1OO2
 
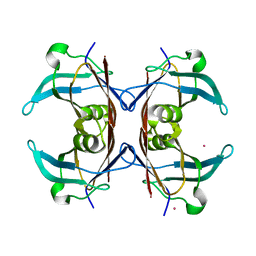 | | Crystal structure of transthyretin from Sparus aurata | | Descriptor: | CADMIUM ION, transthyretin | | Authors: | Pasquato, N, Ramazzina, I, Folli, C, Battistutta, R, Berni, R, Zanotti, G. | | Deposit date: | 2003-03-03 | | Release date: | 2004-01-20 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Distinctive binding and structural properties of piscine transthyretin.
Febs Lett., 555, 2003
|
|
6GQG
 
 | |
1NZE
 
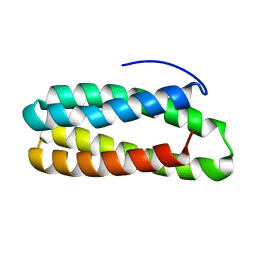 | | Crystal structure of PsbQ polypeptide of photosystem II from higher plants | | Descriptor: | Oxygen-evolving enhancer protein 3, ZINC ION | | Authors: | Calderone, V, Trabucco, M, Vujicic, A, Battistutta, R, Giacometti, G.M, Andreucci, F, Barbato, R, Zanotti, G. | | Deposit date: | 2003-02-17 | | Release date: | 2003-08-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of the PsbQ protein of photosystem II from higher plants
Embo Rep., 4, 2003
|
|
1QQF
 
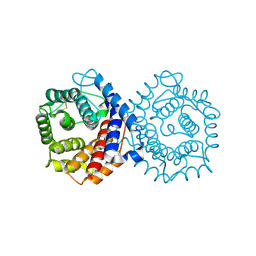 | | N-TERMINALLY TRUNCATED C3D,G FRAGMENT OF THE COMPLEMENT SYSTEM | | Descriptor: | PROTEIN (COMPLEMENT C3DG) | | Authors: | Zanotti, G, Bassetto, A, Battistutta, R, Stoppini, M, Berni, R. | | Deposit date: | 1999-06-04 | | Release date: | 2000-07-31 | | Last modified: | 2015-01-21 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure at 1.44 A resolution of an N-terminally truncated form of the rat serum complement C3d fragment.
Biochim.Biophys.Acta, 1478, 2000
|
|
6GQH
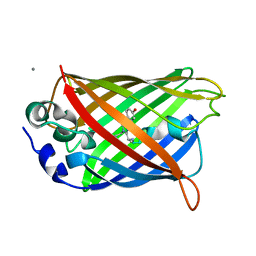 
 | |
1M2R
 
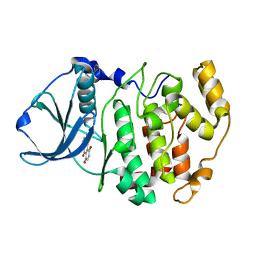 | | Crystal structure of 5,8-di-amino-1,4-di-hydroxy-anthraquinone/CK2 kinase complex | | Descriptor: | 5,8-DI-AMINO-1,4-DIHYDROXY-ANTHRAQUINONE, CASEIN kinase II, alpha chain | | Authors: | De Moliner, E, Moro, S, Sarno, S, Zagotto, G, Zanotti, G, Pinna, L.A, Battistutta, R. | | Deposit date: | 2002-06-25 | | Release date: | 2003-06-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Inhibition of protein kinase CK2 by anthraquinone-related compounds. A
structural insight
J.Biol.Chem., 278, 2003
|
|
1M2P
 
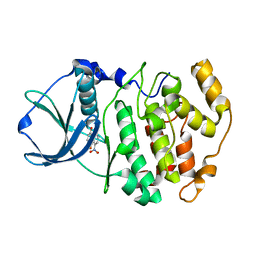 | | Crystal structure of 1,8-di-hydroxy-4-nitro-anthraquinone/CK2 kinase complex | | Descriptor: | 1,8-DI-HYDROXY-4-NITRO-ANTHRAQUINONE, Casein kinase II, alpha chain | | Authors: | De Moliner, E, Moro, S, Sarno, S, Zagotto, G, Zanotti, G, Pinna, L.A, Battistutta, R. | | Deposit date: | 2002-06-25 | | Release date: | 2003-06-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Inhibition of protein kinase CK2 by anthraquinone-related compounds. A
structural insight
J.Biol.Chem., 278, 2003
|
|
1M2Q
 
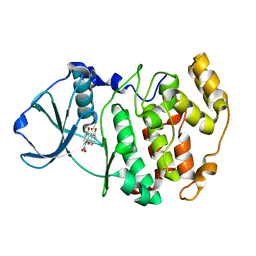 | | Crystal structure of 1,8-di-hydroxy-4-nitro-xanten-9-one/CK2 kinase complex | | Descriptor: | 1,8-DI-HYDROXY-4-NITRO-XANTHEN-9-ONE, Casein kinase II, alpha chain | | Authors: | De Moliner, E, Sarno, S, Moro, S, Zagotto, G, Zanotti, G, Pinna, L.A, Battistutta, R. | | Deposit date: | 2002-06-25 | | Release date: | 2003-06-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Inhibition of protein kinase CK2 by anthraquinone-related compounds. A
structural insight
J.Biol.Chem., 278, 2003
|
|
3PVG
 
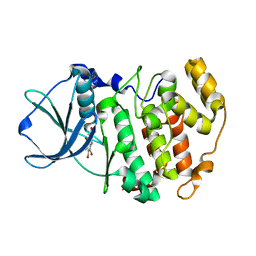 | | Crystal structure of Z. mays CK2 alpha subunit in complex with the inhibitor 4,5,6,7-tetrabromo-1-carboxymethylbenzimidazole (K68) | | Descriptor: | (4,5,6,7-tetrabromo-1H-benzimidazol-1-yl)acetic acid, Casein kinase II subunit alpha | | Authors: | Papinutto, E, Franchin, C, Battistutta, R. | | Deposit date: | 2010-12-07 | | Release date: | 2010-12-15 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | ATP site-directed inhibitors of protein kinase CK2: an update.
Curr Top Med Chem, 11, 2011
|
|
1LTR
 
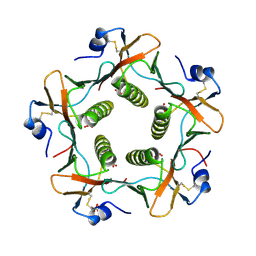 | | CRYSTAL STRUCTURE OF THE B SUBUNIT OF HUMAN HEAT-LABILE ENTEROTOXIN FROM E. COLI CARRYING A PEPTIDE WITH ANTI-HSV ACTIVITY | | Descriptor: | HEAT-LABILE ENTEROTOXIN, SULFATE ION | | Authors: | Matkovic-Calogovic, D, Loreggian, A, Palu, G, Zanotti, G. | | Deposit date: | 1998-07-31 | | Release date: | 1999-02-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (3.04 Å) | | Cite: | Crystal structure of the B subunit of Escherichia coli heat-labile enterotoxin carrying peptides with anti-herpes simplex virus type 1 activity.
J.Biol.Chem., 274, 1999
|
|
3CWY
 
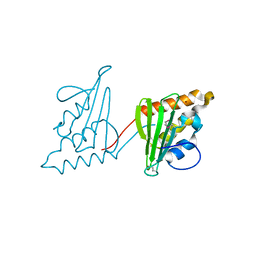 | | Structure of CagD from H. pylori pathogenicity island crystallized in the presence of Cu(II) ions | | Descriptor: | COPPER (II) ION, protein CagD | | Authors: | Cendron, L, Zanotti, G, Angelini, A, Barison, N, Couturier, M, Stein, M. | | Deposit date: | 2008-04-23 | | Release date: | 2008-12-30 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | The Helicobacter pylori CagD (HP0545, Cag24) protein is essential for CagA translocation and maximal induction of interleukin-8 secretion.
J.Mol.Biol., 386, 2009
|
|
3CWX
 
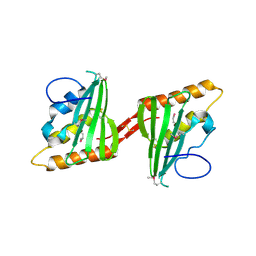 | | Crystal structure of cagd from helicobacter pylori pathogenicity island | | Descriptor: | protein CagD | | Authors: | Cendron, L, Zanotti, G, Angelini, A, Barison, N, Couturier, M, Stein, M. | | Deposit date: | 2008-04-23 | | Release date: | 2008-12-30 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Helicobacter pylori CagD (HP0545, Cag24) protein is essential for CagA translocation and maximal induction of interleukin-8 secretion.
J.Mol.Biol., 386, 2009
|
|
1O9R
 
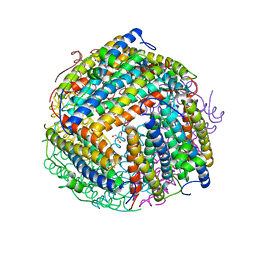 | | The X-ray crystal structure of Agrobacterium tumefaciens Dps, a member of the family that protect DNA without binding | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, AGROBACTERIUM TUMEFACIENS DPS, ... | | Authors: | Ilari, A, Ceci, P, Chiancone, E. | | Deposit date: | 2002-12-18 | | Release date: | 2003-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | The Dps Protein of Agrobacterium Tumefaciens Does not Bind to DNA But Protects It Toward Oxidative Cleavage: X-Ray Crystal Structure, Iron Binding, and Hydroxyl-Radical Scavenging Properties
J.Biol.Chem., 278, 2003
|
|
2PNC
 
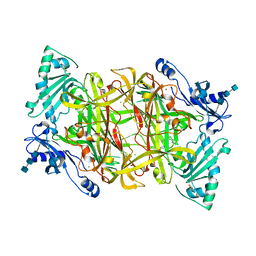 | | Crystal Structure of Bovine Plasma Copper-Containing Amine Oxidase in Complex with Clonidine | | Descriptor: | 2,6-DICHLORO-N-IMIDAZOLIDIN-2-YLIDENEANILINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Cendron, L, Holt, A, Smith, D.J, Zanotti, G, Rigo, A, Di Paolo, M.L. | | Deposit date: | 2007-04-24 | | Release date: | 2008-02-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Multiple binding sites for substrates and modulators of semicarbazide-sensitive amine oxidases: kinetic consequences
Mol.Pharmacol., 73, 2008
|
|
6RCB
 
 | |
6HOV
 
 | |
2PYB
 
 | |
1NA7
 
 | |
6ISJ
 
 | | Crystal structure of CX-4945 bound CK2 alpha from C. neoformans | | Descriptor: | 5-[(3-chlorophenyl)amino]benzo[c][2,6]naphthyridine-8-carboxylic acid, Casein kinase II subunit alpha, SULFATE ION | | Authors: | Cho, H.S. | | Deposit date: | 2018-11-16 | | Release date: | 2019-11-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of CX-4945 bound CK2 alpha from C. neoformans
To be published
|
|
