2IUI
 
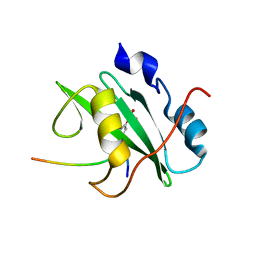 | | Crystal structure of the PI3-kinase p85 N-terminal SH2 domain in complex with PDGFR phosphotyrosyl peptide | | Descriptor: | Phosphatidylinositol 3-kinase regulatory subunit alpha, Platelet-derived growth factor receptor beta | | Authors: | Nolte, R.T, Eck, M.J, Schlessinger, J, Shoelson, S.E, Harrison, S.C. | | Deposit date: | 2006-06-03 | | Release date: | 2006-06-06 | | Last modified: | 2021-04-28 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of the Pi 3-Kinase P85 Amino- Terminal Sh2 Domain and its Phosphopeptide Complexes
Nat.Struct.Biol., 3, 1996
|
|
2JIT
 
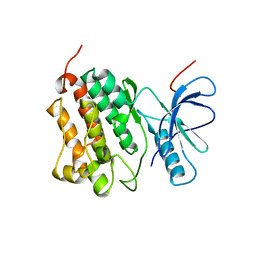 | | Crystal structure of EGFR kinase domain T790M mutation | | Descriptor: | EPIDERMAL GROWTH FACTOR RECEPTOR | | Authors: | Yun, C.-H, Mengwasser, K.E, Toms, A.V, Woo, M.S, Greulich, H, Wong, K.-K, Meyerson, M, Eck, M.J. | | Deposit date: | 2007-07-01 | | Release date: | 2008-01-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The T790M Mutation in Egfr Kinase Causes Drug Resistance by Increasing the Affinity for ATP.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
2JIU
 
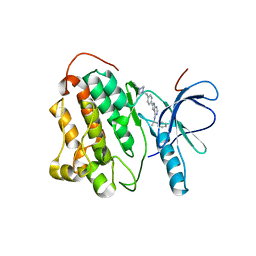 | | Crystal structure of EGFR kinase domain T790M mutation in complex with AEE788 | | Descriptor: | 6-{4-[(4-ETHYLPIPERAZIN-1-YL)METHYL]PHENYL}-N-[(1R)-1-PHENYLETHYL]-7H-PYRROLO[2,3-D]PYRIMIDIN-4-AMINE, EPIDERMAL GROWTH FACTOR RECEPTOR | | Authors: | Yun, C.-H, Mengwasser, K.E, Toms, A.V, Woo, M.S, Greulich, H, Wong, K.-K, Meyerson, M, Eck, M.J. | | Deposit date: | 2007-07-01 | | Release date: | 2008-01-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | The T790M Mutation in Egfr Kinase Causes Drug Resistance by Increasing the Affinity for ATP.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
2JKK
 
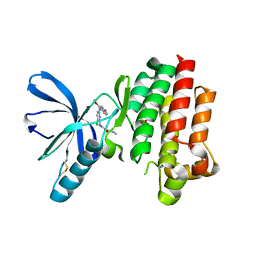 | |
2JKM
 
 | | Focal Adhesion Kinase catalytic domain in complex with bis-anilino pyrimidine inhibitor | | Descriptor: | 2-{[5-CHLORO-2-({(1E,4R)-2-METHOXY-4-[(3R)-3-(METHYLAMINO)PYRROLIDIN-1-YL]CYCLOHEXA-2,5-DIEN-1-YLIDENE}AMINO)PYRIMIDIN-4-YL]AMINO}-N-(1-METHYLETHYL)BENZENESULFONAMIDE, FOCAL ADHESION KINASE 1 | | Authors: | Lietha, D, Eck, M.J. | | Deposit date: | 2008-08-28 | | Release date: | 2008-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Crystal Structures of the Fak Kinase in Complex with Tae226 and Related Bis-Anilino Pyrimidine Inhibitors Reveal a Helical Dfg Conformation.
Plos One, 3, 2008
|
|
2JKQ
 
 | | Focal Adhesion Kinase catalytic domain in complex with bis-anilino pyrimidine inhibitor | | Descriptor: | 4-(1,4'-bipiperidin-1'-yl)-7-({5-chloro-2-[(2-methoxyphenyl)amino]pyrimidin-4-yl}amino)-2-methyl-2,3-dihydro-1H-isoindol-1-one, FOCAL ADHESION KINASE 1 | | Authors: | Lietha, D, Eck, M.J. | | Deposit date: | 2008-08-29 | | Release date: | 2008-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structures of the Fak Kinase in Complex with Tae226 and Related Bis-Anilino Pyrimidine Inhibitors Reveal a Helical Dfg Conformation.
Plos One, 3, 2008
|
|
2IUH
 
 | | Crystal structure of the PI3-kinase p85 N-terminal SH2 domain in complex with c-Kit phosphotyrosyl peptide | | Descriptor: | C-KIT PHOSPHOTYROSYL PEPTIDE, PHOSPHATIDYLINOSITOL 3-KINASE REGULATORY ALPHA SUBUNIT | | Authors: | Nolte, R.T, Eck, M.J, Schlessinger, J, Shoelson, S.E, Harrison, S.C. | | Deposit date: | 2006-06-03 | | Release date: | 2006-06-06 | | Last modified: | 2021-04-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Pi 3-Kinase P85 Amino-Terminal Sh2 Domain and its Phosphopeptide Complexes
Nat.Struct.Biol., 3, 1996
|
|
2JIV
 
 | | Crystal structure of EGFR kinase domain T790M mutation in compex with HKI-272 | | Descriptor: | CHLORIDE ION, EPIDERMAL GROWTH FACTOR RECEPTOR, N-(4-{[3-chloro-4-(pyridin-2-ylmethoxy)phenyl]amino}-3-cyano-7-ethoxyquinolin-6-yl)-4-(dimethylamino)butanamide | | Authors: | Yun, C.-H, Mengwasser, K.E, Toms, A.V, Li, Y, Woo, M.S, Greulich, H, Wong, K.-K, Meyerson, M, Eck, M.J. | | Deposit date: | 2007-07-02 | | Release date: | 2008-01-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | The T790M Mutation in Egfr Kinase Causes Drug Resistance by Increasing the Affinity for ATP.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
2JKO
 
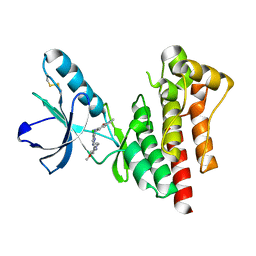 | | Focal Adhesion Kinase catalytic domain in complex with bis-anilino pyrimidine inhibitor | | Descriptor: | 7-{[(2Z,5S)-5-CHLORO-2-{[2-METHOXY-4-(4-METHYLPIPERAZIN-1-YL)PHENYL]IMINO}-2,5-DIHYDROPYRIMIDIN-4-YL]AMINO}-2-METHYL-2,3-DIHYDRO-1H-ISOINDOL-1-ONE, FOCAL ADHESION KINASE 1 | | Authors: | Lietha, D, Eck, M.J. | | Deposit date: | 2008-08-28 | | Release date: | 2008-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal Structures of the Fak Kinase in Complex with Tae226 and Related Bis-Anilino Pyrimidine Inhibitors Reveal a Helical Dfg Conformation.
Plos One, 3, 2008
|
|
7JXW
 
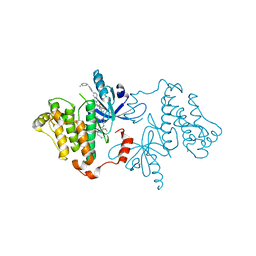 | | EGFR kinase (T790M/V948R) in complex with osimertinib and JBJ-09-063 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-{6-[4-(1-methylpiperidin-4-yl)phenyl]-1-oxo-1,3-dihydro-2H-isoindol-2-yl}-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, N-(2-{[2-(dimethylamino)ethyl](methyl)amino}-4-methoxy-5-{[4-(1-methyl-1H-indol-3-yl)pyrimidin-2-yl]amino}phenyl)prop-2-enamide | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-28 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7JXI
 
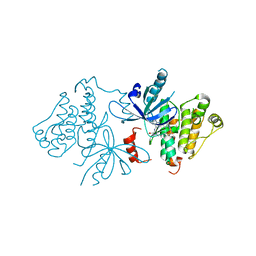 | | EGFR kinase (T790M/V948R) in complex with PF-06747775 | | Descriptor: | Epidermal growth factor receptor, N-[(3R,4R)-4-fluoro-1-{6-[(3-methoxy-1-methyl-1H-pyrazol-4-yl)amino]-9-methyl-9H-purin-2-yl}pyrrolidin-3-yl]propanamide | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-27 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7JXM
 
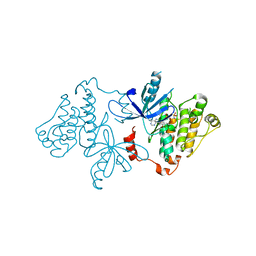 | | EGFR kinase (T790M/V948R) in complex with osimertinib and EAI045 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-(1-oxo-1,3-dihydro-2H-isoindol-2-yl)-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-27 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.192 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7JXP
 
 | | EGFR kinase (T790M/V948R) in complex with osimertinib and JBJ-04-125-02 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-{1-oxo-6-[4-(piperazin-1-yl)phenyl]-1,3-dihydro-2H-isoindol-2-yl}-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-27 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7JXL
 
 | | EGFR kinase (T790M/V948R) in complex with AZ5104 | | Descriptor: | Epidermal growth factor receptor, N-(2-{[2-(dimethylamino)ethyl](methyl)amino}-5-{[4-(1H-indol-3-yl)pyrimidin-2-yl]amino}-4-methoxyphenyl)propanamide | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-27 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7JXK
 
 | | EGFR kinase (T790M/V948R) in complex with PF-06747775 and JBJ-04-125-02 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-{1-oxo-6-[4-(piperazin-1-yl)phenyl]-1,3-dihydro-2H-isoindol-2-yl}-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-08-27 | | Release date: | 2021-09-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7K1I
 
 | | EGFR kinase (L858R/V948R) in complex with allosteric inhibitor JBJ-09-063 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-{6-[4-(1-methylpiperidin-4-yl)phenyl]-1-oxo-1,3-dihydro-2H-isoindol-2-yl}-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-09-07 | | Release date: | 2021-09-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.202 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7K1H
 
 | | EGFR L858R/V948R in complex with osimertinib and allosteric inhibitor JBJ-09-063 | | Descriptor: | (2R)-2-(5-fluoro-2-hydroxyphenyl)-2-{6-[4-(1-methylpiperidin-4-yl)phenyl]-1-oxo-1,3-dihydro-2H-isoindol-2-yl}-N-(1,3-thiazol-2-yl)acetamide, Epidermal growth factor receptor, N-(2-{[2-(dimethylamino)ethyl](methyl)amino}-4-methoxy-5-{[4-(1-methyl-1H-indol-3-yl)pyrimidin-2-yl]amino}phenyl)prop-2-enamide | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-09-07 | | Release date: | 2021-09-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors
Nat Commun, 13, 2022
|
|
7KXZ
 
 | | Active conformation of EGFR kinase in complex with BI-4020 | | Descriptor: | (20R)-10,15,20-trimethyl-2-[(4-methylpiperazin-1-yl)methyl]-18,19,20,21-tetrahydro-15H,17H-12,8-(metheno)pyrazolo[3',4':2,3][1,5,10,12]oxatriazacycloheptadecino[12,11-a]benzimidazol-7(6H)-one, CHLORIDE ION, Epidermal growth factor receptor | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-12-06 | | Release date: | 2022-01-19 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Analysis of the Macrocyclic Inhibitor BI-4020 Binding to EGFR Kinase.
Chemmedchem, 2024
|
|
7KY0
 
 | | Inactive conformation of EGFR (T790M/V948R) kinase in complex with BI-4020 | | Descriptor: | (20R)-10,15,20-trimethyl-2-[(4-methylpiperazin-1-yl)methyl]-18,19,20,21-tetrahydro-15H,17H-12,8-(metheno)pyrazolo[3',4':2,3][1,5,10,12]oxatriazacycloheptadecino[12,11-a]benzimidazol-7(6H)-one, Epidermal growth factor receptor | | Authors: | Beyett, T.S, Eck, M.J. | | Deposit date: | 2020-12-06 | | Release date: | 2022-01-19 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Analysis of the Macrocyclic Inhibitor BI-4020 Binding to EGFR Kinase.
Chemmedchem, 2024
|
|
1LCK
 
 | |
1LCJ
 
 | |
4F9G
 
 | | Crystal structure of STING complex with Cyclic di-GMP. | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Transmembrane protein 173 | | Authors: | Kabaleeswaran, V, Wu, H. | | Deposit date: | 2012-05-18 | | Release date: | 2012-07-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Cyclic di-GMP Sensing via the Innate Immune Signaling Protein STING.
Mol.Cell, 46, 2012
|
|
4F9E
 
 | |
3R7G
 
 | | Crystal structure of Spire KIND domain in complex with the tail of FMN2 | | Descriptor: | Formin-2, Protein spire homolog 1 | | Authors: | Kreutz, B, Vizcarra, C.L, Rodal, A.A, Toms, A.V, Lu, J, Quinlan, M.E, Eck, M.J. | | Deposit date: | 2011-03-22 | | Release date: | 2011-07-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the Spire KIND domain and insights into its interaction with Fmn-family formins
Proc.Natl.Acad.Sci.USA, 2011
|
|
3RBW
 
 | | Crystal structure of Spire KIND domain | | Descriptor: | Protein spire homolog 1 | | Authors: | Vizcarra, C.L, Kreutz, B, Rodal, A.A, Toms, A.V, Lu, J, Zheng, W, Quinlan, M.E, Eck, M.J. | | Deposit date: | 2011-03-30 | | Release date: | 2011-07-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure of the Spire KIND domain and insights into its interaction with Fmn-family formins
Proc.Natl.Acad.Sci.USA, 2011
|
|
