3AV0
 
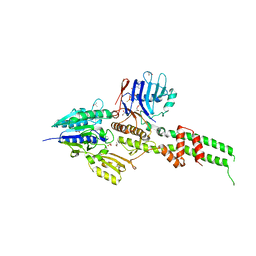 | | Crystal structure of Mre11-Rad50 bound to ATP S | | Descriptor: | DNA double-strand break repair protein mre11, DNA double-strand break repair rad50 ATPase, GLYCEROL, ... | | Authors: | Lim, H.S, Kim, J.S, Cho, Y. | | Deposit date: | 2011-02-18 | | Release date: | 2011-05-25 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal Structure of the Mre11-Rad50-ATP S Complex: Understanding the Interplay between Mre11 and Rad50
To be Published
|
|
3AUY
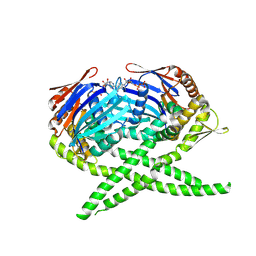 
 | | Crystal structure of Rad50 bound to ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA double-strand break repair rad50 ATPase, MAGNESIUM ION | | Authors: | Lim, H.S, Cho, Y. | | Deposit date: | 2011-02-18 | | Release date: | 2011-05-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of the Mre11-Rad50-ATP S Complex: Understanding the Interplay between Mre11 and Rad50
To be Published
|
|
3WMK
 
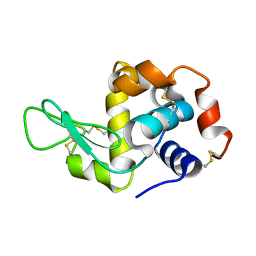 | |
3AUX
 
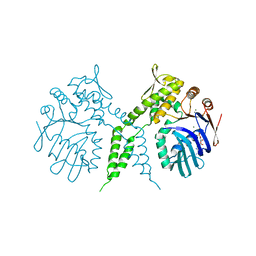 | | Crystal structure of Rad50 bound to ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA double-strand break repair rad50 ATPase, MAGNESIUM ION | | Authors: | Lim, H.S, Cho, Y. | | Deposit date: | 2011-02-17 | | Release date: | 2011-05-25 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of the Mre11-Rad50-ATP S Complex:Understanding the Interplay between Mre11 and Rad50
To be Published
|
|
2PL2
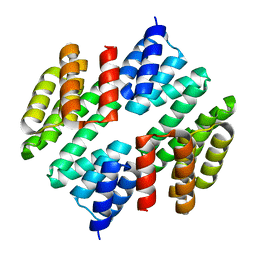 
 | | Crystal structure of TTC0263: a thermophilic TPR protein in Thermus thermophilus HB27 | | Descriptor: | Hypothetical conserved protein TTC0263 | | Authors: | Lim, H, Kim, K, Han, D, Oh, J. | | Deposit date: | 2007-04-18 | | Release date: | 2008-03-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of TTC0263, a thermophilic TPR protein from Thermus thermophilus HB27.
Mol.Cell, 24, 2007
|
|
4NHI
 
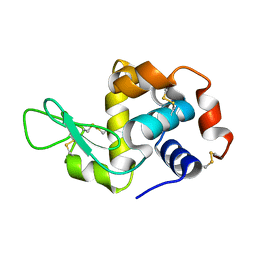 | |
5Y7W
 
 | | Crystal structure of the Nco-A1 PAS-B domain with YL-2 | | Descriptor: | Nuclear receptor coactivator 1, YL-2 peptide | | Authors: | Lee, Y.J, Yoon, H.S, Lee, J.H, Bae, J.H, Song, J.Y, Lim, H.S. | | Deposit date: | 2017-08-18 | | Release date: | 2017-11-15 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Targeted Inhibition of the NCOA1/STAT6 Protein-Protein Interaction
J. Am. Chem. Soc., 139, 2017
|
|
4WFQ
 
 | | Crystal structure of TFIIH subunit | | Descriptor: | GLYCEROL, SULFATE ION, Suppressor of stem-loop protein 1 | | Authors: | Cho, Y, Kim, J.S, Lim, H.S. | | Deposit date: | 2014-09-17 | | Release date: | 2015-02-18 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the Rad3/XPD regulatory domain of Ssl1/p44
J.Biol.Chem., 290, 2015
|
|
7BXA
 
 | | Crystal structure of PD-1 in complex with tislelizumab Fab | | Descriptor: | Programmed cell death protein 1, heavy chain, light chain | | Authors: | Heo, Y.S, Lee, S.H, Lim, H, Lee, H.T, Kim, Y.J, Park, E.B. | | Deposit date: | 2020-04-18 | | Release date: | 2020-06-10 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Crystal structure of PD-1 in complex with an antibody-drug tislelizumab used in tumor immune checkpoint therapy.
Biochem.Biophys.Res.Commun., 527, 2020
|
|
2Q7F
 
 | |
2EHO
 
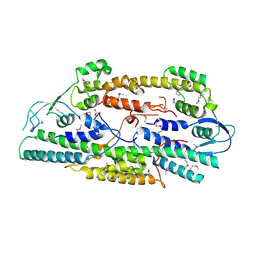 | | Crystal structure of human GINS complex | | Descriptor: | DNA replication complex GINS protein PSF1, DNA replication complex GINS protein PSF2, GINS complex subunit 3, ... | | Authors: | Choi, J.M, Lim, H.S, Kim, J.J, Song, O.K, Cho, Y. | | Deposit date: | 2007-03-07 | | Release date: | 2007-06-19 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of the human GINS complex
Genes Dev., 21, 2007
|
|
4YOA
 
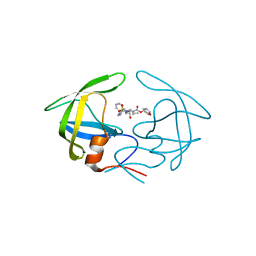 | | Crsystal structure HIV-1 Protease MDR769 L33F Complexed with darunavir | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, HIV-1 Protease | | Authors: | Kuiper, B.D, Keusch, B, Dewdney, T.G, Chordia, P, Brunzelle, J.S, Ross, K, Kovari, I.A, MacArthur, R, Salimnia, H, Kovari, L.C. | | Deposit date: | 2015-03-11 | | Release date: | 2015-07-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.697 Å) | | Cite: | The L33F darunavir resistance mutation acts as a molecular anchor reducing the flexibility of the HIV-1 protease 30s and 80s loops.
Biochem Biophys Rep, 2, 2015
|
|
4YOB
 
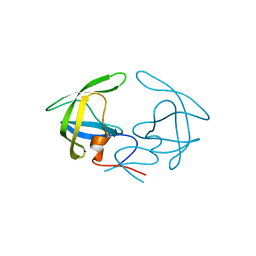 | | Crystal Structure of Apo HIV-1 Protease MDR769 L33F | | Descriptor: | HIV-1 Protease | | Authors: | Kuiper, B.D, Keusch, B, Dewdney, T.G, Chordia, P, Ross, K, Brunzelle, J.S, Kovari, I.A, MacArthur, R, Salimnia, H, Kovari, L.C. | | Deposit date: | 2015-03-11 | | Release date: | 2015-07-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.504 Å) | | Cite: | The L33F darunavir resistance mutation acts as a molecular anchor reducing the flexibility of the HIV-1 protease 30s and 80s loops.
Biochem Biophys Rep, 2, 2015
|
|
4HHL
 
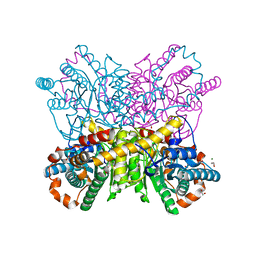 | | High resolution crystal structure of Glucose Isomerase from Streptomyces sp. SK | | Descriptor: | 1,2-ETHANEDIOL, COBALT (II) ION, MAGNESIUM ION, ... | | Authors: | Ben Hlima, H, Riguet, J, Haser, R, Aghajari, N. | | Deposit date: | 2012-10-10 | | Release date: | 2013-03-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Identification of critical residues for the activity and thermostability of Streptomyces sp. SK glucose isomerase.
Appl.Microbiol.Biotechnol., 97, 2013
|
|
4HHM
 
 | | Crystal structure of a mutant, G219A, of Glucose Isomerase from Streptomyces sp. SK | | Descriptor: | COBALT (II) ION, MAGNESIUM ION, Xylose isomerase | | Authors: | Ben Hlima, H, Riguet, J, Haser, R, Aghajari, N. | | Deposit date: | 2012-10-10 | | Release date: | 2013-03-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Identification of critical residues for the activity and thermostability of Streptomyces sp. SK glucose isomerase.
Appl.Microbiol.Biotechnol., 97, 2013
|
|
