6UMV
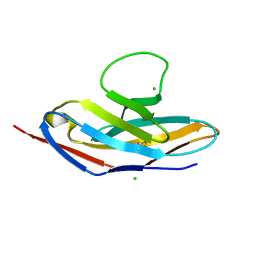 
 | | Human apo PD-1 double mutant | | Descriptor: | CHLORIDE ION, Programmed cell death protein 1 | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2019-10-10 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.424 Å) | | Cite: | A high-affinity human PD-1/PD-L2 complex informs avenues for small-molecule immune checkpoint drug discovery.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6UMT
 
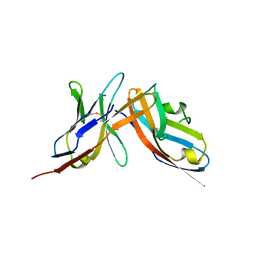 | | High-affinity human PD-1 PD-L2 complex | | Descriptor: | MAGNESIUM ION, Programmed cell death 1 ligand 2, Programmed cell death protein 1 | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2019-10-10 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.986 Å) | | Cite: | A high-affinity human PD-1/PD-L2 complex informs avenues for small-molecule immune checkpoint drug discovery.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6UMU
 
 | | Human apo PD-1 triple mutant | | Descriptor: | CHLORIDE ION, Programmed cell death protein 1 | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2019-10-10 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.183 Å) | | Cite: | A high-affinity human PD-1/PD-L2 complex informs avenues for small-molecule immune checkpoint drug discovery.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
7UX0
 
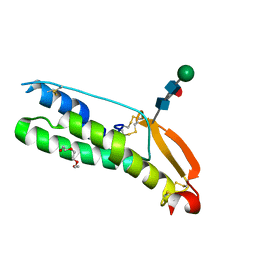 | | Human Sperm TMEM95 Ectodomain | | Descriptor: | Sperm-egg fusion protein TMEM95, TRIETHYLENE GLYCOL, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2022-05-04 | | Release date: | 2022-10-12 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Human sperm TMEM95 binds eggs and facilitates membrane fusion.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
5FD9
 
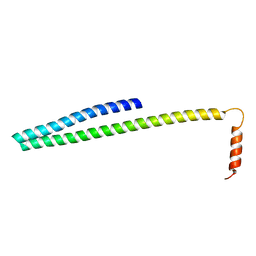 | | X-ray Crystal Structure of ESCRT-III Snf7 core domain (conformation B) | | Descriptor: | Vacuolar-sorting protein SNF7 | | Authors: | Tang, S, Henne, W.M, Borbat, P.P, Buchkovich, N.J, Freed, J.H, Mao, Y, Fromme, J.C, Emr, S.D. | | Deposit date: | 2015-12-15 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments.
Elife, 4, 2015
|
|
5FD7
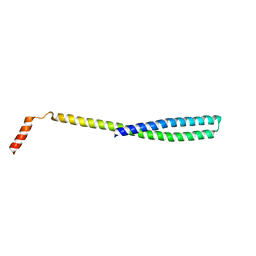 
 | | X-ray Crystal Structure of ESCRT-III Snf7 core domain (conformation A) | | Descriptor: | Vacuolar-sorting protein SNF7 | | Authors: | Tang, S, Henne, W.M, Borbat, P.P, Buchkovich, N.J, Freed, J.H, Mao, Y, Fromme, J.C, Emr, S.D. | | Deposit date: | 2015-12-15 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments.
Elife, 4, 2015
|
|
6CYJ
 
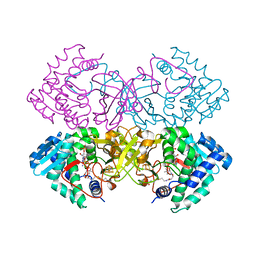 | | Mycobacterium tuberculosis transcriptional regulator | | Descriptor: | HTH-type transcriptional regulator PrpR, IRON/SULFUR CLUSTER, SUCCINYL-COENZYME A | | Authors: | Tang, S, Sacchettini, J. | | Deposit date: | 2018-04-05 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.699 Å) | | Cite: | Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.
Nucleic Acids Res., 47, 2019
|
|
6CZ6
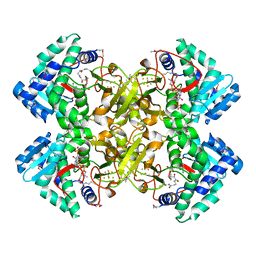 
 | | Mycobacterium tuberculosis transcriptional regulator | | Descriptor: | COENZYME A, HTH-type transcriptional regulator PrpR, IRON/SULFUR CLUSTER | | Authors: | Tang, S, Sacchettini, J. | | Deposit date: | 2018-04-08 | | Release date: | 2019-04-17 | | Last modified: | 2019-10-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.
Nucleic Acids Res., 47, 2019
|
|
6D2S
 
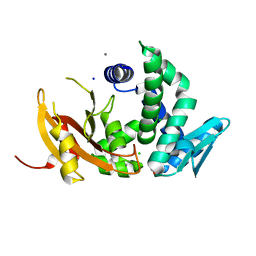 | | Mycobacterium tuberculosis transcriptional regulator | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Tang, S, Sacchettini, J. | | Deposit date: | 2018-04-13 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.819 Å) | | Cite: | Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.
Nucleic Acids Res., 47, 2019
|
|
6CYY
 
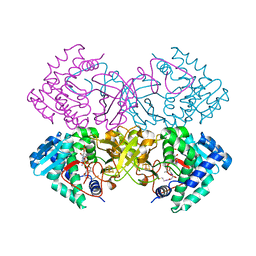 | | Mycobacterium tuberculosis transcriptional regulator | | Descriptor: | COENZYME A, HTH-type transcriptional regulator PrpR, IRON/SULFUR CLUSTER | | Authors: | Tang, S, Sacchettini, J. | | Deposit date: | 2018-04-07 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.506 Å) | | Cite: | Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.
Nucleic Acids Res., 47, 2019
|
|
6D8I
 
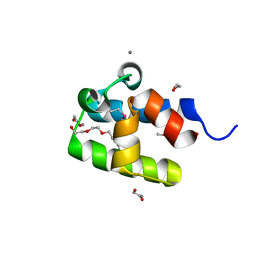 | |
6D8J
 
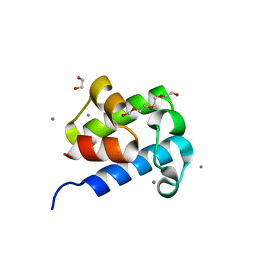 | |
7Q8T
 
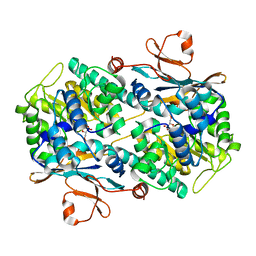 | | Crystal structure of NAMPT bound to ligand TSY535(compound 9a) | | Descriptor: | Nicotinamide phosphoribosyltransferase, SULFATE ION, [(2~{R},3~{S},4~{R},5~{S})-3,4-bis(oxidanyl)-5-[4-[[[4-(phenylsulfonyl)phenyl]carbamoylamino]methyl]phenyl]oxolan-2-yl]methyl dihydrogen phosphate | | Authors: | Kraemer, A, Tang, S, Butterworth, S, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2021-11-11 | | Release date: | 2021-11-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Chemistry-led investigations into the mode of action of NAMPT activators, resulting in the discovery of non-pyridyl class NAMPT activators.
Acta Pharm Sin B, 13, 2023
|
|
7OEX
 
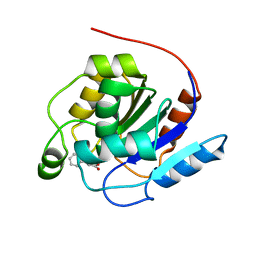 | |
4YIL
 
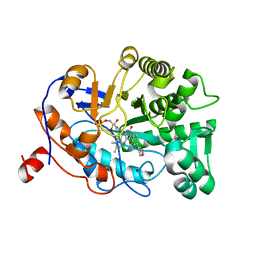 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL 3-CYANO-3-(4-FLUOROPHENYL)ACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | FLAVIN MONONUCLEOTIDE, MAGNESIUM ION, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-02 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.461 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
4YNC
 
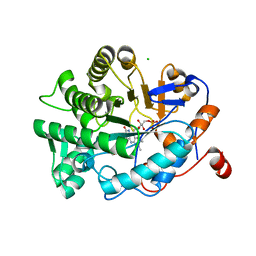 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL-3-CYANO-3-PHENYLACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-09 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.498 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
6UX6
 
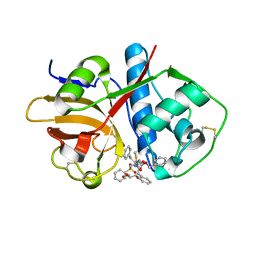 | | Cruzain covalently bound by a vinylsulfone compound | | Descriptor: | Cruzipain, GLYCEROL, Nalpha-[(benzyloxy)carbonyl]-N-[(2S)-1-phenyl-4-(phenylsulfonyl)butan-2-yl]-L-phenylalaninamide | | Authors: | Zhai, X, Tang, S, Chenna, B.C, Meek, T.D, Sacchettini, J.C. | | Deposit date: | 2019-11-06 | | Release date: | 2020-11-25 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Cruzain covalently bound by a vinylsulfone compound
To Be Published
|
|
8F3A
 
 | | HIV-1 gp41 coiled-coil pocket IQN17 | | Descriptor: | ACETIC ACID, CHLORIDE ION, IQN17 | | Authors: | Bruun, T.U.J, Tang, S, Fernandez, D, Kim, P.S. | | Deposit date: | 2022-11-09 | | Release date: | 2023-03-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structure-guided stabilization improves the ability of the HIV-1 gp41 hydrophobic pocket to elicit neutralizing antibodies.
J.Biol.Chem., 299, 2023
|
|
8F3B
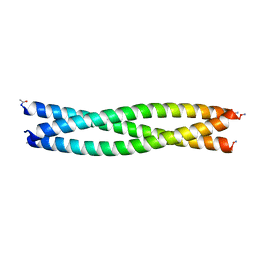 
 | |
1XMH
 
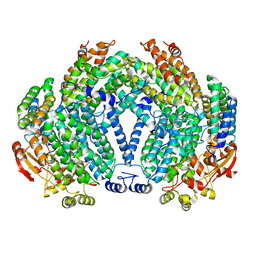 | | Structure of Co(II) reconstituted methane monooxygenase hydroxylase from M. capsulatus (Bath) | | Descriptor: | COBALT (II) ION, Methane monooxygenase component A alpha chain, Methane monooxygenase component A beta chain, ... | | Authors: | Sazinsky, M.H, Merkx, M, Cadieux, E, Tang, S, Lippard, S.J. | | Deposit date: | 2004-10-02 | | Release date: | 2005-01-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Preparation and X-ray Structures of Metal-Free, Dicobalt and Dimanganese Forms of Soluble Methane Monooxygenase Hydroxylase from Methylococcus capsulatus (Bath)
Biochemistry, 43, 2004
|
|
1XMF
 
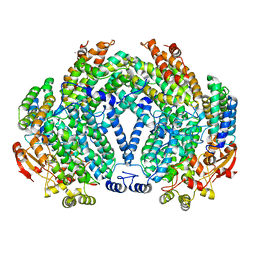 | | Structure of Mn(II)-Soaked Apo Methane Monooxygenase Hydroxylase Crystals from M. capsulatus (Bath) | | Descriptor: | CALCIUM ION, MANGANESE (II) ION, Methane monooxygenase component A alpha chain, ... | | Authors: | Sazinsky, M.H, Merkx, M, Cadieux, E, Tang, S, Lippard, S.J. | | Deposit date: | 2004-10-02 | | Release date: | 2005-01-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Preparation and X-ray Structures of Metal-Free, Dicobalt and Dimanganese Forms of Soluble Methane Monooxygenase Hydroxylase from Methylococcus capsulatus (Bath)
Biochemistry, 43, 2004
|
|
1XMG
 
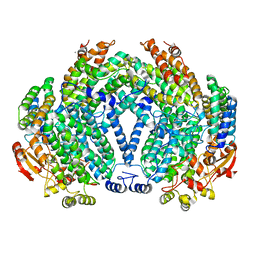 | | Crystal structure of apo methane monooxygenase hydroxylase from M. capsulatus (Bath) | | Descriptor: | CALCIUM ION, Methane monooxygenase component A alpha chain, Methane monooxygenase component A beta chain, ... | | Authors: | Sazinsky, M.H, Merkx, M, Cadieux, E, Tang, S, Lippard, S.J. | | Deposit date: | 2004-10-02 | | Release date: | 2005-01-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Preparation and X-ray Structures of Metal-Free, Dicobalt and Dimanganese Forms of Soluble Methane Monooxygenase Hydroxylase from Methylococcus capsulatus (Bath)
Biochemistry, 43, 2004
|
|
6GG8
 
 | | Mineralocorticoid receptor in complex with (s)-13 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Mineralocorticoid receptor, Nuclear receptor coactivator 1, ... | | Authors: | Edman, K, Aagaard, A, Tangefjord, S. | | Deposit date: | 2018-05-03 | | Release date: | 2019-01-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification of Mineralocorticoid Receptor Modulators with Low Impact on Electrolyte Homeostasis but Maintained Organ Protection.
J.Med.Chem., 62, 2019
|
|
6GGG
 
 | | Mineralocorticoid receptor in complex with (s)-13 | | Descriptor: | 2-[(3~{S})-7-fluoranyl-6-(2-methylpropyl)-4-[(3-oxidanylidene-4~{H}-1,4-benzoxazin-6-yl)carbonyl]-2,3-dihydro-1,4-benzoxazin-3-yl]-~{N}-methyl-ethanamide, CHLORIDE ION, Mineralocorticoid receptor, ... | | Authors: | Edman, K, Aagaard, A, Tangefjord, S. | | Deposit date: | 2018-05-03 | | Release date: | 2019-01-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Identification of Mineralocorticoid Receptor Modulators with Low Impact on Electrolyte Homeostasis but Maintained Organ Protection.
J.Med.Chem., 62, 2019
|
|
6GEV
 
 | | Mineralocorticoid receptor in complex with (s)-13 | | Descriptor: | 6-[[(3~{S})-7-fluoranyl-3-(2-methylpropyl)-2,3-dihydro-1,4-benzoxazin-4-yl]carbonyl]-4~{H}-1,4-benzoxazin-3-one, GLYCEROL, Mineralocorticoid receptor, ... | | Authors: | Edman, K, Aagaard, A, Tangefjord, S. | | Deposit date: | 2018-04-27 | | Release date: | 2019-01-09 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Identification of Mineralocorticoid Receptor Modulators with Low Impact on Electrolyte Homeostasis but Maintained Organ Protection.
J.Med.Chem., 62, 2019
|
|
