5YB4
 
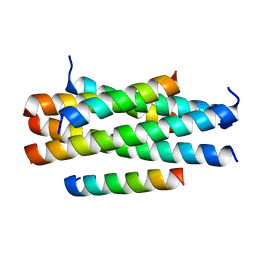 | | Crystal structure of HP23LN36KR | | 分子名称: | HP23L, N36KR | | 著者 | Zhang, X, Wang, X, He, Y. | | 登録日 | 2017-09-03 | | 公開日 | 2018-02-28 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Structural Insights into the Mechanisms of Action of Short-Peptide HIV-1 Fusion Inhibitors Targeting the Gp41 Pocket
Front Cell Infect Microbiol, 8, 2018
|
|
3IYL
 
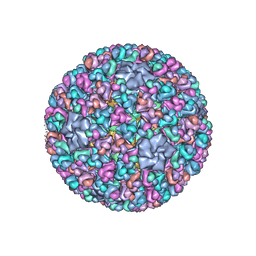 | | Atomic CryoEM Structure of a Nonenveloped Virus Suggests How Membrane Penetration Protein is Primed for Cell Entry | | 分子名称: | Core protein VP6, MYRISTIC ACID, Outer capsid VP4, ... | | 著者 | Zhang, X, Jin, L, Fang, Q, Hui, W, Zhou, Z.H. | | 登録日 | 2010-02-02 | | 公開日 | 2010-05-12 | | 最終更新日 | 2024-11-20 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | 3.3 A cryo-EM structure of a nonenveloped virus reveals a priming mechanism for cell entry.
Cell(Cambridge,Mass.), 141, 2010
|
|
3J4U
 
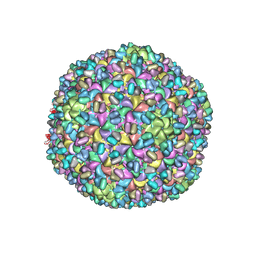 | | A new topology of the HK97-like fold revealed in Bordetella bacteriophage: non-covalent chainmail secured by jellyrolls | | 分子名称: | cementing protein, major capsid protein | | 著者 | Zhang, X, Guo, H, Jin, L, Czornyj, E, Hodes, A, Hui, W.H, Nieh, A.W, Miller, J.F, Zhou, Z.H. | | 登録日 | 2013-10-09 | | 公開日 | 2013-12-25 | | 最終更新日 | 2024-02-21 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5 A resolution.
Elife, 2, 2013
|
|
7EVN
 
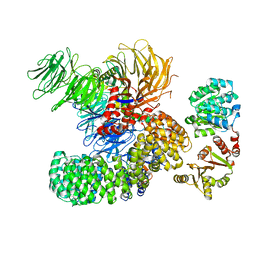 | | The cryo-EM structure of the DDX42-SF3b complex | | 分子名称: | ATP-dependent RNA helicase DDX42, PHD finger-like domain-containing protein 5A, Splicing factor 3B subunit 1, ... | | 著者 | Zhang, X, Zhan, X, Shi, Y. | | 登録日 | 2021-05-21 | | 公開日 | 2022-08-03 | | 最終更新日 | 2025-06-18 | | 実験手法 | ELECTRON MICROSCOPY (2.6 Å) | | 主引用文献 | Structural insights into branch site proofreading by human spliceosome
Nat.Struct.Mol.Biol., 2024
|
|
7EVO
 
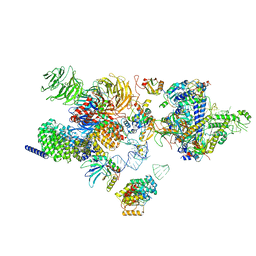 | | The cryo-EM structure of the human 17S U2 snRNP | | 分子名称: | HIV Tat-specific factor 1, PHD finger-like domain-containing protein 5A, RNA helicase, ... | | 著者 | Zhang, X, Zhan, X, Shi, Y. | | 登録日 | 2021-05-21 | | 公開日 | 2022-08-03 | | 最終更新日 | 2024-05-22 | | 実験手法 | ELECTRON MICROSCOPY (2.5 Å) | | 主引用文献 | Structural insights into branch site proofreading by human spliceosome.
Nat.Struct.Mol.Biol., 2024
|
|
3AH5
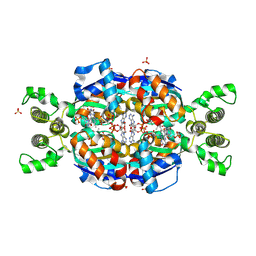 
 | | Crystal Structure of flavin dependent thymidylate synthase ThyX from helicobacter pylori complexed with FAD and dUMP | | 分子名称: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, FLAVIN-ADENINE DINUCLEOTIDE, SULFATE ION, ... | | 著者 | Zhang, X, Zhang, J, Hu, Y, Zou, Q, Wang, D. | | 登録日 | 2010-04-14 | | 公開日 | 2011-04-20 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Crystal structure and functional analysis of a flavin dependent thymidylate synthase from helicobacter pylori
To be Published
|
|
5Z57
 
 | | Cryo-EM structure of the human activated spliceosome (late Bact) at 6.5 angstrom | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, ALANINE, BUD13 homolog, ... | | 著者 | Zhang, X, Yan, C, Zhan, X, Li, L, Lei, J, Shi, Y. | | 登録日 | 2018-01-17 | | 公開日 | 2018-09-19 | | 最終更新日 | 2024-10-30 | | 実験手法 | ELECTRON MICROSCOPY (6.5 Å) | | 主引用文献 | Structure of the human activated spliceosome in three conformational states.
Cell Res., 28, 2018
|
|
5Z56
 
 | | cryo-EM structure of a human activated spliceosome (mature Bact) at 5.1 angstrom. | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, BUD13 homolog, Cell division cycle 5-like protein, ... | | 著者 | Zhang, X, Yan, C, Zhan, X, Li, L, Lei, J, Shi, Y. | | 登録日 | 2018-01-17 | | 公開日 | 2018-09-19 | | 最終更新日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (5.1 Å) | | 主引用文献 | Structure of the human activated spliceosome in three conformational states.
Cell Res., 28, 2018
|
|
5Z58
 
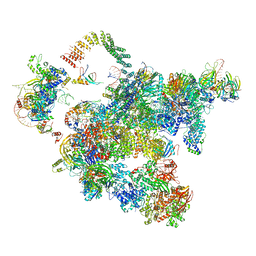 | | Cryo-EM structure of a human activated spliceosome (early Bact) at 4.9 angstrom. | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, BUD13 homolog, Cell division cycle 5-like protein, ... | | 著者 | Zhang, X, Yan, C, Zhan, X, Li, L, Lei, J, Shi, Y. | | 登録日 | 2018-01-17 | | 公開日 | 2018-09-19 | | 最終更新日 | 2024-11-06 | | 実験手法 | ELECTRON MICROSCOPY (4.9 Å) | | 主引用文献 | Structure of the human activated spliceosome in three conformational states.
Cell Res., 28, 2018
|
|
5Z1V
 
 | | Crystal structure of AvrPib | | 分子名称: | AvrPib protein | | 著者 | Zhang, X, He, D, Zhao, Y.X, Taylor, I.A, Peng, Y.L, Yang, J, Liu, J.F. | | 登録日 | 2017-12-28 | | 公開日 | 2018-09-05 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.661 Å) | | 主引用文献 | A positive-charged patch and stabilized hydrophobic core are essential for avirulence function of AvrPib in the rice blast fungus.
Plant J., 96, 2018
|
|
5TEC
 
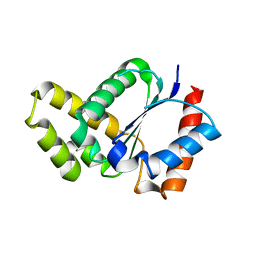 | | Crystal structure of the TIR domain from the Arabidopsis thaliana NLR protein SNC1 | | 分子名称: | Protein SUPPRESSOR OF npr1-1, CONSTITUTIVE 1 | | 著者 | Zhang, X, Bentham, A, Ve, T, Williams, S.J, Kobe, B. | | 登録日 | 2016-09-20 | | 公開日 | 2017-02-01 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Multiple functional self-association interfaces in plant TIR domains.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
9J0X
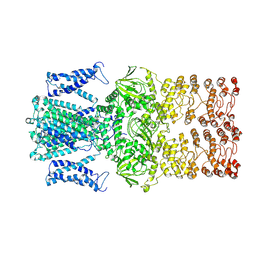 
 | |
9J10
 
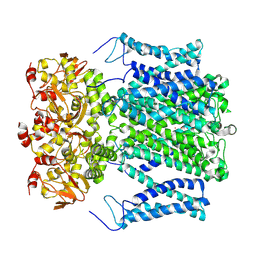 | |
9J0Y
 
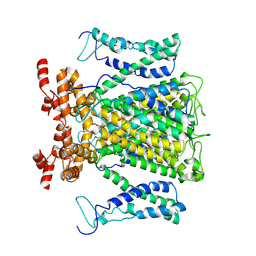 | |
9J0Z
 
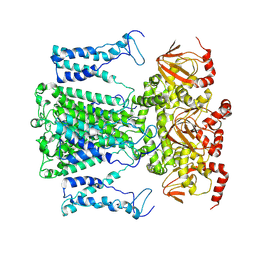 | |
9B5Y
 
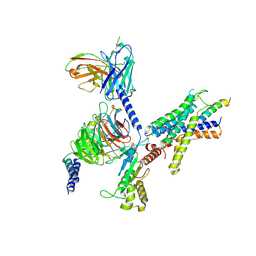 | | Cryo-EM structure of the LAPTH-bound PTH1R in complex with Gq | | 分子名称: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(q), ... | | 著者 | Zhang, X, Liu, H, Zhang, C, Vilardaga, J.-P. | | 登録日 | 2024-03-22 | | 公開日 | 2025-03-19 | | 最終更新日 | 2025-06-04 | | 実験手法 | ELECTRON MICROSCOPY (3.49 Å) | | 主引用文献 | Allosteric mechanism in the distinctive coupling of G q and G s to the parathyroid hormone type 1 receptor.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
5KQA
 
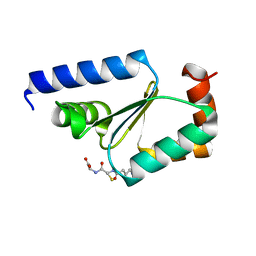 | | Crystal structure of buckwheat glutaredoxin-glutathione complex | | 分子名称: | GLUTATHIONE, Glutaredoxin-glutathione complex | | 著者 | Zhang, X, Wang, W, Zhao, Y, Wang, Z, Wang, H. | | 登録日 | 2016-07-06 | | 公開日 | 2017-07-05 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | Structural insights into the binding of buckwheat glutaredoxin with GSH and regulation of its catalytic activity
J. Inorg. Biochem., 173, 2017
|
|
8T3V
 
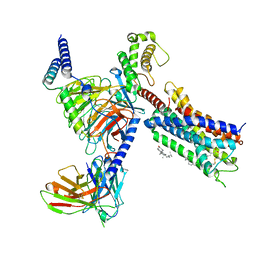 | | Cryo-EM structure of the DHA bound FFA1-Gq complex | | 分子名称: | CHOLESTEROL, DOCOSA-4,7,10,13,16,19-HEXAENOIC ACID, Free fatty acid receptor 1, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2023-06-07 | | 公開日 | 2024-01-24 | | 最終更新日 | 2024-11-06 | | 実験手法 | ELECTRON MICROSCOPY (3.39 Å) | | 主引用文献 | Structural basis for the ligand recognition and signaling of free fatty acid receptors.
Sci Adv, 10, 2024
|
|
8T3O
 
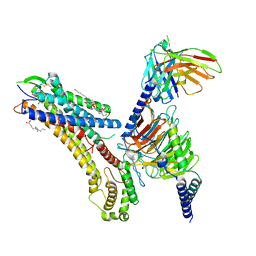 | | Cryo-EM structure of the TUG-891 bound FFA4-Gq complex | | 分子名称: | (2R)-1-{[(R)-hydroxy{[(1R,2R,3R,4R,5S,6R)-2,3,5,6-tetrahydroxy-4-(phosphonooxy)cyclohexyl]oxy}phosphoryl]oxy}-3-(octadecanoyloxy)propan-2-yl (5Z,8Z,11Z,14Z)-icosa-5,8,11,14-tetraenoate, 3-{4-[(4-fluoro-4'-methyl[1,1'-biphenyl]-2-yl)methoxy]phenyl}propanoic acid, Free fatty acid receptor 4, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2023-06-07 | | 公開日 | 2024-01-17 | | 最終更新日 | 2025-05-14 | | 実験手法 | ELECTRON MICROSCOPY (3.06 Å) | | 主引用文献 | Structural basis for the ligand recognition and signaling of free fatty acid receptors.
Sci Adv, 10, 2024
|
|
8T3Q
 
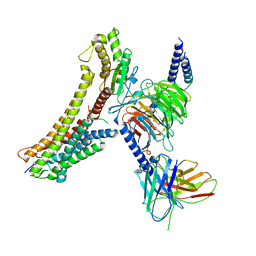 | | Cryo-EM structure of the DHA bound FFA4-Gq complex | | 分子名称: | DOCOSA-4,7,10,13,16,19-HEXAENOIC ACID, Free fatty acid receptor 4, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2023-06-07 | | 公開日 | 2024-01-24 | | 最終更新日 | 2025-05-28 | | 実験手法 | ELECTRON MICROSCOPY (3.14 Å) | | 主引用文献 | Structural basis for the ligand recognition and signaling of free fatty acid receptors.
Sci Adv, 10, 2024
|
|
8T3S
 
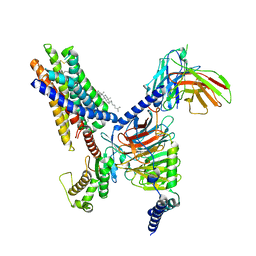 | | Cryo-EM structure of the Butyrate bound FFA2-Gq complex | | 分子名称: | CHOLESTEROL, Free fatty acid receptor 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2023-06-07 | | 公開日 | 2024-01-24 | | 最終更新日 | 2025-05-28 | | 実験手法 | ELECTRON MICROSCOPY (3.07 Å) | | 主引用文献 | Structural basis for the ligand recognition and signaling of free fatty acid receptors.
Sci Adv, 10, 2024
|
|
8SG1
 
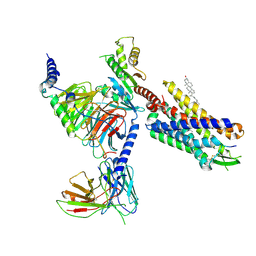 | | Cryo-EM structure of CMKLR1 signaling complex | | 分子名称: | CHOLESTEROL, Chemerin 9, Chemerin-like receptor 1, ... | | 著者 | Zhang, X, Zhang, C. | | 登録日 | 2023-04-11 | | 公開日 | 2023-11-01 | | 最終更新日 | 2025-05-28 | | 実験手法 | ELECTRON MICROSCOPY (2.94 Å) | | 主引用文献 | Structural basis of G protein-Coupled receptor CMKLR1 activation and signaling induced by a chemerin-derived agonist.
Plos Biol., 21, 2023
|
|
9CLW
 
 | | Cryo-EM structure of Gq-coupled FFA2 in complex with TUG-1375 and 4-CMTB | | 分子名称: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, (2~{S})-2-(4-chlorophenyl)-3-methyl-~{N}-(1,3-thiazol-2-yl)butanamide, CHOLESTEROL, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2024-07-12 | | 公開日 | 2025-06-25 | | 最終更新日 | 2025-07-02 | | 実験手法 | ELECTRON MICROSCOPY (3.19 Å) | | 主引用文献 | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
9CM7
 
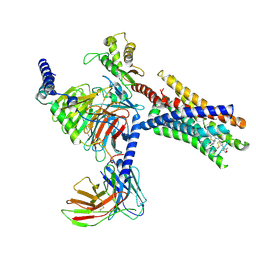 | | Cryo-EM structure of Gi-coupled FFA2 in complex with TUG-1375 and AZ-1729 | | 分子名称: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, Free fatty acid receptor 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2024-07-12 | | 公開日 | 2025-06-25 | | 最終更新日 | 2025-07-02 | | 実験手法 | ELECTRON MICROSCOPY (3.29 Å) | | 主引用文献 | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
9CM3
 
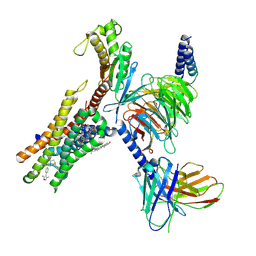 | | Cryo-EM structure of Gq-coupled FFA2 in complex with TUG-1375 and compound 187 | | 分子名称: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, 4-[(2R,6S)-2,6-dimethylmorpholin-4-yl]-7-(2-fluorobenzene-1-sulfonyl)-2-methyl-5H-pyrrolo[3,2-d]pyrimidin-6-amine, CHOLESTEROL, ... | | 著者 | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | 登録日 | 2024-07-12 | | 公開日 | 2025-06-25 | | 最終更新日 | 2025-07-09 | | 実験手法 | ELECTRON MICROSCOPY (3.06 Å) | | 主引用文献 | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
