7ZBN
 
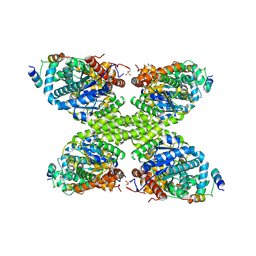 | |
3K2B
 
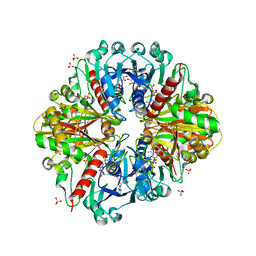 | | Crystal structure of photosynthetic A4 isoform glyceraldehyde-3-phosphate dehydrogenase complexed with NAD, from Arabidopsis thaliana | | Descriptor: | Glyceraldehyde-3-phosphate dehydrogenase A, chloroplastic, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Fermani, S, Falini, G, Thumiger, A, Sparla, F, Marri, L, Trost, P. | | Deposit date: | 2009-09-29 | | Release date: | 2010-06-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of photosynthetic glyceraldehyde-3-phosphate dehydrogenase (isoform A4) from Arabidopsis thaliana in complex with NAD
Acta Crystallogr.,Sect.F, 66, 2010
|
|
3QV1
 
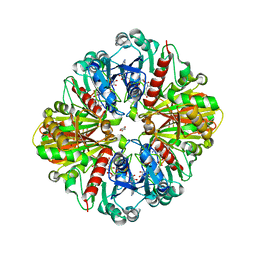 | | Crystal structure of the binary complex of photosyntetic A4 glyceraldehyde 3-phosphate dehydrogenase (GAPDH) with cp12-2, both from Arabidopsis thaliana. | | Descriptor: | 1,2-ETHANEDIOL, CP12 protein, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Thumiger, A, Fermani, S, Falini, G, Marri, L, Sparla, F, Trost, P. | | Deposit date: | 2011-02-25 | | Release date: | 2012-02-29 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational Selection and Folding-upon-binding of Intrinsically Disordered Protein CP12 Regulate Photosynthetic Enzymes Assembly.
J.Biol.Chem., 287, 2012
|
|
3RVD
 
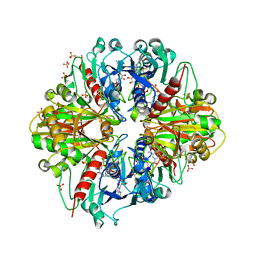 | | Crystal structure of the binary complex, obtained by soaking, of photosyntetic a4 glyceraldehyde 3-phosphate dehydrogenase (gapdh) with cp12-2, both from arabidopsis thaliana. | | Descriptor: | Glyceraldehyde-3-phosphate dehydrogenase A, chloroplastic, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Fermani, S, Thumiger, A, Falini, G, Marri, L, Sparla, F, Trost, P. | | Deposit date: | 2011-05-06 | | Release date: | 2012-04-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Conformational Selection and Folding-upon-binding of Intrinsically Disordered Protein CP12 Regulate Photosynthetic Enzymes Assembly.
J.Biol.Chem., 287, 2012
|
|
2LJ9
 
 | |
4HLN
 
 | | Structure of barley starch synthase I in complex with maltooligosaccharide | | Descriptor: | Starch synthase I, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Cuesta-Seijo, J.A, Nielsen, M.M, Marri, L, Tanaka, H, Palcic, M.M. | | Deposit date: | 2012-10-17 | | Release date: | 2013-06-05 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of starch synthase I from barley: insight into regulatory mechanisms of starch synthase activity
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4R06
 
 | | Crystal Structure of SR2067 bound to PPARgamma | | Descriptor: | 1-(naphthalen-1-ylsulfonyl)-N-[(1S)-1-phenylpropyl]-1H-indole-5-carboxamide, Peroxisome proliferator-activated receptor gamma, SULFATE ION | | Authors: | Marrewijk, L, Kamenecka, T, Griffin, P.R, Bruning, J.B. | | Deposit date: | 2014-07-30 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | SR2067 Reveals a Unique Kinetic and Structural Signature for PPAR gamma Partial Agonism.
Acs Chem.Biol., 11, 2016
|
|
7A54
 
 | |
7AVI
 
 | | Crystal structure of SOS1 in complex with compound 2 | | Descriptor: | 3-propan-2-yl-~{N}-[(1~{R})-1-(3-sulfamoylphenyl)ethyl]-[1,2]oxazolo[5,4-b]pyridine-5-carboxamide, Son of sevenless homolog 1 | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-05 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7AVT
 
 | | Crystal structure of SOS1 in complex with compound 7 | | Descriptor: | IMIDAZOLE, Son of sevenless homolog 1, ~{N}-[(1~{R})-1-(3-aminophenyl)ethyl]-6,7-dimethoxy-2-methyl-quinazolin-4-amine | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-06 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7AVS
 
 | | Crystal structure of SOS1 in complex with compound 6 | | Descriptor: | 6,7-dimethoxy-2-methyl-~{N}-[(1~{R})-1-[3-(trifluoromethyl)phenyl]ethyl]quinazolin-4-amine, IMIDAZOLE, Son of sevenless homolog 1 | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-06 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7AVV
 
 | | Crystal structure of SOS1 in complex with compound 9 | | Descriptor: | IMIDAZOLE, Son of sevenless homolog 1, ~{N}-[(1~{R})-1-[3-azanyl-5-(trifluoromethyl)phenyl]ethyl]-2-methyl-quinazolin-4-amine | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-06 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7AVU
 
 | | Crystal structure of SOS1 in complex with compound 8 | | Descriptor: | IMIDAZOLE, Son of sevenless homolog 1, ~{N}-[(1~{R})-1-[3-azanyl-5-(trifluoromethyl)phenyl]ethyl]-6,7-dimethoxy-2-methyl-quinazolin-4-amine | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-06 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7AVL
 
 | | Crystal structure of SOS1 in complex with compound 4 | | Descriptor: | 6,7-dimethoxy-2-methyl-~{N}-[(1~{R})-1-phenylethyl]quinazolin-4-amine, IMIDAZOLE, Son of sevenless homolog 1 | | Authors: | Bader, G, Kessler, D, Wolkerstorfer, B. | | Deposit date: | 2020-11-05 | | Release date: | 2021-03-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.718 Å) | | Cite: | One Atom Makes All the Difference: Getting a Foot in the Door between SOS1 and KRAS.
J.Med.Chem., 64, 2021
|
|
7QUO
 
 | | FimH lectin domain in complex with oligomannose-6 | | Descriptor: | FimH, NICKEL (II) ION, SULFATE ION, ... | | Authors: | Bouckaert, J, Bourenkov, G.P. | | Deposit date: | 2022-01-18 | | Release date: | 2023-02-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural insights into a cooperative switch between one and two FimH bacterial adhesins binding pauci- and high-mannose type N-glycan receptors
J.Biol.Chem., 299, 2023
|
|
7BHD
 
 | | FimH in complex with alpha1,6 core-fucosylated oligomannose-3, crystallized in the trigonal space group | | Descriptor: | NICKEL (II) ION, SULFATE ION, Type 1 fimbrin D-mannose specific adhesin, ... | | Authors: | Bridot, C, Bouckaert, J, Krammer, E.-M. | | Deposit date: | 2021-01-11 | | Release date: | 2022-07-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural insights into a cooperative switch between one and two FimH bacterial adhesins binding pauci- and high-mannose type N-glycan receptors.
J.Biol.Chem., 299, 2023
|
|
6WXY
 
 | | crystal structure of cA6-bound Card1 | | Descriptor: | Card1, cA6 | | Authors: | Rostol, J, Xie, W, Patel, D.J, Marraffini, L. | | Deposit date: | 2020-05-12 | | Release date: | 2020-12-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Card1 nuclease provides defence during type III CRISPR immunity.
Nature, 590, 2021
|
|
6WXW
 
 | |
6WXX
 
 | | crystal structure of cA4-activated Card1 | | Descriptor: | Card1, MANGANESE (II) ION, cA4 | | Authors: | Rostol, J, Xie, W, Patel, D.J, Marraffini, L. | | Deposit date: | 2020-05-12 | | Release date: | 2020-12-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The Card1 nuclease provides defence during type III CRISPR immunity.
Nature, 590, 2021
|
|
6XL1
 
 | | crystal structure of cA4-activated Card1(D294N) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Card1, MANGANESE (II) ION, ... | | Authors: | Rostol, J, Xie, W, Patel, D.J, Marraffini, L. | | Deposit date: | 2020-06-27 | | Release date: | 2020-12-30 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Card1 nuclease provides defence during type III CRISPR immunity.
Nature, 590, 2021
|
|
7A5X
 
 | | Two copies of the catalytic domain of NanA sialidase from Streptococcus pneumoniae juxtaposed in the P212121 space group, in complex with DANA derivatized with a PEG linker on the glycerol group. | | Descriptor: | (4~{S},5~{R},6~{R})-5-acetamido-6-[(1~{R},2~{S})-3-[[1-(2-ethoxyethyl)-1,2,3-triazol-4-yl]methylsulfanyl]-1,2-bis(oxidanyl)propyl]-4-oxidanyl-oxane-2-carboxylic acid, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Bridot, C, Bouckaert, J. | | Deposit date: | 2020-08-24 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Polyvalent Transition-State Analogues of Sialyl Substrates Strongly Inhibit Bacterial Sialidases*.
Chemistry, 27, 2021
|
|
8BXY
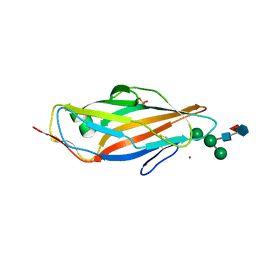 
 | | FimH in complex with alpha1,6 core-fucosylated oligomannose-3, crystallized in the trigonal space group | | Descriptor: | NICKEL (II) ION, SULFATE ION, Type 1 fimbrin D-mannose specific adhesin, ... | | Authors: | Bridot, C, Bouckaert, J, Krammer, E.-M. | | Deposit date: | 2022-12-11 | | Release date: | 2023-04-12 | | Last modified: | 2023-04-26 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural insights into a cooperative switch between one and two FimH bacterial adhesins binding pauci- and high-mannose type N-glycan receptors.
J.Biol.Chem., 299, 2023
|
|
8BY3
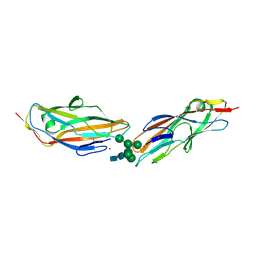 
 | | FimH lectin domain in complex with oligomannose-6 | | Descriptor: | NICKEL (II) ION, SULFATE ION, Type 1 fimbrin D-mannose specific adhesin, ... | | Authors: | Bouckaert, J, Bourenkov, G.P. | | Deposit date: | 2022-12-11 | | Release date: | 2023-04-12 | | Last modified: | 2023-04-26 | | Method: | X-RAY DIFFRACTION (3.186 Å) | | Cite: | Structural insights into a cooperative switch between one and two FimH bacterial adhesins binding pauci- and high-mannose type N-glycan receptors.
J.Biol.Chem., 299, 2023
|
|
8BVD
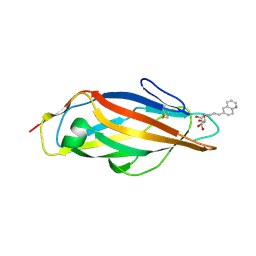 
 | | FimH lectin domain in complex with mannose C-linked to quinoline | | Descriptor: | (2R,3S,4R,5S,6R)-2-(hydroxymethyl)-6-[(E)-3-quinolin-6-ylprop-2-enyl]oxane-3,4,5-triol, Type 1 fimbrin D-mannose specific adhesin | | Authors: | Bouckaert, J, Bridot, C. | | Deposit date: | 2022-12-03 | | Release date: | 2023-02-15 | | Last modified: | 2023-03-08 | | Method: | X-RAY DIFFRACTION (2.995 Å) | | Cite: | Insightful Improvement in the Design of Potent Uropathogenic E. coli FimH Antagonists.
Pharmaceutics, 15, 2023
|
|
7PCD
 
 | | HER2 IN COMPLEX WITH A COVALENT INHIBITOR | | Descriptor: | 1-[4-[4-[[3,5-bis(chloranyl)-4-([1,2,4]triazolo[1,5-a]pyridin-7-yloxy)phenyl]amino]pyrimido[5,4-d]pyrimidin-6-yl]piperazin-1-yl]-4-(3-fluoranylazetidin-1-yl)butan-1-one, Receptor tyrosine-protein kinase erbB-2 | | Authors: | Bader, G. | | Deposit date: | 2021-08-03 | | Release date: | 2022-07-27 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Discovery of potent and selective HER2 inhibitors with efficacy against HER2 exon 20 insertion-driven tumors, which preserve wild-type EGFR signaling.
Nat Cancer, 3, 2022
|
|
