6HXP
 
 | |
6HXN
 
 | |
6HXL
 
 | |
6HXQ
 
 | |
6HXK
 | |
6HXM
 
 | |
1AL3
 
 | |
1EDD
 
 | |
1EDE
 
 | |
1EDB
 
 | |
2DHE
 
 | |
2DHD
 
 | |
2DHC
 
 | |
2EDC
 
 | |
2EDA
 
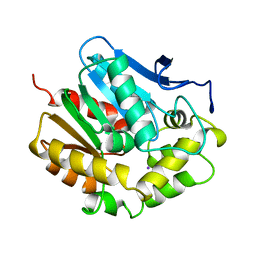 | |
7S5B
 
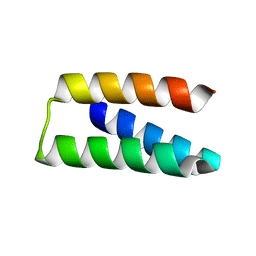 | |
7RDH
 | |
7Z3P
 
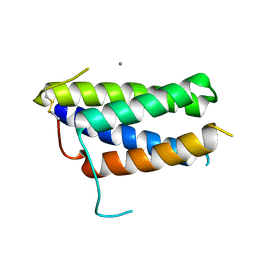 | | Crystal structure of the mouse leptin:LepR-CRH2 encounter complex to 1.95 A resolution. | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, Leptin, ... | | Authors: | Tsirigotaki, A, Verschueren, K, Savvides, S.N, Verstraete, K. | | Deposit date: | 2022-03-02 | | Release date: | 2023-03-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.943 Å) | | Cite: | Mechanism of receptor assembly via the pleiotropic adipokine Leptin.
Nat.Struct.Mol.Biol., 30, 2023
|
|
7Z3R
 
 | | Crystal structure of the mouse leptin:LepR-IgCRH2 complex to 2.95 A resolution. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Leptin, Leptin receptor | | Authors: | Verstraete, K, Verschueren, K, Savvides, S.N, Tsirigotaki, A. | | Deposit date: | 2022-03-02 | | Release date: | 2023-03-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.951 Å) | | Cite: | Mechanism of receptor assembly via the pleiotropic adipokine Leptin.
Nat.Struct.Mol.Biol., 30, 2023
|
|
7Z3Q
 
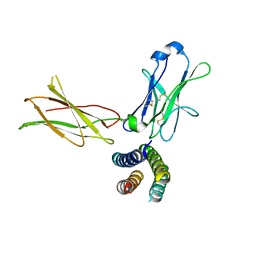 | | Crystal structure of the human leptin:LepR-CRH2 encounter complex to 3.6 A resolution. | | Descriptor: | Leptin, Leptin receptor | | Authors: | Verstraete, K, Verschueren, K, Alexandra, T, Savvides, S.N. | | Deposit date: | 2022-03-02 | | Release date: | 2023-03-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.617 Å) | | Cite: | Mechanism of receptor assembly via the pleiotropic adipokine Leptin.
Nat.Struct.Mol.Biol., 30, 2023
|
|
7N1J
 | |
7N1K
 
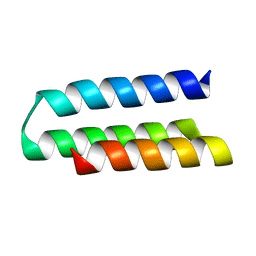 | |
7N3T
 
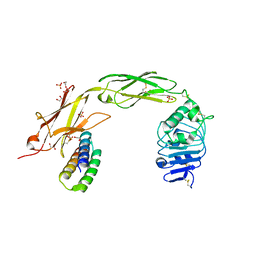 | | TrkA ECD complex with designed miniprotein ligand | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Jude, K.M, Cao, L, Garcia, K.C. | | Deposit date: | 2021-06-01 | | Release date: | 2022-04-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Design of protein-binding proteins from the target structure alone.
Nature, 605, 2022
|
|
6GLX
 
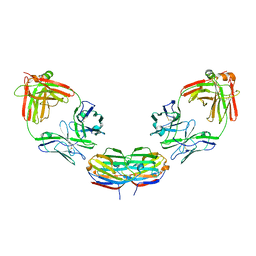 | | Structure of galectin-10 in complex with the Fab fragment of a Charcot-Leyden crystal solubilizing antibody, 4E8 | | Descriptor: | Fab 4E8 - Heavy chain, Fab 4E8 - Light chain, Galectin-10 | | Authors: | Verstraete, K, Verschueren, K, Savvides, S.N. | | Deposit date: | 2018-05-23 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.396 Å) | | Cite: | Protein crystallization promotes type 2 immunity and is reversible by antibody treatment.
Science, 364, 2019
|
|
6GKU
 
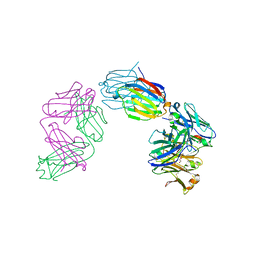 | | Structure of galectin-10 in complex with the Fab fragment of a Charcot-Leyden crystal solubilizing antibody, 6F5 | | Descriptor: | CALCIUM ION, Fab 6F5 - Heavy Chain, Fab 6F5 - Light chain, ... | | Authors: | Verstraete, K, Verschueren, K, Savvides, S.N. | | Deposit date: | 2018-05-21 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Protein crystallization promotes type 2 immunity and is reversible by antibody treatment.
Science, 364, 2019
|
|
