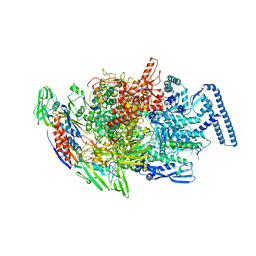
6TYE
 
 | |
6PQ7
 
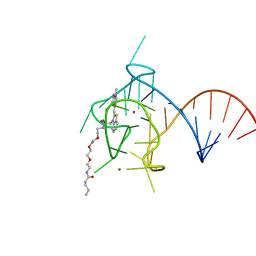 | |
6UTN
 
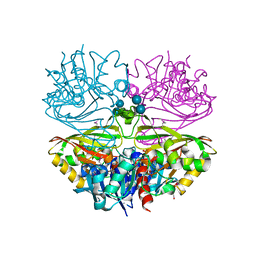 | | Native E. coli Glyceraldehyde 3-phosphate dehydrogenase | | 分子名称: | ACETATE ION, Glyceraldehyde-3-phosphate dehydrogenase, PHOSPHATE ION, ... | | 著者 | Rodriguez-Hernandez, A, Romo-Arevalo, E, Rodriguez-Romero, A. | | 登録日 | 2019-10-29 | | 公開日 | 2019-12-11 | | 最終更新日 | 2024-11-20 | | 実験手法 | X-RAY DIFFRACTION (1.79 Å) | | 主引用文献 | A Novel Substrate-Binding Site in the X-Ray Structure of an Oxidized E. coli Glyceraldehyde 3-Phosphate Dehydrogenase Elucidated by Single-Wavelength Anomalous Dispersion
Crystals, 9, 2019
|
|
6P7M
 
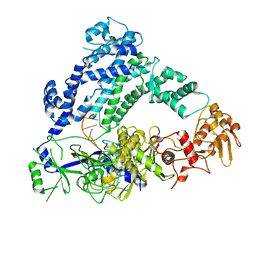 | | Cryo-EM structure of LbCas12a-crRNA: AcrVA4 (1:2 complex) | | 分子名称: | Cas12a, MAGNESIUM ION, anti-CRISPR VA4, ... | | 著者 | Knott, G.J, Liu, J.J, Doudna, J.A. | | 登録日 | 2019-06-06 | | 公開日 | 2019-08-21 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a.
Elife, 8, 2019
|
|
6UTM
 
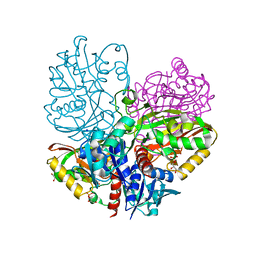 | | Native E. coli Glyceraldehyde 3-phosphate dehydrogenase | | 分子名称: | GLYCEROL, Glyceraldehyde-3-phosphate dehydrogenase, SN-GLYCEROL-3-PHOSPHATE, ... | | 著者 | Rodriguez-Hernandez, A, Romo-Arevalo, E, Rodriguez-Romero, A. | | 登録日 | 2019-10-29 | | 公開日 | 2019-12-11 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (2.14 Å) | | 主引用文献 | A Novel Substrate-Binding Site in the X-Ray Structure of an Oxidized E. coli Glyceraldehyde 3-Phosphate Dehydrogenase Elucidated by Single-Wavelength Anomalous Dispersion
Crystals, 9, 2019
|
|
6P7N
 
 | | Cryo-EM structure of LbCas12a-crRNA: AcrVA4 (2:2 complex) | | 分子名称: | Cas12a, MAGNESIUM ION, anti-CRISPR VA4, ... | | 著者 | Knott, G.J, Liu, J.J, Doudna, J.A. | | 登録日 | 2019-06-06 | | 公開日 | 2019-08-21 | | 最終更新日 | 2023-08-16 | | 実験手法 | ELECTRON MICROSCOPY (4.9 Å) | | 主引用文献 | Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a.
Elife, 8, 2019
|
|
6UTO
 
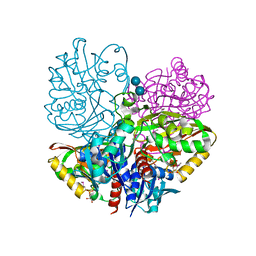 | | Native E. coli Glyceraldehyde 3-phosphate dehydrogenase | | 分子名称: | ACETATE ION, Glyceraldehyde-3-phosphate dehydrogenase, SN-GLYCEROL-3-PHOSPHATE, ... | | 著者 | Rodriguez-Hernandez, A, Romo-Arevalo, E, Rodriguez-Romero, A. | | 登録日 | 2019-10-29 | | 公開日 | 2019-12-11 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (1.64 Å) | | 主引用文献 | A Novel Substrate-Binding Site in the X-Ray Structure of an Oxidized E. coli Glyceraldehyde 3-Phosphate Dehydrogenase Elucidated by Single-Wavelength Anomalous Dispersion
Crystals, 9, 2019
|
|
6UP0
 
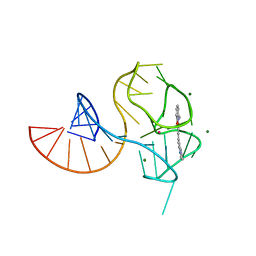 | |
7ZFM
 
 | | Engineered Protein Targeting the Zika Viral Envelope Fusion Loop | | 分子名称: | 1,2-ETHANEDIOL, ACETIC ACID, HEXAETHYLENE GLYCOL, ... | | 著者 | Athayde, D, Archer, M, Viana, I.F.T, Adan, W.C.S, Xavier, L.S.S, Lins, R.D. | | 登録日 | 2022-04-01 | | 公開日 | 2022-08-17 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.711 Å) | | 主引用文献 | In Vitro Neutralisation of Zika Virus by an Engineered Protein Targeting the Viral Envelope Fusion Loop
SSRN, 2022
|
|
8TJG
 
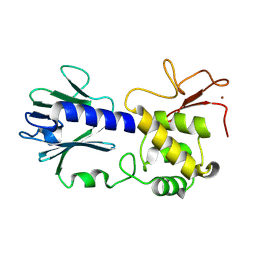 | |
6PW8
 
 | |
6VBK
 
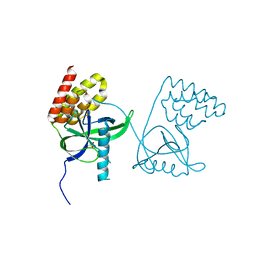 | | Crystal structure of N-terminal domain of Mycobacterium tuberculosis complex Lon protease | | 分子名称: | GLYCEROL, Lon211 | | 著者 | Bi, F.K, Chen, C, Chen, X.Y, Guo, C.Y, Lin, D.H. | | 登録日 | 2019-12-19 | | 公開日 | 2020-12-23 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Crystal structure of the N domain of Lon protease from Mycobacterium avium complex.
Protein Sci., 28, 2019
|
|
8A67
 
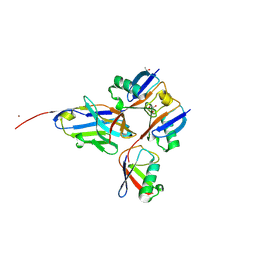 | |
7PPM
 
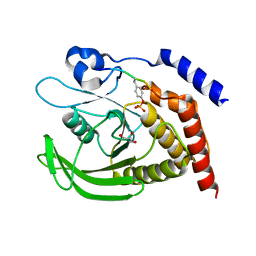 | | SHP2 catalytic domain in complex with IRS1 (889-901) phosphopeptide (pSer-892, pTyr-896) | | 分子名称: | GLYCEROL, Insulin receptor substrate 1, Tyrosine-protein phosphatase non-receptor type 11,Tyrosine-protein phosphatase non-receptor type 11 | | 著者 | Sok, P, Zeke, A, Remenyi, A. | | 登録日 | 2021-09-14 | | 公開日 | 2022-09-07 | | 最終更新日 | 2024-11-20 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Structural insights into the pSer/pThr dependent regulation of the SHP2 tyrosine phosphatase in insulin and CD28 signaling.
Nat Commun, 13, 2022
|
|
7PPN
 
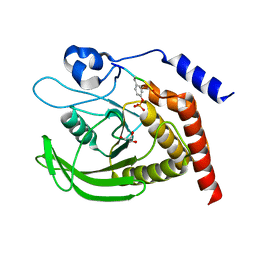 | | SHP2 catalytic domain in complex with CD28 (183-198) phosphopeptide (pTyr-191, p-Thr-195) | | 分子名称: | GLYCEROL, T-cell-specific surface glycoprotein CD28, Tyrosine-protein phosphatase non-receptor type 11,Tyrosine-protein phosphatase non-receptor type 11 | | 著者 | Sok, P, Zeke, A, Remenyi, A. | | 登録日 | 2021-09-14 | | 公開日 | 2022-09-07 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structural insights into the pSer/pThr dependent regulation of the SHP2 tyrosine phosphatase in insulin and CD28 signaling.
Nat Commun, 13, 2022
|
|
7PPL
 
 | | SHP2 catalytic domain in complex with IRS1 (625-639) phosphopeptide (pTyr-632, pSer-636) | | 分子名称: | ETHANOL, GLYCEROL, Insulin receptor substrate 1, ... | | 著者 | Sok, P, Zeke, A, Remenyi, A. | | 登録日 | 2021-09-14 | | 公開日 | 2022-09-07 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.53 Å) | | 主引用文献 | Structural insights into the pSer/pThr dependent regulation of the SHP2 tyrosine phosphatase in insulin and CD28 signaling.
Nat Commun, 13, 2022
|
|
7PUX
 
 | |
9GBF
 
 | | X-RAY structure of PHDvC5HCH tandem domain of NSD2 | | 分子名称: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, Histone-lysine N-methyltransferase NSD2, ... | | 著者 | Musco, G, Cocomazzi, P, Berardi, A, Knapp, S, Kramer, A. | | 登録日 | 2024-07-31 | | 公開日 | 2024-12-18 | | 最終更新日 | 2025-01-22 | | 実験手法 | X-RAY DIFFRACTION (1.76 Å) | | 主引用文献 | The C-terminal PHDVC5HCH tandem domain of NSD2 is a combinatorial reader of unmodified H3K4 and tri-methylated H3K27 that regulates transcription of cell adhesion genes in multiple myeloma.
Nucleic Acids Res., 53, 2025
|
|
7PV9
 
 | |
9NRC
 | | TMPRSS6 in complex with REGN7999 Fab and REGN8023 Fab | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | 著者 | Saotome, K, Franklin, M.C. | | 登録日 | 2025-03-14 | | 公開日 | 2025-07-09 | | 実験手法 | ELECTRON MICROSCOPY (3.29 Å) | | 主引用文献 | A TMPRSS6-inhibiting mAb improves disease in a beta-thalassemia mouse model and reduces iron in healthy humans.
JCI Insight, 10, 2025
|
|
7PTH
 
 | |
7R7M
 
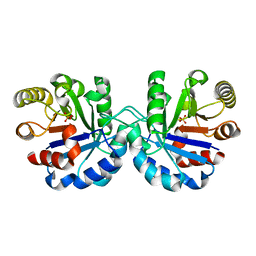 | |
2H63
 
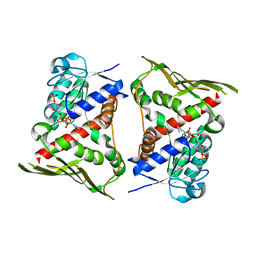 | | Crystal Structure of Human Biliverdin Reductase A | | 分子名称: | Biliverdin reductase A, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | 著者 | Kavanagh, K, Elkins, J, Ugochukwu, E, Guo, K, Pilka, E, Lukacik, P, Smee, C, Papagrigoriou, E, Bunkoczi, G, Sundstrom, M, Arrowsmith, C, Weigelt, J, Edwards, A, von Delft, F, Oppermann, U, Structural Genomics Consortium (SGC) | | 登録日 | 2006-05-30 | | 公開日 | 2006-07-04 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Crystal Structure of Human Biliverdin Reductase A
To be Published
|
|
2I3Y
 
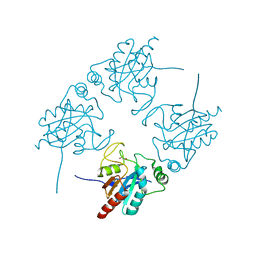 | | Crystal structure of human glutathione peroxidase 5 | | 分子名称: | 1,2-ETHANEDIOL, Epididymal secretory glutathione peroxidase | | 著者 | Kavanagh, K.L, Johansson, C, Rojkova, A, Umeano, C, Bunkoczi, G, Gileadi, O, von Delft, F, Weigelt, J, Arrowsmith, C, Sundstrom, M, Edwards, A, Oppermann, U, Structural Genomics Consortium (SGC) | | 登録日 | 2006-08-21 | | 公開日 | 2006-09-12 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Crystal structure of human glutathione peroxidase 5
To be published
|
|
9H3I
 | |
