1W4C
 
 | | P4 protein from Bacteriophage PHI12 apo state | | Descriptor: | NTPASE P4 | | Authors: | Mancini, E.J, Kainov, D.E, Grimes, J.M, Tuma, R, Bamford, D.H, Stuart, D.I. | | Deposit date: | 2004-07-22 | | Release date: | 2004-11-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic Snapshots of an RNA Packaging Motor Reveal Conformational Changes Linking ATP Hydrolysis to RNA Translocation
Cell(Cambridge,Mass.), 118, 2004
|
|
1WAC
 
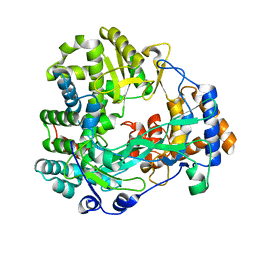 | | Back-priming mode of Phi6 RNA-dependent RNA polymerase | | Descriptor: | P2 PROTEIN | | Authors: | Laurila, M.R.L, Salgado, P.S, Stuart, D.I, Grimes, J.M, Bamford, D.H. | | Deposit date: | 2004-10-26 | | Release date: | 2005-01-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Back-Priming Mode of Phi6 RNA-Dependent RNA Polymerase
J.Gen.Virol., 86, 2005
|
|
1W00
 
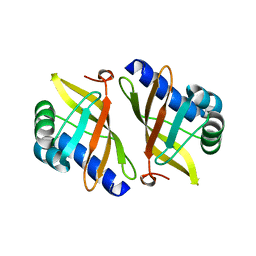 | | Crystal structure of mutant enzyme D103L of Ketosteroid Isomerase from Pseudomonas putida biotype B | | Descriptor: | STEROID DELTA-ISOMERASE | | Authors: | Kim, D.H, Jang, D.S, Nam, G.H, Oh, B.H, Choi, K.Y. | | Deposit date: | 2004-05-30 | | Release date: | 2005-05-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Double-Mutant Cycle Analysis of a Hydrogen Bond Network in Ketosteroid Isomerase from Pseudomonas Putida Biotype B
Biochem.J., 382, 2004
|
|
1VND
 
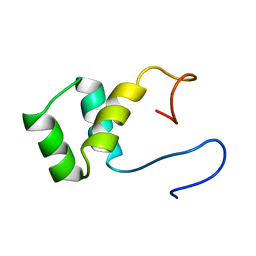 | | VND/NK-2 PROTEIN (HOMEODOMAIN), NMR | | Descriptor: | VND/NK-2 PROTEIN | | Authors: | Tsao, D.H.H, Gruschus, J.M, Wang, L.-H, Nirenberg, M, Ferretti, J.A. | | Deposit date: | 1996-05-22 | | Release date: | 1996-11-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The three-dimensional solution structure of the NK-2 homeodomain from Drosophila.
J.Mol.Biol., 251, 1995
|
|
1W48
 
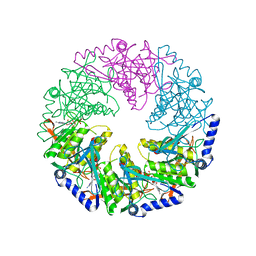 | | P4 protein from Bacteriophage PHI12 in complex with AMPcPP | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, NTPASE P4 | | Authors: | Mancini, E.J, Kainov, D.E, Grimes, J.M, Tuma, R, Bamford, D.H, Stuart, D.I. | | Deposit date: | 2004-07-22 | | Release date: | 2004-10-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Atomic Snapshots of an RNA Packaging Motor Reveal Conformational Changes Linking ATP Hydrolysis to RNA Translocation
Cell(Cambridge,Mass.), 118, 2004
|
|
1W47
 
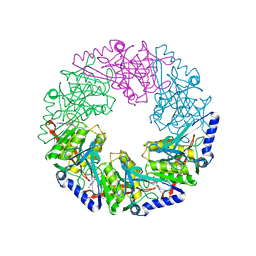 | | P4 protein from Bacteriophage PHI12 in complex with ADP and MN | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MANGANESE (II) ION, NTPASE P4 | | Authors: | Mancini, E.J, Kainov, D.E, Grimes, J.M, Tuma, R, Bamford, D.H, Stuart, D.I. | | Deposit date: | 2004-07-22 | | Release date: | 2004-11-04 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic Snapshots of an RNA Packaging Motor Reveal Conformational Changes Linking ATP Hydrolysis to RNA Translocation
Cell(Cambridge,Mass.), 118, 2004
|
|
1VQB
 
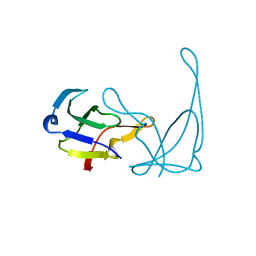 | | GENE V PROTEIN (SINGLE-STRANDED DNA BINDING PROTEIN) | | Descriptor: | GENE V PROTEIN | | Authors: | Skinner, M.M, Zhang, H, Leschnitzer, D.H, Guan, Y, Bellamy, H, Sweet, R.M, Gray, C.W, Konings, R.N.H, Wang, A.H.-J, Terwilliger, T.C. | | Deposit date: | 1996-08-14 | | Release date: | 1997-02-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of the gene V protein of bacteriophage f1 determined by multiwavelength x-ray diffraction on the selenomethionyl protein.
Proc.Natl.Acad.Sci.USA, 91, 1994
|
|
1W44
 
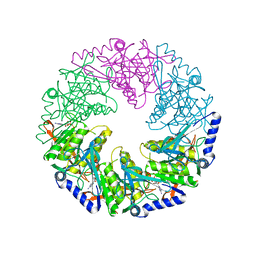 | | P4 protein from Bacteriophage PHI12 in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, NTPASE P4 | | Authors: | Mancini, E.J, Kainov, D.E, Grimes, J.M, Tuma, R, Bamford, D.H, Stuart, D.I. | | Deposit date: | 2004-07-22 | | Release date: | 2004-10-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Atomic Snapshots of an RNA Packaging Motor Reveal Conformational Changes Linking ATP Hydrolysis to RNA Translocation
Cell(Cambridge,Mass.), 118, 2004
|
|
1W4A
 
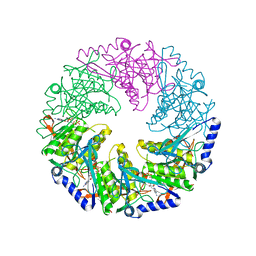 | | P4 protein from PHI12 in complex with AMPcPP and Mn | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, MANGANESE (II) ION, NTPASE P4 | | Authors: | Mancini, E.J, Kainov, D.E, Grimes, J.M, Tuma, R, Bamford, D.H, Stuart, D.I. | | Deposit date: | 2004-07-22 | | Release date: | 2004-10-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Atomic Snapshots of an RNA Packaging Motor Reveal Conformational Changes Linking ATP Hydrolysis to RNA Translocation
Cell(Cambridge,Mass.), 118, 2004
|
|
1VAD
 
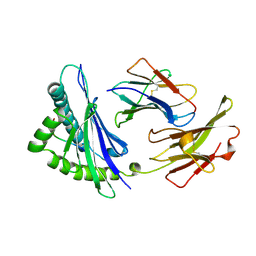 | |
1VAC
 
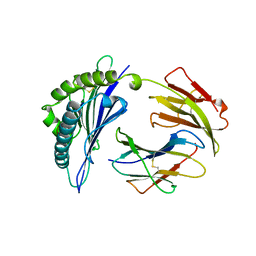 | |
1XV0
 
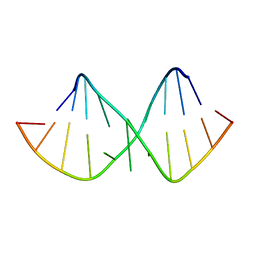 | | Solution NMR structure of RNA internal loop with three consecutive sheared GA pairs in 5'GGUGGAGGCU/3'PCCGAAGCCG | | Descriptor: | 5'-R(*GP*CP*CP*GP*AP*AP*GP*CP*CP*(P5P)-3', 5'-R(*GP*GP*UP*GP*GP*AP*GP*GP*CP*U)-3' | | Authors: | Chen, G, Znosko, B.M, Kennedy, S.D, Krugh, T.R, Turner, D.H. | | Deposit date: | 2004-10-26 | | Release date: | 2004-11-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of an RNA Internal Loop with Three Consecutive Sheared GA Pairs
Biochemistry, 44, 2005
|
|
1XF1
 
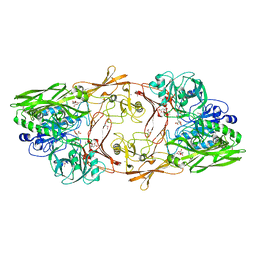 | | Structure of C5a peptidase- a key virulence factor from Streptococcus | | Descriptor: | ACETATE ION, C5a peptidase, CALCIUM ION, ... | | Authors: | Brown, C.K, Gu, Z.Y, Cleary, P.P, Matsuka, Y, Olmstead, S, Ohlendorf, D.H, Earhart, C.A. | | Deposit date: | 2004-09-13 | | Release date: | 2005-11-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the streptococcal cell wall C5a peptidase
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1XPM
 
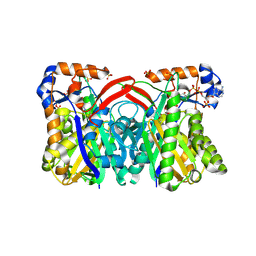 | | Crystal Structure of Staphylococcus aureus HMG-COA Synthase with HMG-CoA and Acetoacetyl-COA and Acetylated Cysteine | | Descriptor: | 3-HYDROXY-3-METHYLGLUTARYL-COENZYME A, 3-hydroxy-3-methylglutaryl CoA synthase, ACETOACETYL-COENZYME A, ... | | Authors: | Theisen, M.J, Misra, I, Saadat, D, Campobasso, N, Miziorko, H.M, Harrison, D.H.T. | | Deposit date: | 2004-10-08 | | Release date: | 2004-11-02 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | 3-hydroxy-3-methylglutaryl-CoA synthase intermediate complex observed in "real-time"
Proc.Natl.Acad.Sci.USA, 47, 2004
|
|
1XAU
 
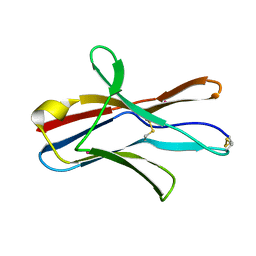 | |
1XGD
 
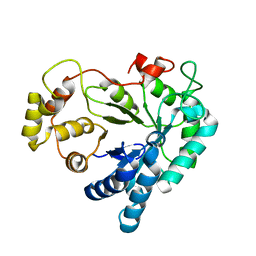 | | Apo R268A human aldose reductase | | Descriptor: | Aldose reductase | | Authors: | Brownlee, J.M, Bohren, K.M, Milne, A.C, Gabbay, K.H, Harrison, D.H.T. | | Deposit date: | 2004-09-16 | | Release date: | 2005-03-29 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structure of Apo R268A human aldose reductase: Hinges and latches that control the kinetic mechanism
Biochim.Biophys.Acta, 1748, 2005
|
|
1XAK
 
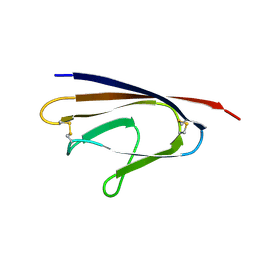 | |
1XPK
 
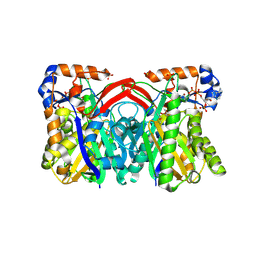 | | CRYSTAL STRUCTURE OF STAPHYLOCOCCUS AUREUS HMG-COA SYNTHASE WITH HMG-CoA AND WITH ACETOACETYL-COA AND ACETYLATED CYSTEINE | | Descriptor: | 3-HYDROXY-3-METHYLGLUTARYL-COENZYME A, 3-hydroxy-3-methylglutaryl CoA synthase, ACETOACETYL-COENZYME A, ... | | Authors: | Theisen, M.J, Misra, I, Saadat, D, Campobasso, N, Miziorko, H.M, Harrison, D.H.T. | | Deposit date: | 2004-10-08 | | Release date: | 2004-11-02 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | 3-hydroxy-3-methylglutaryl-CoA synthase intermediate complex observed in "real-time"
Proc.Natl.Acad.Sci.USA, 47, 2004
|
|
1Y3G
 
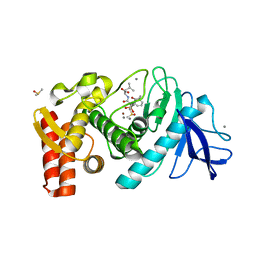 | | Crystal Structure of a Silanediol Protease Inhibitor Bound to Thermolysin | | Descriptor: | (2S)-2-{[(AMINOMETHYL)(DIHYDROXY)SILYL]METHYL}-4-METHYLPENTANAL, 3-PHENYLPROPANAL, CALCIUM ION, ... | | Authors: | Juers, D.H, Kim, J, Matthews, B.W, Sieburth, S.M. | | Deposit date: | 2004-11-24 | | Release date: | 2006-01-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Analysis of Silanediols as Transition-State-Analogue Inhibitors of the Benchmark Metalloprotease Thermolysin(,).
Biochemistry, 44, 2005
|
|
1YK8
 
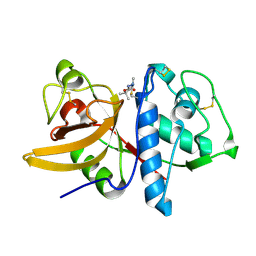 | | Cathepsin K complexed with a cyanamide-based inhibitor | | Descriptor: | Cathepsin K, TERT-BUTYL 2-CYANO-2-METHYLHYDRAZINECARBOXYLATE | | Authors: | Barrett, D.G, Deaton, D.N, Hassell, A.M, McFadyen, R.B, Miller, A.B, Miller, L.R, Payne, J.A, Shewchuk, L.M, Willard, D.H, Wright, L.L. | | Deposit date: | 2005-01-17 | | Release date: | 2005-07-19 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Acyclic cyanamide-based inhibitors of cathepsin K.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
1YK7
 
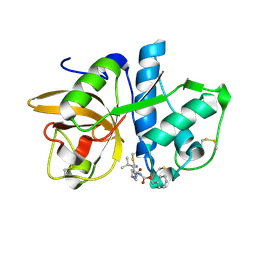 | | Cathepsin K complexed with a cyanopyrrolidine inhibitor | | Descriptor: | Cathepsin K, N2-[(BENZYLOXY)CARBONYL]-N1-[(3S)-1-CYANOPYRROLIDIN-3-YL]-L-LEUCINAMIDE | | Authors: | Barrett, D.G, Deaton, D.N, Hassell, A.M, McFadyen, R.B, Miller, A.B, Miller, L.R, Shewchuk, L.M, Tavares, F.X, Willard, D.H, Wright, L.L. | | Deposit date: | 2005-01-17 | | Release date: | 2005-03-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Novel and potent cyclic cyanamide-based cathepsin K inhibitors.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
1YKL
 
 | | Protocatechuate 3,4-Dioxygenase Y408C mutant bound to DHB | | Descriptor: | 3,4-DIHYDROXYBENZOIC ACID, FE (III) ION, Protocatechuate 3,4-dioxygenase alpha chain, ... | | Authors: | Brown, C.K, Ohlendorf, D.H. | | Deposit date: | 2005-01-18 | | Release date: | 2005-08-16 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Roles of the equatorial tyrosyl iron ligand of protocatechuate 3,4-dioxygenase in catalysis
Biochemistry, 44, 2005
|
|
1YKK
 
 | | Protocatechuate 3,4-Dioxygenase Y408C Mutant | | Descriptor: | FE (III) ION, Protocatechuate 3,4-dioxygenase alpha chain, Protocatechuate 3,4-dioxygenase beta chain | | Authors: | Brown, C.K, Ohlendorf, D.H. | | Deposit date: | 2005-01-18 | | Release date: | 2005-08-16 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Roles of the equatorial tyrosyl iron ligand of protocatechuate 3,4-dioxygenase in catalysis
Biochemistry, 44, 2005
|
|
1YKT
 
 | | Trypsin/Bpti complex mutant | | Descriptor: | CALCIUM ION, Pancreatic trypsin inhibitor, SULFATE ION, ... | | Authors: | Brown, C.K, Ohlendorf, D.H. | | Deposit date: | 2005-01-18 | | Release date: | 2006-04-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Partially folded bovine pancreatic trypsin inhibitor analogues attain fully native structures when co-crystallized with S195A rat trypsin
J.Mol.Biol., 375, 2008
|
|
1YKM
 
 | | Protocatechuate 3,4-Dioxygenase Y408E mutant | | Descriptor: | FE (III) ION, Protocatechuate 3,4-dioxygenase alpha chain, Protocatechuate 3,4-dioxygenase beta chain | | Authors: | Brown, C.K, Ohlendorf, D.H. | | Deposit date: | 2005-01-18 | | Release date: | 2005-08-16 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Roles of the equatorial tyrosyl iron ligand of protocatechuate 3,4-dioxygenase in catalysis
Biochemistry, 44, 2005
|
|
