2KIR
 
 | | Solution structure of a designer toxin, mokatoxin-1 | | Descriptor: | Designer toxin | | Authors: | Biancalana, M, Koide, A, Takacs, Z, Goldstein, S, Koide, S. | | Deposit date: | 2009-05-07 | | Release date: | 2009-12-29 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | A designer ligand specific for Kv1.3 channels from a scorpion neurotoxin-based library.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
2P4A
 
 | | X-ray structure of a camelid affinity matured single-domain vhh antibody fragment in complex with RNASE A | | Descriptor: | ANTIBODY CAB-RN05, Ribonuclease pancreatic, SULFATE ION | | Authors: | Tereshko, V, Koide, A, Uysal, S, Koide, S. | | Deposit date: | 2007-03-11 | | Release date: | 2007-08-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Exploring the capacity of minimalist protein interfaces: interface energetics and affinity maturation to picomolar KD of a single-domain antibody with a flat paratope.
J.Mol.Biol., 373, 2007
|
|
2P44
 
 | | Complex of a camelid single-domain vhh antibody fragment with RNASE A at 1.8A resolution: SE5A-mono-1 crystal form with five se-met sites (M34, M51, F68M, M83, L86M) in vhh scaffold | | Descriptor: | ANTIBODY CAB-RN05, Ribonuclease pancreatic | | Authors: | Tereshko, V, Uysal, S, Koide, A, Margalef, K, Koide, S, Kossiakoff, A.A. | | Deposit date: | 2007-03-11 | | Release date: | 2008-03-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Toward chaperone-assisted crystallography: protein engineering enhancement of crystal packing and X-ray phasing capabilities of a camelid single-domain antibody (VHH) scaffold
Protein Sci., 17, 2008
|
|
2P43
 
 | | Complex of a camelid single-domain vhh antibody fragment with RNASE A at 1.65A resolution: SE3-mono-1 crystal form with three se-met sites (M34, M51, M83) in vhh scaffold | | Descriptor: | ANTIBODY CAB-RN05, Ribonuclease pancreatic | | Authors: | Tereshko, V, Uysal, S, Koide, A, Margalef, K, Koide, S, Kossiakoff, A.A. | | Deposit date: | 2007-03-11 | | Release date: | 2008-03-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Toward chaperone-assisted crystallography: protein engineering enhancement of crystal packing and X-ray phasing capabilities of a camelid single-domain antibody (VHH) scaffold
Protein Sci., 17, 2008
|
|
2P49
 
 | | Complex of a camelid single-domain vhh antibody fragment with RNASE A at 1.4A resolution: native mono_1 crystal form | | Descriptor: | ANTIBODY CAB-RN05, PHOSPHATE ION, Ribonuclease pancreatic | | Authors: | Tereshko, V, Uysal, S, Margalef, K, Koide, A, Kossiakoff, A.A, Koide, S. | | Deposit date: | 2007-03-11 | | Release date: | 2007-08-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Exploring the capacity of minimalist protein interfaces: interface energetics and affinity maturation to picomolar KD of a single-domain antibody with a flat paratope.
J.Mol.Biol., 373, 2007
|
|
2P42
 
 | | Complex of a camelid single-domain vhh antibody fragment with RNASE A at 1.8A resolution: SE3-mono-2 crystal form with three se-met sites (M34, M51, M83) in vhh scaffold | | Descriptor: | ANTIBODY CAB-RN05, MAGNESIUM ION, Ribonuclease pancreatic | | Authors: | Tereshko, V, Uysal, S, Koide, A, Margalef, K, Koide, S, Kossiakoff, A.A. | | Deposit date: | 2007-03-11 | | Release date: | 2008-03-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Toward chaperone-assisted crystallography: protein engineering enhancement of crystal packing and X-ray phasing capabilities of a camelid single-domain antibody (VHH) scaffold
Protein Sci., 17, 2008
|
|
2ZQ3
 
 | | The crystal structure of the orthorhombic form of hen egg white lysozyme at 1.6 angstroms resolution | | Descriptor: | Lysozyme C, SODIUM ION | | Authors: | Aibara, S, Suzuki, A, Kidera, A, Shibata, K, Hirose, M. | | Deposit date: | 2008-08-03 | | Release date: | 2008-09-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The crystal structure of the orthorhombic form of hen egg white lysozyme at 1.5 angstroms resolution
To be Published
|
|
2ZQ4
 
 | | The crystal structure of the orthorhombic form of hen egg white lysozyme at 2.0 angstroms resolution | | Descriptor: | Lysozyme C | | Authors: | Aibara, S, Suzuki, A, Kidera, A, Shibata, K, Yamane, T, Hirose, M. | | Deposit date: | 2008-08-03 | | Release date: | 2008-09-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of the orthorhombic form of hen egg white lysozyme at 1.5 angstroms resolution
To be Published
|
|
3VL1
 
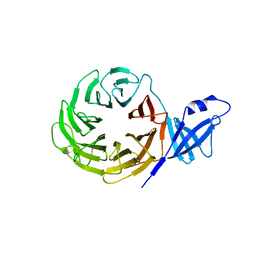 | | Crystal structure of yeast Rpn14 | | Descriptor: | 26S proteasome regulatory subunit RPN14 | | Authors: | Kim, S, Nishide, A, Saeki, Y, Takagi, K, Tanaka, K, Kato, K, Mizushima, T. | | Deposit date: | 2011-11-28 | | Release date: | 2012-05-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | New crystal structure of the proteasome-dedicated chaperone Rpn14 at 1.6 A resolution
Acta Crystallogr.,Sect.F, 68, 2012
|
|
9G8R
 
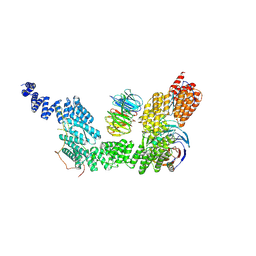 | | human SKI7-SKI238 complex in the open state | | Descriptor: | Isoform 2 of HBS1-like protein, Superkiller complex protein 2, Superkiller complex protein 3, ... | | Authors: | Koegel, A, Keidel, A, Loukeri, M.J, Kuhn, C.C, Langer, L.M, Schaefer, I.B, Conti, E. | | Deposit date: | 2024-07-23 | | Release date: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural basis of mRNA decay by the human exosome-ribosome supercomplex
Nature, 2024
|
|
9G8Q
 
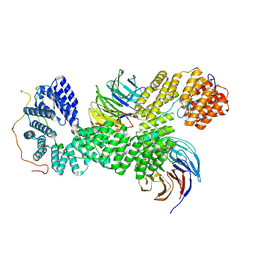 | | 40S-bound human SKI238 complex in the open state (Gatekeeping module) | | Descriptor: | Helicase SKI2W, Superkiller complex protein 3, WD repeat-containing protein 61 | | Authors: | Koegel, A, Keidel, A, Loukeri, M.J, Kuhn, C.C, Langer, L.M, Schaefer, I.B, Conti, E. | | Deposit date: | 2024-07-23 | | Release date: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Structural basis of mRNA decay by the human exosome-ribosome supercomplex
Nature, 2024
|
|
9G8N
 
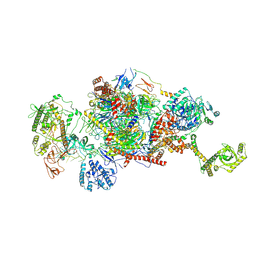 | | 80S-bound human Ski2-exosome complex | | Descriptor: | CrPV-IRES RNA, DIS3-like exonuclease 1, Exosome complex component CSL4, ... | | Authors: | Koegel, A, Keidel, A, Loukeri, M.J, Kuhn, C.C, Langer, L.M, Schaefer, I.B, Conti, E. | | Deposit date: | 2024-07-23 | | Release date: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Structural basis of mRNA decay by the human exosome-ribosome supercomplex
Nature, 2024
|
|
9G8P
 
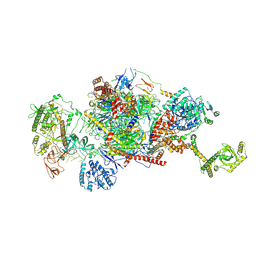 | | 40S-bound human SKI2-exosome complex | | Descriptor: | CrPV-IRES RNA, DIS3-like exonuclease 1, Exosome complex component CSL4, ... | | Authors: | Koegel, A, Keidel, A, Loukeri, M.J, Kuhn, C.C, Langer, L.M, Schaefer, I.B, Conti, E. | | Deposit date: | 2024-07-23 | | Release date: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Structural basis of mRNA decay by the human exosome-ribosome supercomplex
Nature, 2024
|
|
5FN0
 
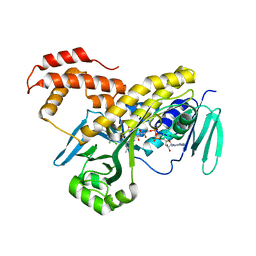 | | Crystal structure of Pseudomonas fluorescens kynurenine-3- monooxygenase (KMO) in complex with GSK180 | | Descriptor: | 3-(5,6-DICHLORO-2-OXOBENZO[D]OXAZOL-3(2H)-YL)PROPANOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, KYNURENINE 3-MONOOXYGENASE | | Authors: | Mole, D.J, Webster, S.P, Uings, I, Zheng, X, Binnie, M, Wilson, K, Hutchinson, J.P, Mirguet, O, Walker, A, Beaufils, B, Ancellin, N, Trottet, L, Beneton, V, Mowat, C.G, Wilkinson, M, Rowland, P, Haslam, C, McBride, A, Homer, N.Z.M, Baily, J.E, Sharp, M.G.F, Garden, O.J, Hughes, J, Howie, S.E.M, Holmes, D, Liddle, J, Iredale, J.P. | | Deposit date: | 2015-11-10 | | Release date: | 2016-01-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Kynurenine-3-Monooxygenase Inhibition Prevents Multiple Organ Failure in Rodent Models of Acute Pancreatitis.
Nat.Med. (N.Y.), 22, 2016
|
|
2PYF
 
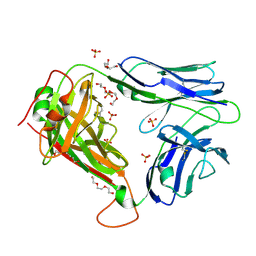 | | Crystal Structures of High Affinity Human T-Cell Receptors Bound to pMHC RevealNative Diagonal Binding Geometry Unbound TCR Clone 5-1 | | Descriptor: | SULFATE ION, T-Cell Receptor, Alpha Chain, ... | | Authors: | Sami, M, Rizkallah, P.J, Dunn, S, Li, Y, Moysey, R, Vuidepot, A, Baston, E, Todorov, P, Molloy, P, Gao, F, Boulter, J.M, Jakobsen, B.K. | | Deposit date: | 2007-05-16 | | Release date: | 2007-09-25 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of high affinity human T-cell receptors bound to peptide major
histocompatibility complex reveal native diagonal binding geometry
Protein Eng.Des.Sel., 20, 2007
|
|
4ZH5
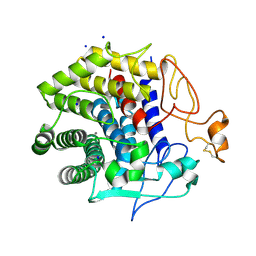 
 | | Crystal structure of Endoglucanase from Perinereis brevicirris with Cellobiose | | Descriptor: | CALCIUM ION, CHLORIDE ION, Endoglucanase, ... | | Authors: | Fewings, R.S, Swiderska, A, Sanchez-Weatherby, J, Sorensen, T.-M, Schnorr, K.M, Kneale, G.G, McGeehan, J.E. | | Deposit date: | 2015-04-24 | | Release date: | 2016-06-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Biophysical and structural characterisation of the endoglucanase from Perinereis brevicirris
To Be Published
|
|
4AJX
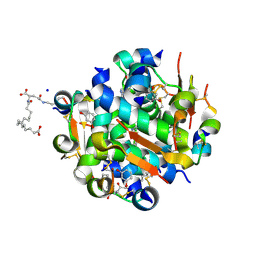 
 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | IMIDAZOLE, INSULIN, N-(16-Carboxyhexadecanoyl)-L-glutamic acid, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-20 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
7YA7
 
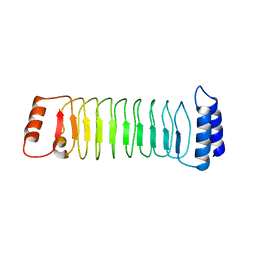 | | The crystal structure of IpaH1.4 LRR domain | | Descriptor: | RING-type E3 ubiquitin transferase | | Authors: | Hiragi, K, Nishide, A, Takagi, K, Iwai, K, Kim, M, Mizushima, T. | | Deposit date: | 2022-06-27 | | Release date: | 2023-02-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural insight into the recognition of the linear ubiquitin assembly complex by Shigella E3 ligase IpaH1.4/2.5.
J.Biochem., 173, 2023
|
|
4AKJ
 
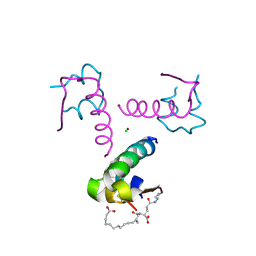 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | CHLORIDE ION, INSULIN A CHAIN, INSULIN B CHAIN, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-23 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
1M4U
 
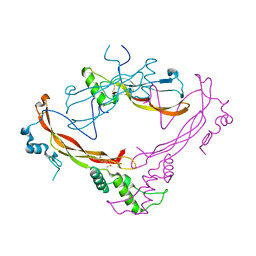 | | Crystal structure of Bone Morphogenetic Protein-7 (BMP-7) in complex with the secreted antagonist Noggin | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Bone Morphogenetic Protein-7, Noggin | | Authors: | Groppe, J, Greenwald, J, Wiater, E, Rodriguez-Leon, J, Economides, A.N, Kwiatkowski, W, Affolter, M, Vale, W.W, Izpisua-Belmonte, J.C, Choe, S. | | Deposit date: | 2002-07-03 | | Release date: | 2002-12-18 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Structural Basis of BMP Signalling Inhibition by the Cystine Knot Protein Noggin
Nature, 420, 2002
|
|
4AJZ
 
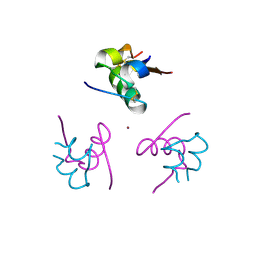 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | CHLORIDE ION, INSULIN A CHAIN, INSULIN B CHAIN, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-21 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
4AK0
 
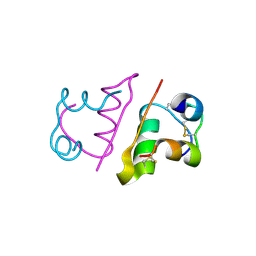 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | INSULIN A CHAIN, INSULIN B CHAIN | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-21 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
2Z33
 
 | | Solution structure of the DNA complex of PhoB DNA-binding/transactivation Domain | | Descriptor: | 5'-D(*AP*CP*AP*GP*AP*TP*TP*TP*AP*TP*GP*AP*CP*AP*GP*T)-3', 5'-D(*AP*CP*TP*GP*TP*CP*AP*TP*AP*AP*AP*TP*CP*TP*GP*T)-3', Phosphate regulon transcriptional regulatory protein phoB | | Authors: | Yamane, T, Okamura, H, Ikeguchi, M, Nishimura, Y, Kidera, A. | | Deposit date: | 2007-05-31 | | Release date: | 2008-04-22 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Water-mediated interactions between DNA and PhoB DNA-binding/transactivation domain: NMR-restrained molecular dynamics in explicit water environment.
Proteins, 71, 2008
|
|
1IWZ
 
 | | Crystal Structure Analysis of Human lysozyme at 178K. | | Descriptor: | CHLORIDE ION, LYSOZYME C | | Authors: | Joti, Y, Nakasako, M, Kidera, A, Go, N. | | Deposit date: | 2002-06-03 | | Release date: | 2002-09-04 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Nonlinear temperature dependence of the crystal structure of lysozyme: correlation between coordinate shifts and thermal factors.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1IWV
 
 | | Crystal Structure Analysis of Human lysozyme at 147K. | | Descriptor: | CHLORIDE ION, LYSOZYME C | | Authors: | Joti, Y, Nakasako, M, Kidera, A, Go, N. | | Deposit date: | 2002-06-03 | | Release date: | 2002-09-04 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Nonlinear temperature dependence of the crystal structure of lysozyme: correlation between coordinate shifts and thermal factors.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
