2RGP
 
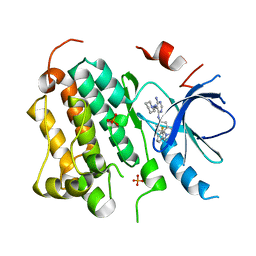 | | Structure of EGFR in complex with hydrazone, a potent dual inhibitor | | Descriptor: | Epidermal growth factor receptor, N-[1-(3-fluorobenzyl)-1H-indazol-5-yl]-5-[(piperidin-1-ylamino)methyl]pyrimidine-4,6-diamine, PHOSPHATE ION | | Authors: | Abad, M.C. | | Deposit date: | 2007-10-04 | | Release date: | 2008-08-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | 4-Amino-6-arylamino-pyrimidine-5-carbaldehyde hydrazones as potent ErbB-2/EGFR dual kinase inhibitors.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3BEL
 
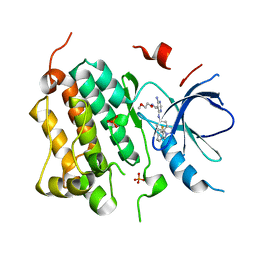 | | X-ray structure of EGFR in complex with oxime inhibitor | | Descriptor: | 4-amino-6-{[1-(3-fluorobenzyl)-1H-indazol-5-yl]amino}pyrimidine-5-carbaldehyde O-(2-methoxyethyl)oxime, Epidermal growth factor receptor, PHOSPHATE ION | | Authors: | Abad, M.C, Xu, G, Neeper, M.P, Struble, G.T, Gaul, M.D, Connolly, P.J. | | Deposit date: | 2007-11-19 | | Release date: | 2008-07-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery of novel 4-amino-6-arylaminopyrimidine-5-carbaldehyde oximes as dual inhibitors of EGFR and ErbB-2 protein tyrosine kinases.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3K6P
 
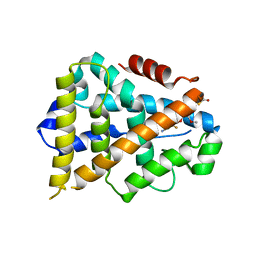 | | Estrogen Related Receptor alpha in Complex with an Ether Based Ligand | | Descriptor: | 4-(4-{[(5R)-2,4-dioxo-1,3-thiazolidin-5-yl]methyl}-2-methoxyphenoxy)-3-(trifluoromethyl)benzonitrile, Steroid hormone receptor ERR1 | | Authors: | Abad, M.C, Patch, R.J. | | Deposit date: | 2009-10-09 | | Release date: | 2010-10-20 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.996 Å) | | Cite: | Development of Diaryl Ether based ligands for Estrogen Related Receptor alpha as Potential Anti-Diabetic Agents.
To be Published
|
|
4C9Y
 
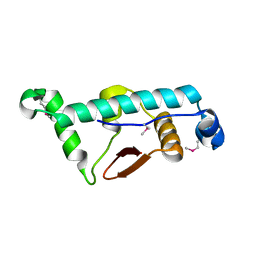 | | Structural Basis for the microtubule binding of the human kinetochore Ska complex | | Descriptor: | SPINDLE AND KINETOCHORE-ASSOCIATED PROTEIN 1 | | Authors: | Abad, M, Medina, B, Santamaria, A, Zou, J, Plasberg-Hill, C, Madhumalar, A, Jayachandran, U, Redli, P.M, Rappsilber, J, Nigg, E.A, Jeyaprakash, A.A. | | Deposit date: | 2013-10-04 | | Release date: | 2014-01-22 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural Basis for Microtubule Recognition by the Human Kinetochore Ska Complex.
Nat.Commun., 5, 2014
|
|
3QA2
 
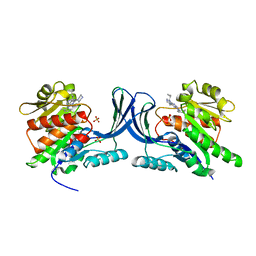 | | X-ray Structure of ketohexokinase in complex with a pyrimidopyrimidine analog 2 | | Descriptor: | Ketohexokinase, N~8~-(cyclopropylmethyl)-N~4~-(2-methylphenyl)-2-(piperazin-1-yl)pyrimido[5,4-d]pyrimidine-4,8-diamine, SULFATE ION | | Authors: | Abad, M.C. | | Deposit date: | 2011-01-10 | | Release date: | 2012-01-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.519 Å) | | Cite: | Inhibitors of Ketohexokinase: Discovery of Pyrimidinopyrimidines with Specific Substitution that Complements the ATP-Binding Site.
ACS Med Chem Lett, 2, 2011
|
|
3RO4
 
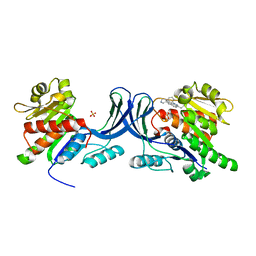 | |
4CA0
 
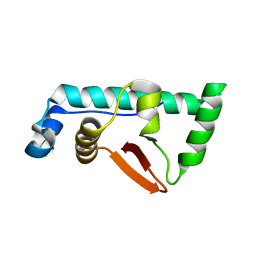 | | Structural Basis for the microtubule binding of the human kinetochore Ska complex | | Descriptor: | SPINDLE AND KINETOCHORE-ASSOCIATED PROTEIN 1 | | Authors: | Abad, M, Medina, B, Santamaria, A, Zou, J, Plasberg-Hill, C, Madhumalar, A, Jayachandran, U, Redli, P.M, Rappsilber, J, Nigg, E.A, Jeyaprakash, A.A. | | Deposit date: | 2013-10-04 | | Release date: | 2014-01-22 | | Method: | X-RAY DIFFRACTION (2.259 Å) | | Cite: | Structural Basis for Microtubule Recognition by the Human Kinetochore Ska Complex.
Nat.Commun., 5, 2014
|
|
2RA0
 
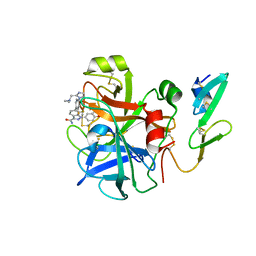 | | X-ray Structure of FXa in complex with 7-fluoroindazole | | Descriptor: | 1-(3-amino-1,2-benzisoxazol-5-yl)-6-(4-{2-[(dimethylamino)methyl]-1H-imidazol-1-yl}-2-fluorophenyl)-7-fluoro-1H-indazole-3-carboxamide, Coagulation factor X | | Authors: | Abad, M.C. | | Deposit date: | 2007-09-14 | | Release date: | 2008-01-29 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | 7-fluoroindazoles as potent and selective inhibitors of factor xa.
J.Med.Chem., 51, 2008
|
|
1KI0
 
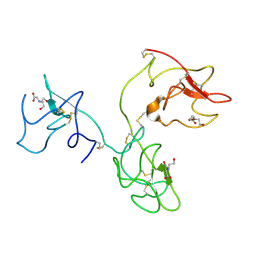 | | The X-ray Structure of Human Angiostatin | | Descriptor: | ANGIOSTATIN, BICINE | | Authors: | Abad, M.C, Arni, R.K, Grella, D.K, Castellino, F.J, Tulinsky, A, Geiger, J.H. | | Deposit date: | 2001-12-02 | | Release date: | 2002-05-29 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The X-ray crystallographic structure of the angiogenesis inhibitor angiostatin.
J.Mol.Biol., 318, 2002
|
|
2P7G
 
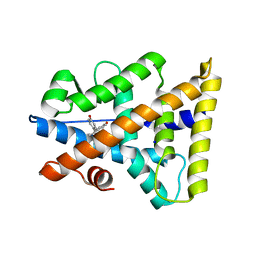 | | X-ray Structure of Estrogen Related Receptor g in complex with Bisphenol A. | | Descriptor: | 4,4'-PROPANE-2,2-DIYLDIPHENOL, Estrogen-related receptor gamma, SODIUM ION | | Authors: | Abad, M.C. | | Deposit date: | 2007-03-20 | | Release date: | 2008-02-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural determination of estrogen-related receptor gamma in the presence of phenol derivative compounds.
J.Steroid Biochem.Mol.Biol., 108, 2008
|
|
2P7A
 
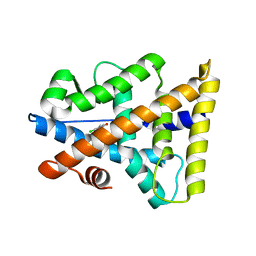 | |
2P7Z
 
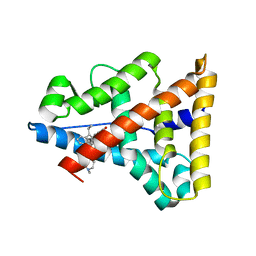 | | Estrogen Related Receptor Gamma in complex with 4-hydroxy-tamoxifen | | Descriptor: | 4-HYDROXYTAMOXIFEN, Estrogen-related receptor gamma | | Authors: | Abad, M.C. | | Deposit date: | 2007-03-21 | | Release date: | 2008-02-26 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural determination of estrogen-related receptor gamma in the presence of phenol derivative compounds.
J.Steroid Biochem.Mol.Biol., 108, 2008
|
|
1M7X
 
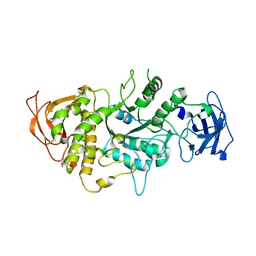 | | The X-ray Crystallographic Structure of Branching Enzyme | | Descriptor: | 1,4-alpha-glucan Branching Enzyme | | Authors: | Abad, M.C, Binderup, K, Rios-Steiner, J, Arni, R.K, Preiss, J, Geiger, J.H. | | Deposit date: | 2002-07-23 | | Release date: | 2002-09-18 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The X-ray crystallographic structure of Escherichia coli branching enzyme
J.Biol.Chem., 277, 2002
|
|
3N7O
 
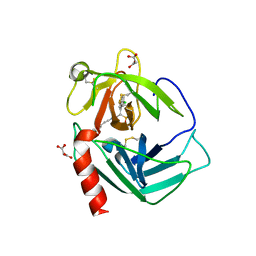 | | X-ray structure of human chymase in complex with small molecule inhibitor. | | Descriptor: | (S)-[(1S)-1-(5-chloro-1-benzothiophen-3-yl)-2-{[(E)-2-(3,4-difluorophenyl)ethenyl]amino}-2-oxoethyl]methylphosphinic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Chymase, ... | | Authors: | Abad, M.C, Kervinen, J, Crysler, C, Bayoumy, S, Spurlino, J, Deckman, I, Greco, M.N, Maryanoff, B.E, Degaravilla, L. | | Deposit date: | 2010-05-27 | | Release date: | 2010-07-21 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Potency variation of small-molecule chymase inhibitors across species.
Biochem. Pharmacol., 80, 2010
|
|
3NCA
 
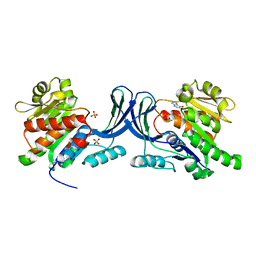 | | X-ray structure of ketohexokinase in complex with a thieno pyridinol compound | | Descriptor: | Ketohexokinase, SULFATE ION, thieno[3,2-b]pyridin-7-ol | | Authors: | Abad, M.C, Gibbs, A.C, Spurlino, J.C. | | Deposit date: | 2010-06-04 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Electron density guided fragment-based lead discovery of ketohexokinase inhibitors.
J.Med.Chem., 53, 2010
|
|
3NBV
 
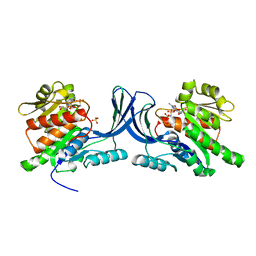 | |
3NBW
 
 | | X-ray structure of ketohexokinase in complex with a pyrazole compound | | Descriptor: | 5-amino-3-(methylsulfanyl)-1-phenyl-1H-pyrazole-4-carbonitrile, GLYCEROL, Ketohexokinase, ... | | Authors: | Abad, M.C, Gibbs, A.C, Spurlino, J.C. | | Deposit date: | 2010-06-04 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.341 Å) | | Cite: | Electron density guided fragment-based lead discovery of ketohexokinase inhibitors.
J.Med.Chem., 53, 2010
|
|
3NC2
 
 | |
3NC9
 
 | |
3QAI
 
 | | X-ray Structure of ketohexokinase in complex with a pyrimidopyrimidine analog 3 | | Descriptor: | Ketohexokinase, N~8~-(cyclopropylmethyl)-2-(2,6-diazaspiro[3.3]hept-2-yl)-N~4~-[2-(methylsulfanyl)phenyl]pyrimido[5,4-d]pyrimidine-4,8-diamine, SULFATE ION | | Authors: | Abad, M.C. | | Deposit date: | 2011-01-11 | | Release date: | 2012-01-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Inhibitors of Ketohexokinase: Discovery of Pyrimidinopyrimidines with Specific Substitution that Complements the ATP-Binding Site.
ACS Med Chem Lett, 2, 2011
|
|
3Q92
 
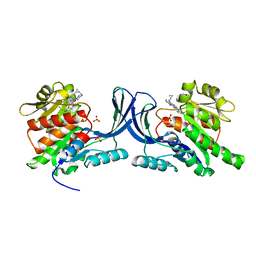 | | X-ray Structure of ketohexokinase in complex with a pyrimidopyrimidine analog 1 | | Descriptor: | Ketohexokinase, N~8~-(cyclopropylmethyl)-N~4~-[2-(methylsulfanyl)phenyl]-2-(piperazin-1-yl)pyrimido[5,4-d]pyrimidine-4,8-diamine, SULFATE ION | | Authors: | Abad, M.C. | | Deposit date: | 2011-01-07 | | Release date: | 2012-01-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Inhibitors of Ketohexokinase: Discovery of Pyrimidinopyrimidines with Specific Substitution that Complements the ATP-Binding Site.
ACS Med Chem Lett, 2, 2011
|
|
3QYA
 
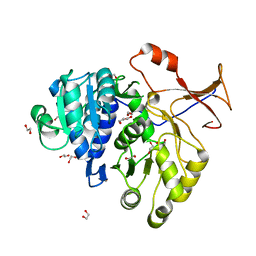 | | Crystal structure of a red-emitter mutant of Lampyris turkestanicus luciferase | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Kheirabadi, M, Gohlke, U, Hossein Khani, S, Heinemann, U, Naderi-Manesh, H. | | Deposit date: | 2011-03-03 | | Release date: | 2012-03-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Crystal structure of native and a mutant of Lampyris turkestanicus luciferase implicate in bioluminescence color shift.
Biochim.Biophys.Acta, 1834, 2013
|
|
2RDL
 
 | | Hamster Chymase 2 | | Descriptor: | Chymase 2, METHOXYSUCCINYL-ALA-ALA-PRO-ALA-CHLOROMETHYLKETONE INHIBITOR, SULFATE ION | | Authors: | Spurlino, J, Abad, M, Kervinen, J. | | Deposit date: | 2007-09-24 | | Release date: | 2007-10-30 | | Last modified: | 2012-12-12 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis for elastolytic substrate specificity in rodent alpha-chymases.
J.Biol.Chem., 283, 2008
|
|
6XWT
 
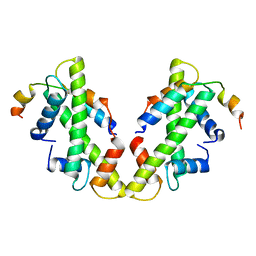 | | drosophila melanogaster CENP-A/H4 bound to N-terminal CAL1 fragment | | Descriptor: | Chromosome alignment defect 1, Histone H3-like centromeric protein cid, Histone H4 | | Authors: | Jeyaprakash, A.A, Medina-Pritchard, B, Lazou, V, Zou, J, Byron, O, Abad, M.A, Rappsilber, J, Heun, P. | | Deposit date: | 2020-01-24 | | Release date: | 2020-04-01 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.47 Å) | | Cite: | Structural basis for centromere maintenance by Drosophila CENP-A chaperone CAL1.
Embo J., 39, 2020
|
|
6XWV
 
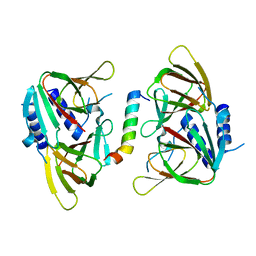 | | Crystal structure of drosophila melanogaster CENP-C bound to CAL1 | | Descriptor: | Calmodulin, Ryanodine Receptor 2 | | Authors: | Jeyaprakash, A.A, Medina-Pritchard, B, Lazou, V, Zou, J, Byron, O, Abad, M.A, Rappsilber, J, Heun, P. | | Deposit date: | 2020-01-24 | | Release date: | 2020-04-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Structural basis for centromere maintenance by Drosophila CENP-A chaperone CAL1.
Embo J., 39, 2020
|
|
