2GSS
 
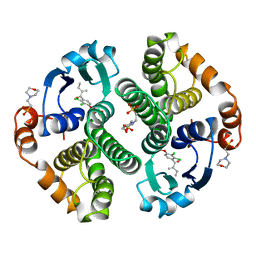 | | HUMAN GLUTATHIONE S-TRANSFERASE P1-1 IN COMPLEX WITH ETHACRYNIC ACID | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ETHACRYNIC ACID, GLUTATHIONE S-TRANSFERASE P1-1, ... | | Authors: | Oakley, A.J, Rossjohn, J, Parker, M.W. | | Deposit date: | 1996-10-29 | | Release date: | 1997-11-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The three-dimensional structure of the human Pi class glutathione transferase P1-1 in complex with the inhibitor ethacrynic acid and its glutathione conjugate.
Biochemistry, 36, 1997
|
|
3CRY
 
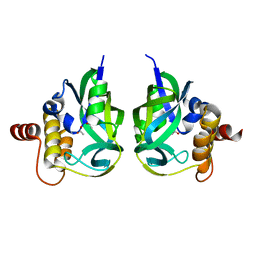 | | Gamma-glutamyl cyclotransferase | | Descriptor: | ACETATE ION, Gamma-glutamyl cyclotransferase | | Authors: | Oakley, A.J, Board, P.G. | | Deposit date: | 2008-04-08 | | Release date: | 2008-04-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The identification and structural characterization of C7orf24 as gamma-glutamyl cyclotransferase. An essential enzyme in the gamma-glutamyl cycle
J.Biol.Chem., 283, 2008
|
|
1MJ5
 
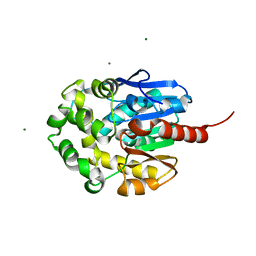 | | LINB (haloalkane dehalogenase) from sphingomonas paucimobilis UT26 at atomic resolution | | Descriptor: | 1,3,4,6-tetrachloro-1,4-cyclohexadiene hydrolase, CHLORIDE ION, MAGNESIUM ION | | Authors: | Oakley, A.J, Damborsky, J, Wilce, M.C. | | Deposit date: | 2002-08-27 | | Release date: | 2003-08-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (0.95 Å) | | Cite: | Crystal structure of haloalkane dehalogenase LinB from Sphingomonas paucimobilis UT26 at 0.95 A resolution: dynamics of catalytic residues.
Biochemistry, 43, 2004
|
|
6CBT
 
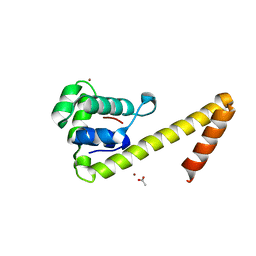 | |
6CBR
 
 | |
6CBS
 
 | |
3JUC
 
 | | Human gamma-glutamylamine cyclotransferase complex with 5-oxoproline | | Descriptor: | AIG2-like domain-containing protein 1, NITRATE ION, PYROGLUTAMIC ACID | | Authors: | Oakley, A.J. | | Deposit date: | 2009-09-15 | | Release date: | 2010-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Identification and characterization of {gamma}-glutamylamine cyclotransferase: An enzyme responsible for {gamma}-glutamyl-{epsilon}-lysine catabolism
J.Biol.Chem., 285, 2010
|
|
3JUD
 
 | | Human gamma-glutamylamine cyclotransferase, E82Q mutant | | Descriptor: | AIG2-like domain-containing protein 1, NITRATE ION | | Authors: | Oakley, A.J. | | Deposit date: | 2009-09-15 | | Release date: | 2010-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (0.98 Å) | | Cite: | Identification and characterization of {gamma}-glutamylamine cyclotransferase: An enzyme responsible for {gamma}-glutamyl-{epsilon}-lysine catabolism
J.Biol.Chem., 285, 2010
|
|
3JUB
 
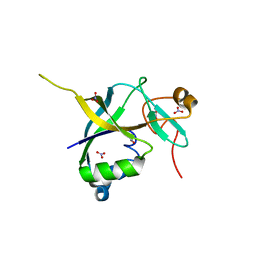 | | Human gamma-glutamylamine cyclotransferase | | Descriptor: | AIG2-like domain-containing protein 1, NITRATE ION | | Authors: | Oakley, A.J. | | Deposit date: | 2009-09-15 | | Release date: | 2010-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Identification and characterization of {gamma}-glutamylamine cyclotransferase: An enzyme responsible for {gamma}-glutamyl-{epsilon}-lysine catabolism
J.Biol.Chem., 285, 2010
|
|
1IGO
 
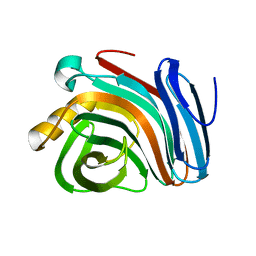 | | Family 11 xylanase | | Descriptor: | SULFATE ION, family 11 xylanase | | Authors: | Oakley, A.J, Thomson, C, Heinrich, T, Dunlop, R, Wilce, M.C.J. | | Deposit date: | 2001-04-18 | | Release date: | 2002-04-18 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Characterization of a family 11 xylanase from Bacillus subtillis B230 used for paper bleaching.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1MMI
 
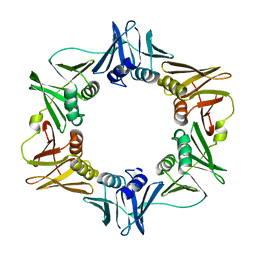 | | E. COLI DNA POLYMERASE BETA SUBUNIT | | Descriptor: | DNA polymerase III, beta chain | | Authors: | Oakley, A.J, Prosselkov, P, Wijffels, G, Beck, J.L, Wilce, M.C.J, Dixon, N.E. | | Deposit date: | 2002-09-04 | | Release date: | 2003-09-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.848 Å) | | Cite: | Flexibility revealed by the 1.85 A crystal structure of the beta sliding-clamp subunit of Escherichia coli DNA polymerase III.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
3GSS
 
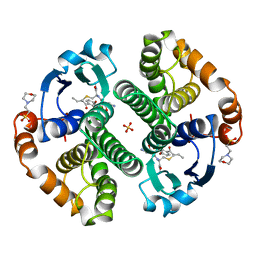 | | HUMAN GLUTATHIONE S-TRANSFERASE P1-1 IN COMPLEX WITH ETHACRYNIC ACID-GLUTATHIONE CONJUGATE | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ETHACRYNIC ACID, GLUTATHIONE, ... | | Authors: | Oakley, A.J, Rossjohn, J, Parker, M.W. | | Deposit date: | 1996-10-29 | | Release date: | 1997-11-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The three-dimensional structure of the human Pi class glutathione transferase P1-1 in complex with the inhibitor ethacrynic acid and its glutathione conjugate.
Biochemistry, 36, 1997
|
|
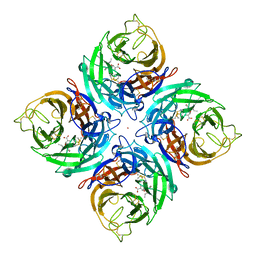
3K39
 | |
3K36
 
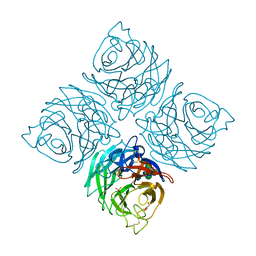 | |
3K4P
 
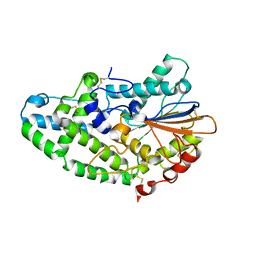 | | Aspergillus niger Phytase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A | | Authors: | Oakley, A.J. | | Deposit date: | 2009-10-06 | | Release date: | 2010-06-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The structure of Aspergillus niger phytase PhyA in complex with a phytate mimetic
Biochem.Biophys.Res.Commun., 397, 2010
|
|
3K4Q
 
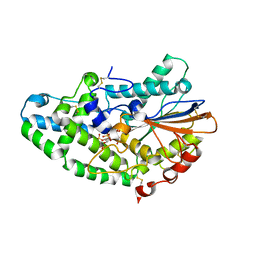 | | Aspergillus niger Phytase in complex with myo-inositol hexakis sulfate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A, D-MYO-INOSITOL-HEXASULPHATE | | Authors: | Oakley, A.J. | | Deposit date: | 2009-10-06 | | Release date: | 2010-06-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The structure of Aspergillus niger phytase PhyA in complex with a phytate mimetic
Biochem.Biophys.Res.Commun., 397, 2010
|
|
3K37
 
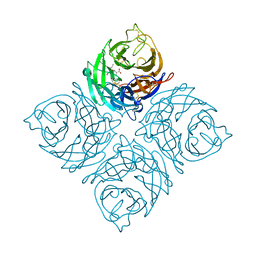 | | Crystal Structure of B/Perth Neuraminidase in complex with Peramivir | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-(1-ACETYLAMINO-2-ETHYL-BUTYL)-4-GUANIDINO-2-HYDROXY-CYCLOPENTANECARBOXYLIC ACID, CALCIUM ION, ... | | Authors: | Oakley, A.J, McKimm-Breschkin, J.L. | | Deposit date: | 2009-10-02 | | Release date: | 2010-09-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Basis of Resistance to Neuraminidase Inhibitors of Influenza B Viruses.
J.Med.Chem., 2010
|
|
3K38
 
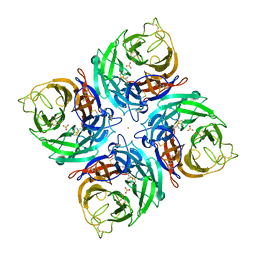 | |
3K3A
 
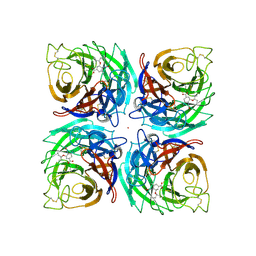 | |
1R5A
 
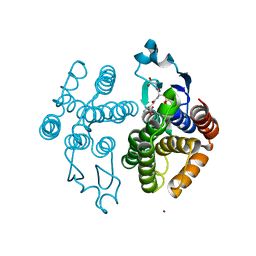 | | Glutathione S-transferase | | Descriptor: | COPPER (II) ION, GLUTATHIONE SULFONIC ACID, glutathione transferase | | Authors: | Oakley, A.J. | | Deposit date: | 2003-10-09 | | Release date: | 2003-10-28 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Identification, characterization and structure of a new Delta class glutathione transferase isoenzyme.
Biochem.J., 388, 2005
|
|
22GS
 
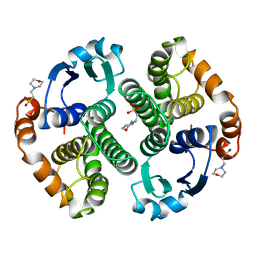 | | HUMAN GLUTATHIONE S-TRANSFERASE P1-1 Y49F MUTANT | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLUTATHIONE S-TRANSFERASE P1-1 | | Authors: | Oakley, A.J. | | Deposit date: | 1998-03-10 | | Release date: | 1999-03-23 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Thermodynamic description of the effect of the mutation Y49F on human glutathione transferase P1-1 in binding with glutathione and the inhibitor S-hexylglutathione.
J.Biol.Chem., 278, 2003
|
|
1V2A
 
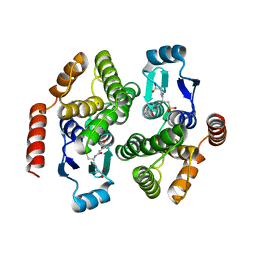 | | Glutathione S-transferase 1-6 from Anopheles dirus species B | | Descriptor: | GLUTATHIONE SULFONIC ACID, glutathione transferase gst1-6 | | Authors: | Oakley, A.J. | | Deposit date: | 2003-10-10 | | Release date: | 2003-10-28 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Identification, characterization and structure of a new Delta class glutathione transferase isoenzyme.
Biochem.J., 388, 2005
|
|
7SSG
 
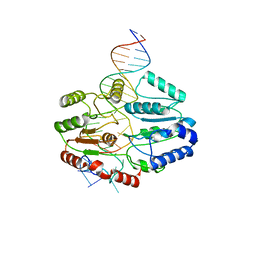 | | Mfd DNA complex | | Descriptor: | DNA (5'-D(P*TP*GP*GP*CP*GP*GP*CP*GP*AP*GP*GP*C)-3'), DNA (5'-D(P*TP*TP*GP*CP*CP*TP*CP*GP*CP*TP*GP*CP*CP*A)-3'), Transcription-repair-coupling factor | | Authors: | Oakley, A.J, Xu, Z.-Q. | | Deposit date: | 2021-11-11 | | Release date: | 2022-05-25 | | Last modified: | 2022-07-06 | | Method: | ELECTRON MICROSCOPY (5.2 Å) | | Cite: | Mechanism of transcription modulation by the transcription-repair coupling factor.
Nucleic Acids Res., 50, 2022
|
|
5V3Q
 
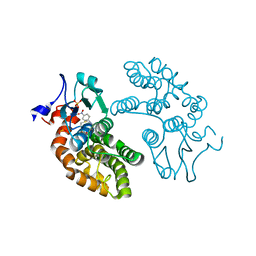 | | Human GSTO1-1 complexed with ML175 | | Descriptor: | Glutathione S-transferase omega-1, N-{3-[(2-chloro-acetyl)-(4-nitro-phenyl)-amino]-propyl}-2,2,2-trifluoro-acetamide, SULFATE ION | | Authors: | Oakley, A.J. | | Deposit date: | 2017-03-07 | | Release date: | 2017-12-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | GSTO1-1 plays a pro-inflammatory role in models of inflammation, colitis and obesity.
Sci Rep, 7, 2017
|
|
5WCE
 
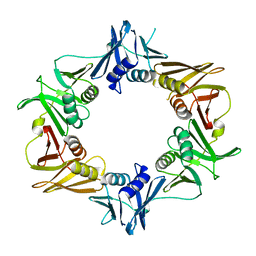 | | Caulobacter crescentus pol III beta | | Descriptor: | DNA polymerase III subunit beta | | Authors: | Oakley, A.J. | | Deposit date: | 2017-06-30 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of Pol III beta from Caulobacter crescentus
To Be Published
|
|
