5AXQ
 
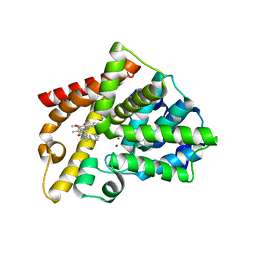 | | Crystal structure of the catalytic domain of PDE10A complexed with highly potent and brain-penetrant PDE10A Inhibitor with 2-oxindole scaffold | | Descriptor: | 1-(cyclopropylmethyl)-4-fluoranyl-5-[5-methoxy-4-oxidanylidene-3-(2-phenylpyrazol-3-yl)pyridazin-1-yl]-3,3-dimethyl-indol-2-one, 3-[3-fluoranyl-4-[5-methoxy-4-oxidanylidene-3-(2-phenylpyrazol-3-yl)pyridazin-1-yl]phenyl]-1,3-oxazolidin-2-one, MAGNESIUM ION, ... | | Authors: | Oki, H, Zama, Y. | | Deposit date: | 2015-07-31 | | Release date: | 2015-11-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Design and synthesis of a novel 2-oxindole scaffold as a highly potent and brain-penetrant phosphodiesterase 10A inhibitor
Bioorg.Med.Chem., 23, 2015
|
|
7TLN
 
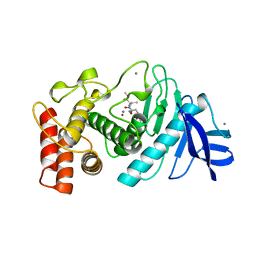 | | STRUCTURAL ANALYSIS OF THE INHIBITION OF THERMOLYSIN BY AN ACTIVE-SITE-DIRECTED IRREVERSIBLE INHIBITOR | | Descriptor: | 2-(ACETYL-HYDROXY-AMINO)-4-METHYL-PENTANOIC ACID METHYL ESTER, CALCIUM ION, THERMOLYSIN, ... | | Authors: | Matthews, B.W, Holmes, M.A, Tronrud, D.E. | | Deposit date: | 1983-01-27 | | Release date: | 1983-03-09 | | Last modified: | 2022-11-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural analysis of the inhibition of thermolysin by an active-site-directed irreversible inhibitor.
Biochemistry, 22, 1983
|
|
5AXP
 
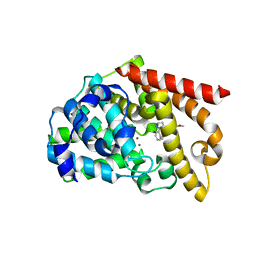 | | Crystal structure of the catalytic domain of PDE10A complexed with 1-(2-fluoro-4-(2-oxo-1,3-oxazolidin-3-yl)phenyl)-5-methoxy-3-(1-phenyl-1H-pyrazol-5-yl)pyridazin-4(1H)-one | | Descriptor: | 3-[3-fluoranyl-4-[5-methoxy-4-oxidanylidene-3-(2-phenylpyrazol-3-yl)pyridazin-1-yl]phenyl]-1,3-oxazolidin-2-one, MAGNESIUM ION, ZINC ION, ... | | Authors: | Oki, H, Zama, Y. | | Deposit date: | 2015-07-31 | | Release date: | 2015-11-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design and synthesis of a novel 2-oxindole scaffold as a highly potent and brain-penetrant phosphodiesterase 10A inhibitor
Bioorg.Med.Chem., 23, 2015
|
|
1ETL
 
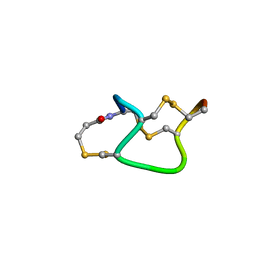 | |
1ETM
 
 | |
8TLN
 
 | |
6IIW
 
 | | Crystal structure of human UHRF1 PHD finger in complex with PAF15 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, E3 ubiquitin-protein ligase UHRF1, PCNA-associated factor, ... | | Authors: | Arita, K, Kori, S. | | Deposit date: | 2018-10-07 | | Release date: | 2019-10-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.699 Å) | | Cite: | Two distinct modes of DNMT1 recruitment ensure stable maintenance DNA methylation.
Nat Commun, 11, 2020
|
|
1BHI
 
 | | STRUCTURE OF TRANSACTIVATION DOMAIN OF CRE-BP1/ATF-2, NMR, 20 STRUCTURES | | Descriptor: | CRE-BP1 | | Authors: | Nagadoi, A, Nakazawa, K, Uda, H, Maekawa, T, Ishii, S, Nishimura, Y. | | Deposit date: | 1998-06-09 | | Release date: | 1999-06-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the transactivation domain of ATF-2 comprising a zinc finger-like subdomain and a flexible subdomain.
J.Mol.Biol., 287, 1999
|
|
5X7X
 
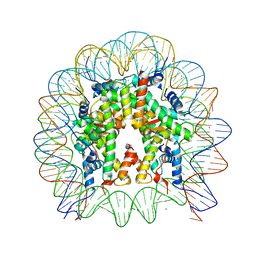 | | The crystal structure of the nucleosome containing H3.3 at 2.18 angstrom resolution | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A type 1-B/E, ... | | Authors: | Arimura, Y, Taguchi, H, Kurumizaka, H. | | Deposit date: | 2017-02-27 | | Release date: | 2017-04-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.184 Å) | | Cite: | Crystal Structure and Characterization of Novel Human Histone H3 Variants, H3.6, H3.7, and H3.8
Biochemistry, 56, 2017
|
|
6KXV
 
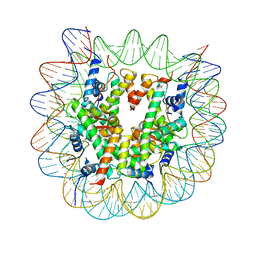 | | Crystal structure of a nucleosome containing Leishmania histone H3 | | Descriptor: | DNA (146-MER), Histone H2A type 1-B/E, Histone H2B type 1-J, ... | | Authors: | Dacher, M, Taguchi, H, Kujirai, T, Kurumizaka, H. | | Deposit date: | 2019-09-13 | | Release date: | 2020-07-22 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.63 Å) | | Cite: | Incorporation and influence of Leishmania histone H3 in chromatin.
Nucleic Acids Res., 47, 2019
|
|
5Y0C
 
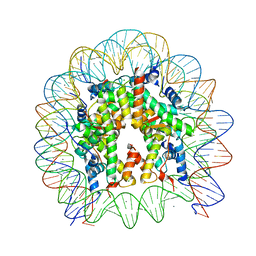 | | Crystal Structure of the human nucleosome at 2.09 angstrom resolution | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A type 1-B/E, ... | | Authors: | Kurumizaka, H, Arimura, Y, Fujita, R, Noda, M. | | Deposit date: | 2017-07-16 | | Release date: | 2018-07-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.087 Å) | | Cite: | Cancer-associated mutations of histones H2B, H3.1 and H2A.Z.1 affect the structure and stability of the nucleosome.
Nucleic Acids Res., 46, 2018
|
|
5Y0D
 
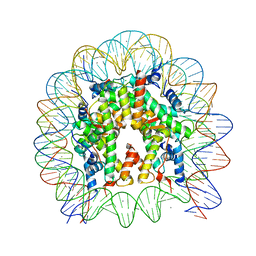 | | Crystal Structure of the human nucleosome containing the H2B E76K mutant | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A type 1-B/E, ... | | Authors: | Kurumizaka, H, Arimura, Y, Fujita, R, Noda, M. | | Deposit date: | 2017-07-16 | | Release date: | 2018-07-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Cancer-associated mutations of histones H2B, H3.1 and H2A.Z.1 affect the structure and stability of the nucleosome.
Nucleic Acids Res., 46, 2018
|
|
2DZV
 
 | |
2DZT
 
 | |
4ZK8
 
 | | Copper-containing nitrite reductase from thermophilic bacterium Geobacillus thermodenitrificans (Re-refined) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Fukuda, Y, Inoue, T. | | Deposit date: | 2015-04-30 | | Release date: | 2015-05-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Structural insights into the function of a thermostable copper-containing nitrite reductase
J.Biochem., 155, 2014
|
|
2DZU
 
 | |
5Z30
 
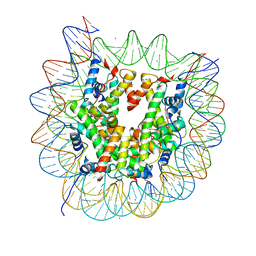 | | The crystal structure of the nucleosome containing a cancer-associated histone H2A.Z R80C mutant | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A.Z, ... | | Authors: | Horikoshi, N, Arimura, Y, Kurumizaka, H. | | Deposit date: | 2018-01-05 | | Release date: | 2018-07-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Cancer-associated mutations of histones H2B, H3.1 and H2A.Z.1 affect the structure and stability of the nucleosome.
Nucleic Acids Res., 46, 2018
|
|
2DZS
 
 | |
2DZW
 
 | |
2E09
 
 | |
2DZP
 
 | |
2DZX
 
 | |
5TMN
 
 | | Slow-and fast-binding inhibitors of thermolysin display different modes of binding. crystallographic analysis of extended phosphonamidate transition-state analogues | | Descriptor: | CALCIUM ION, N-[(S)-({[(benzyloxy)carbonyl]amino}methyl)(hydroxy)phosphoryl]-L-leucyl-L-leucine, THERMOLYSIN, ... | | Authors: | Holden, H.M, Tronrud, D.E, Monzingo, A.F, Weaver, L.H, Matthews, B.W. | | Deposit date: | 1987-06-29 | | Release date: | 1989-01-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Slow- and fast-binding inhibitors of thermolysin display different modes of binding: crystallographic analysis of extended phosphonamidate transition-state analogues.
Biochemistry, 26, 1987
|
|
1F24
 
 | |
1F25
 
 | |
