors.
Nat Commun, 3, 2012
|
|
4ESF
 
 | |
3F49
 
 | |
4ESK
 
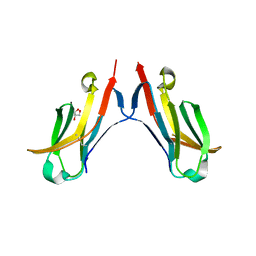 | |
4Y2E
 
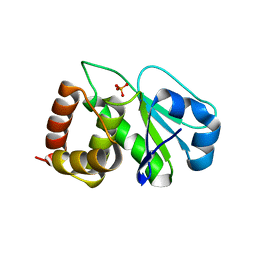 | | Crystal structure of the catalytic domain of human dual-specificity phosphatase 7 (C232S) | | Descriptor: | Dual specificity protein phosphatase 7, PHOSPHATE ION | | Authors: | Lountos, G.T, Austin, B.P, Tropea, J.E, Waugh, D.S. | | Deposit date: | 2015-02-09 | | Release date: | 2015-06-03 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Structure of human dual-specificity phosphatase 7, a potential cancer drug target.
Acta Crystallogr.,Sect.F, 71, 2015
|
|
2IN0
 
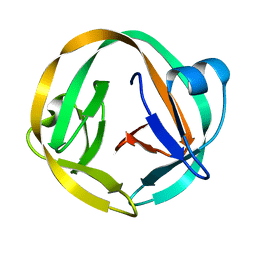 | | crystal structure of Mtu recA intein splicing domain | | Descriptor: | Endonuclease PI-MtuI | | Authors: | Van Roey, P. | | Deposit date: | 2006-10-05 | | Release date: | 2007-05-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystallographic and mutational studies of Mycobacterium tuberculosis recA mini-inteins suggest a pivotal role for a highly conserved aspartate residue.
J.Mol.Biol., 367, 2007
|
|
3F56
 
 | |
8AY1
 
 | | Crystal structure of the C. elegans POFUT2 (CePoFUT2) triple mutant (R298K-R299K-A418C) in complex with the Rattus norvegicus TSR4 single mutant (E10C) from F-spondin | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, GDP-fucose protein O-fucosyltransferase 2,Spondin-1, ... | | Authors: | Hurtado-Guerrero, R, Merino, P. | | Deposit date: | 2022-09-01 | | Release date: | 2022-10-26 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | The Essential Role of Water Molecules in the Reaction Mechanism of Protein O-Fucosyltransferase 2.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
5T11
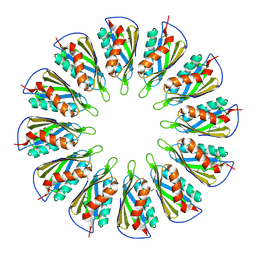 
 | |
1U7K
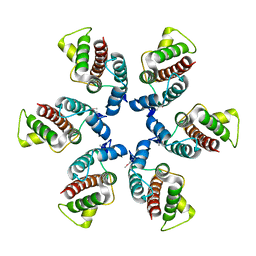 
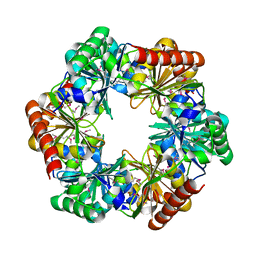
 | | Structure of a hexameric N-terminal domain from murine leukemia virus capsid | | Descriptor: | Gag polyprotein | | Authors: | Mortuza, G.B, Haire, L.F, Stevens, A, Smerdon, S.J, Stoye, J.P, Taylor, I.A. | | Deposit date: | 2004-08-04 | | Release date: | 2004-10-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | High-resolution structure of a retroviral capsid hexameric amino-terminal domain.
Nature, 431, 2004
|
|
