8OJR
 
 | | Solution NMR Structure of Alpha-Synuclein 1-25 Peptide in 50% TFE. | | Descriptor: | Alpha-synuclein | | Authors: | Allen, S.G, Williams, C, Meade, R.M, Tang, T.M.S, Crump, M.P, Mason, J.M. | | Deposit date: | 2023-03-24 | | Release date: | 2023-08-09 | | Method: | SOLUTION NMR | | Cite: | An N-terminal alpha-Synuclein fragment binds lipid vesicles to modulate lipid induced aggregation
Cell Rep Phys Sci, 2023
|
|
6Z30
 
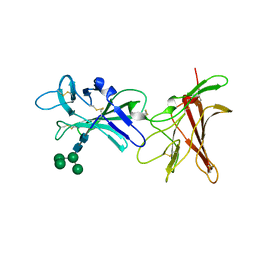 | | Human cation-independent mannose 6-phosphate/ IGF2 receptor domains 9-10 | | Descriptor: | Cation-independent mannose-6-phosphate receptor, alpha-D-mannopyranose-(1-3)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Bochel, A.J, Williams, C, Crump, M.P. | | Deposit date: | 2020-05-19 | | Release date: | 2020-08-19 | | Last modified: | 2020-12-09 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure of the Human Cation-Independent Mannose 6-Phosphate/IGF2 Receptor Domains 7-11 Uncovers the Mannose 6-Phosphate Binding Site of Domain 9.
Structure, 28, 2020
|
|
6Z31
 
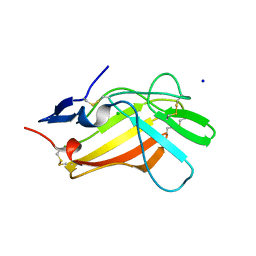 | |
6Z32
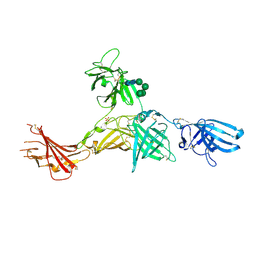 
 | | Human cation-independent mannose 6-phosphate/IGF2 receptor domains 7-11 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Cation-independent mannose-6-phosphate receptor, SULFATE ION, ... | | Authors: | Bochel, A.J, Williams, C, McCoy, A.J, Hoppe, H, Winter, A.J, Nicholls, R.D, Harlos, K, Jones, Y.E, Berger, I, Hassan, B, Crump, M.P. | | Deposit date: | 2020-05-19 | | Release date: | 2020-08-19 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.47 Å) | | Cite: | Structure of the Human Cation-Independent Mannose 6-Phosphate/IGF2 Receptor Domains 7-11 Uncovers the Mannose 6-Phosphate Binding Site of Domain 9.
Structure, 28, 2020
|
|
5NL8
 
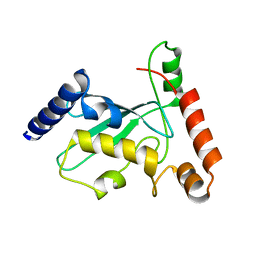 | | Pex4 of Hansenula Polymorpha | | Descriptor: | Ubiquitin-conjugating enzyme E2-21 kDa | | Authors: | Groves, M, Williams, C. | | Deposit date: | 2017-04-04 | | Release date: | 2018-01-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural insights into K48-linked ubiquitin chain formation by the Pex4p-Pex22p complex.
Biochem. Biophys. Res. Commun., 496, 2018
|
|
5IEI
 
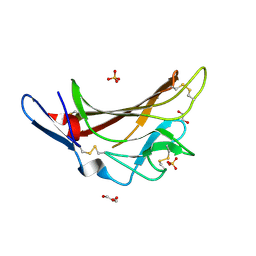 | | X-ray crystallographic structure of a high affinity IGF2 antagonist (Domain11 AB5 RHH) based on human IGF2R domain 11 | | Descriptor: | 1,2-ETHANEDIOL, Cation-independent mannose-6-phosphate receptor, GLYCEROL, ... | | Authors: | Nicholls, R.D, Williams, C, Strickland, M, Frago, S, Hassan, A.B, Crump, M.P. | | Deposit date: | 2016-02-25 | | Release date: | 2016-05-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Functional evolution of IGF2:IGF2R domain 11 binding generates novel structural interactions and a specific IGF2 antagonist.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
2FEC
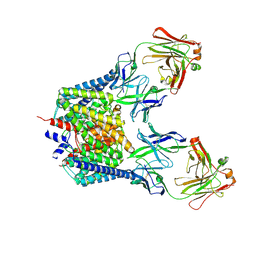 
 | | Structure of the E203Q mutant of the Cl-/H+ exchanger CLC-ec1 from E.Coli | | Descriptor: | Fab fragment, heavy chain, light chain, ... | | Authors: | Accardi, A, Walden, M.P, Nguitragool, W, Jayaram, H, Williams, C, Miller, C. | | Deposit date: | 2005-12-15 | | Release date: | 2006-01-03 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (3.967 Å) | | Cite: | Separate ion pathways in a Cl-/H+ exchanger
J.Gen.Physiol., 126, 2005
|
|
2FEE
 
 | | Structure of the Cl-/H+ exchanger CLC-ec1 from E.Coli in NaBr | | Descriptor: | Fab fragment, heavy chain, light chain, ... | | Authors: | Accardi, A, Walden, M.P, Nguitragool, W, Jayaram, H, Williams, C, Miller, C. | | Deposit date: | 2005-12-15 | | Release date: | 2006-01-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Separate ion pathways in a Cl-/H+ exchanger
J.Gen.Physiol., 126, 2005
|
|
2FED
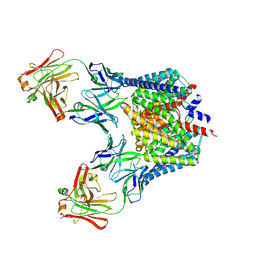 
 | | Structure of the E203Q mutant of the Cl-/H+ exchanger CLC-ec1 from E.Coli | | Descriptor: | Fab fragment, heavy chain, light chain, ... | | Authors: | Accardi, A, Walden, M.P, Nguitragool, W, Jayaram, H, Williams, C, Miller, C. | | Deposit date: | 2005-12-15 | | Release date: | 2006-01-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.317 Å) | | Cite: | Separate ion pathways in a Cl-/H+ exchanger
J.Gen.Physiol., 126, 2005
|
|
2HT2
 
 | | Structure of the Escherichia coli ClC chloride channel Y445H mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab fragment, heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-25 | | Release date: | 2006-09-19 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Synergism Between Halide Binding and Proton Transport in a CLC-type Exchanger.
J.Mol.Biol., 362, 2006
|
|
2HLF
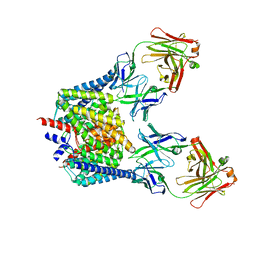 
 | | Structure of the Escherichis coli ClC chloride channel Y445E mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab Fragment, Heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-07 | | Release date: | 2006-09-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Synergism Between Halide Binding and Proton Transport in a CLC-type Exchanger
J.Mol.Biol., 362, 2006
|
|
2HT3
 
 | | Structure of the Escherichia coli ClC chloride channel Y445L mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab fragment, Heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-25 | | Release date: | 2006-09-19 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Synergism between halide binding and proton transport in a CLC-type exchanger
J.Mol.Biol., 362, 2006
|
|
2HT4
 
 | | Structure of the Escherichia coli ClC chloride channel Y445W mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab fragment, Heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-25 | | Release date: | 2006-09-19 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Synergism Between Halide Binding and Proton Transport in a CLC-type Exchanger.
J.Mol.Biol., 362, 2006
|
|
2HTL
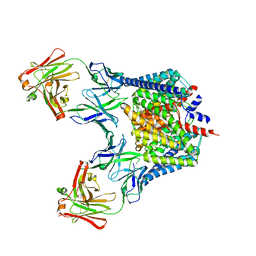 
 | | Structure of the Escherichia coli ClC chloride channel Y445F mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab fragment, Heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-26 | | Release date: | 2006-09-19 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Synergism Between Halide Binding and Proton Transport in a CLC-type Exchanger.
J.Mol.Biol., 362, 2006
|
|
2HTK
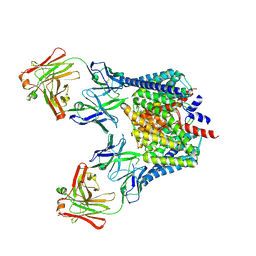 
 | | Structure of the Escherichia coli ClC chloride channel Y445A mutant and Fab complex | | Descriptor: | BROMIDE ION, Fab fragment, Heavy chain, ... | | Authors: | Accardi, A, Lobet, S, Williams, C, Miller, C, Dutzler, R. | | Deposit date: | 2006-07-26 | | Release date: | 2006-09-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.41 Å) | | Cite: | Synergism between halide binding and proton transport in a CLC-type exchanger
J.Mol.Biol., 362, 2006
|
|
4I8A
 
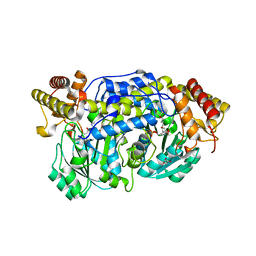 | | Alanine-glyoxylate aminotransferase variant S187F | | Descriptor: | GLYCEROL, Serine-pyruvate aminotransferase | | Authors: | Fodor, K, Oppici, E, Williams, C, Cellini, B, Wilmanns, M. | | Deposit date: | 2012-12-03 | | Release date: | 2013-05-01 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the S187F variant of human liver alanine: Aminotransferase associated with primary hyperoxaluria type I and its functional implications.
Proteins, 81, 2013
|
|
2B4K
 
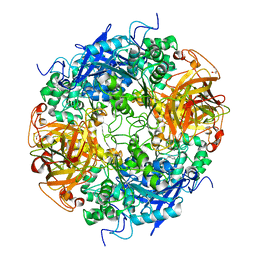 | | Acetobacter turbidans alpha-amino acid ester hydrolase complexed with phenylglycine | | Descriptor: | Alpha-amino acid ester hydrolase, D-PHENYLGLYCINE, GLYCEROL | | Authors: | Barends, T.R.M, Polderman-Tijmes, J.J, Jekel, P.A, Williams, C, Wybenga, G, Janssen, D.B, Dijkstra, B.W. | | Deposit date: | 2005-09-26 | | Release date: | 2005-12-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Acetobacter turbidans alpha-amino acid ester hydrolase: how a single mutation improves an antibiotic-producing enzyme.
J.Biol.Chem., 281, 2006
|
|
2CNR
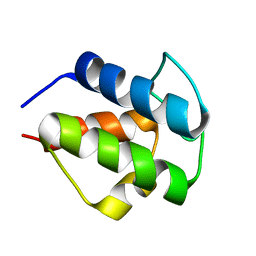 
 | |
2L22
 
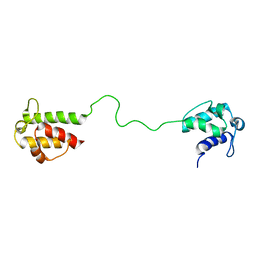 | | Mupirocin didomain ACP | | Descriptor: | Mupirocin didomain Acyl Carrier Protein | | Authors: | Dong, X, Williams, C, Crump, M.P, Wattana-amorn, P. | | Deposit date: | 2010-08-10 | | Release date: | 2012-02-15 | | Last modified: | 2013-11-06 | | Method: | SOLUTION NMR | | Cite: | A conserved motif flags acyl carrier proteins for beta-branching in polyketide synthesis.
Nat.Chem.Biol., 9, 2013
|
|
2LP8
 
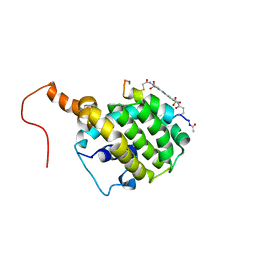 | | SOLUTION STRUCTURE OF AN APOPTOSIS ACTIVATING PHOTOSWITCHABLE BAK PEPTIDE BOUND to BCL-XL | | Descriptor: | 3,3'-(E)-diazene-1,2-diylbis{6-[(chloroacetyl)amino]benzenesulfonic acid}, Bcl-2 homologous antagonist/killer, Bcl-2-like protein 1 | | Authors: | Wysoczanski, P, Mart, R.J, Loveridge, J.E, Williams, C, Whittaker, S.B.-M, Crump, M.P, Allemann, R.K. | | Deposit date: | 2012-02-03 | | Release date: | 2012-04-18 | | Last modified: | 2021-08-18 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of a Photoswitchable Apoptosis Activating Bak Peptide Bound to Bcl-x(L).
J.Am.Chem.Soc., 134, 2012
|
|
2M6T
 
 | | NMR solution structure ensemble of 3-4D mutant domain 11 IGF2R | | Descriptor: | Insulin-like growth factor 2 receptor variant | | Authors: | Strickland, M, Williams, C, Richards, E, Minnall, L, Crump, M.P, Frago, S, Hughes, J, Garner, L, Hoppe, H, Rezgui, D, Zaccheo, O.J, Prince, S.N, Hassan, A.B, Whittaker, S. | | Deposit date: | 2013-04-09 | | Release date: | 2014-10-15 | | Last modified: | 2016-06-01 | | Method: | SOLUTION NMR | | Cite: | Functional evolution of IGF2:IGF2R domain 11 binding generates novel structural interactions and a specific IGF2 antagonist.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
2M68
 
 | | NMR solution structure ensemble of 3-4D mutant domain 11 IGF2R in complex with IGF2 (domain 11 structure only) | | Descriptor: | Insulin-like growth factor 2 receptor variant | | Authors: | Strickland, M, Williams, C, Richards, E, Minnall, L, Crump, M.P, Frago, S, Hughes, J, Garner, L, Hoppe, H, Rezgui, D, Zaccheo, O.J, Prince, S.N, Hassan, A.B, Whittaker, S. | | Deposit date: | 2013-03-27 | | Release date: | 2014-10-15 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Functional evolution of IGF2:IGF2R domain 11 binding generates novel structural interactions and a specific IGF2 antagonist.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
2PNG
 
 | | Type I rat fatty acid synthase acyl carrier protein (ACP) domain | | Descriptor: | Fatty acid synthase (EC 2.3.1.85) | | Authors: | Ploskon, E.A, Arthur, C.J, Evans, S.E, Williams, C, Crosby, J, Crump, M.P. | | Deposit date: | 2007-04-24 | | Release date: | 2007-06-05 | | Last modified: | 2022-08-24 | | Method: | SOLUTION NMR | | Cite: | A Mammalian type I Fatty Acid synthase acyl carrier protein domain does not sequester acyl chains.
J.Biol.Chem., 283, 2008
|
|
1Q1J
 
 | | Crystal Structure Analysis of anti-HIV-1 Fab 447-52D in complex with V3 peptide | | Descriptor: | Fab 447-52D, heavy chain, light chain, ... | | Authors: | Stanfield, R.L, Gorny, M.K, Williams, C, Zolla-Pazner, S, Wilson, I.A. | | Deposit date: | 2003-07-21 | | Release date: | 2004-02-17 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural rationale for the broad neutralization of HIV-1 by human monoclonal antibody 447-52D.
Structure, 12, 2004
|
|
5WIH
 
 | |
