2I47
 
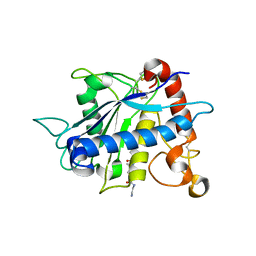 | | Crystal structure of catalytic domain of TACE with inhibitor | | Descriptor: | 4-({[4-(BUT-2-YN-1-YLOXY)PHENYL]SULFONYL}METHYL)-1-[(3,5-DIMETHYLISOXAZOL-4-YL)SULFONYL]-N-HYDROXYPIPERIDINE-4-CARBOXAMIDE, ADAM 17, N-{(2R)-2-[2-(hydroxyamino)-2-oxoethyl]-4-methylpentanoyl}-3-methyl-L-valyl-N-(2-aminoethyl)-L-alaninamide, ... | | Authors: | Xu, W, Condon, J.S, Lovering, F.E. | | Deposit date: | 2006-08-21 | | Release date: | 2006-12-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of potent and selective TACE inhibitors via the S1 pocket.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
9B9S
 
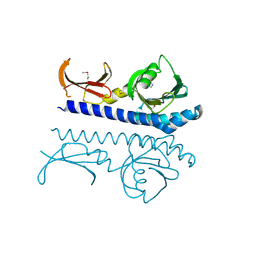 | | Crystal structure of the ligand binding domain of the Halomonas titanicae chemoreceptor Htc10 in complex with guanine | | Descriptor: | Chemotaxis protein, GUANINE | | Authors: | Ramos Ricciuti, F.E, Herrera Seitz, M.K, Gasperotti, A.F, Boyko, A, Jung, K, Bellinzoni, M, Lisa, M.N, Studdert, C.A. | | Deposit date: | 2024-04-03 | | Release date: | 2024-11-20 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | The chemoreceptor controlling the Wsp-like transduction pathway in Halomonas titanicae KHS3 binds and responds to purine derivatives.
Febs J., 292, 2025
|
|
3K0X
 | | Crystal structure of telomere capping protein Ten1 from Saccharomyces pombe | | Descriptor: | IODIDE ION, Protein Ten1 | | Authors: | Gelinas, A.D, Reyes, F.E, Batey, R.T, Wuttke, D.S. | | Deposit date: | 2009-09-25 | | Release date: | 2009-10-27 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Telomere capping proteins are structurally related to RPA with an additional telomere-specific domain.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
6OIF
 | | Cryo-EM structure of human TorsinA filament | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Torsin-1A | | Authors: | Zheng, W, Demircioglu, F.E, Schwartz, T.U, Egelman, E.H. | | Deposit date: | 2019-04-09 | | Release date: | 2019-07-24 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | The AAA + ATPase TorsinA polymerizes into hollow helical tubes with 8.5 subunits per turn.
Nat Commun, 10, 2019
|
|
7JPN
 
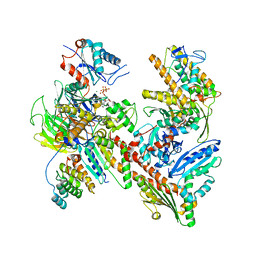 | | Cryo-EM structure of Arpin-bound Arp2/3 complex | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin-related protein 2, Actin-related protein 2/3 complex subunit 1B, ... | | Authors: | van Eeuwen, T, Fregoso, F.E, Dominguez, R, Zimmet, A, Boczkowska, M, Rebowski, G. | | Deposit date: | 2020-08-09 | | Release date: | 2022-02-09 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.24 Å) | | Cite: | Molecular mechanism of Arp2/3 complex inhibition by Arpin.
Nat Commun, 13, 2022
|
|
4L81
 
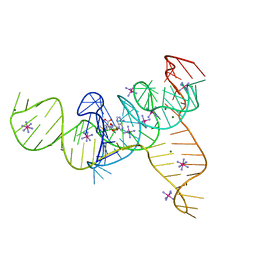 | | Structure of the SAM-I/IV riboswitch (env87(deltaU92, deltaG93)) | | Descriptor: | COBALT HEXAMMINE(III), MAGNESIUM ION, S-ADENOSYLMETHIONINE, ... | | Authors: | Trausch, J.J, Reyes, F.E, Edwards, A.L, Batey, R.T. | | Deposit date: | 2013-06-15 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural basis for diversity in the SAM clan of riboswitches.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
1EFP
 
 | | ELECTRON TRANSFER FLAVOPROTEIN (ETF) FROM PARACOCCUS DENITRIFICANS | | Descriptor: | ADENOSINE MONOPHOSPHATE, FLAVIN-ADENINE DINUCLEOTIDE, PROTEIN (ELECTRON TRANSFER FLAVOPROTEIN) | | Authors: | Roberts, D.L, Salazar, D, Fulmer, J.P, Frerman, F.E, Kim, J.J.-P. | | Deposit date: | 1998-12-18 | | Release date: | 1999-08-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of Paracoccus denitrificans electron transfer flavoprotein: structural and electrostatic analysis of a conserved flavin binding domain.
Biochemistry, 38, 1999
|
|
3SEM
 
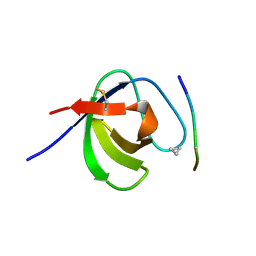 | | SEM5 SH3 DOMAIN COMPLEXED WITH PEPTOID INHIBITOR | | Descriptor: | SEX MUSCLE ABNORMAL PROTEIN 5, SH3 PEPTOID INHIBITOR | | Authors: | Nguyen, J.T, Turck, C.W, Cohen, F.E, Zuckermann, R.N, Lim, W.A. | | Deposit date: | 1998-11-02 | | Release date: | 1999-01-06 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Exploiting the basis of proline recognition by SH3 and WW domains: design of N-substituted inhibitors.
Science, 282, 1998
|
|
5K7P
 
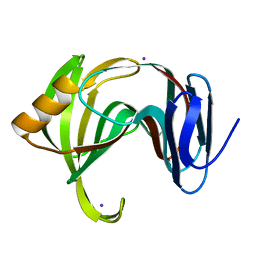 | | MicroED structure of xylanase at 2.3 A resolution | | Descriptor: | Endo-1,4-beta-xylanase 2, IODIDE ION | | Authors: | de la Cruz, M.J, Hattne, J, Shi, D, Seidler, P, Rodriguez, J, Reyes, F.E, Sawaya, M.R, Cascio, D, Eisenberg, D, Gonen, T. | | Deposit date: | 2016-05-26 | | Release date: | 2017-04-05 | | Last modified: | 2024-02-28 | | Method: | ELECTRON CRYSTALLOGRAPHY (2.3 Å) | | Cite: | Atomic-resolution structures from fragmented protein crystals with the cryoEM method MicroED.
Nat. Methods, 14, 2017
|
|
1LE5
 
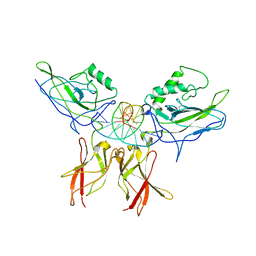 | | Crystal structure of a NF-kB heterodimer bound to an IFNb-kB | | Descriptor: | 5'-D(*AP*AP*GP*GP*AP*AP*TP*TP*TP*CP*CP*C)-3', 5'-D(*TP*GP*GP*GP*AP*AP*AP*TP*TP*CP*CP*T)-3', Nuclear factor NF-kappa-B p50 subunit, ... | | Authors: | Berkowitz, B, Huang, D.B, Chen-Park, F.E, Sigler, P.B, Ghosh, G. | | Deposit date: | 2002-04-09 | | Release date: | 2003-04-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | The X-ray crystal structure of the
NF-kB p50/p65 heterodimer bound
to the Interferon beta-kB site
J.Biol.Chem., 277, 2002
|
|
6GBM
 
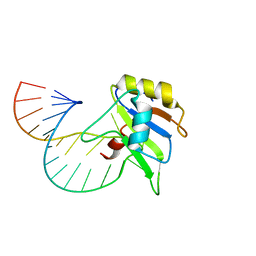 | | Solution structure of FUS-RRM bound to stem-loop RNA | | Descriptor: | RNA (5'-R(*GP*GP*CP*AP*GP*AP*UP*UP*AP*CP*AP*AP*UP*UP*CP*UP*AP*UP*UP*UP*GP*CP*C)-3'), RNA-binding protein FUS | | Authors: | Loughlin, F.E, Allain, F.H.-T. | | Deposit date: | 2018-04-15 | | Release date: | 2019-02-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Solution Structure of FUS Bound to RNA Reveals a Bipartite Mode of RNA Recognition with Both Sequence and Shape Specificity.
Mol. Cell, 73, 2019
|
|
6G99
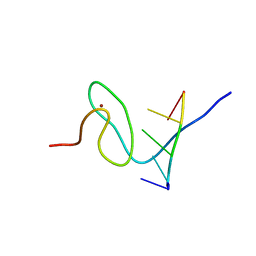 
 | | Solution structure of FUS-ZnF bound to UGGUG | | Descriptor: | RNA (5'-R(*UP*GP*GP*UP*G)-3'), RNA-binding protein FUS, ZINC ION | | Authors: | Loughlin, F.E, Allain, F.H.-T. | | Deposit date: | 2018-04-10 | | Release date: | 2019-02-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Solution Structure of FUS Bound to RNA Reveals a Bipartite Mode of RNA Recognition with Both Sequence and Shape Specificity.
Mol. Cell, 73, 2019
|
|
9B9X
 
 | | Crystal structure of the ligand binding domain of the Halomonas titanicae chemoreceptor Htc10 in complex with hypoxanthine | | Descriptor: | Chemotaxis protein, HYPOXANTHINE | | Authors: | Ramos Ricciuti, F.E, Herrera Seitz, M.K, Gasperotti, A.F, Boyko, A, Jung, K, Bellinzoni, M, Lisa, M.N, Studdert, C.A. | | Deposit date: | 2024-04-03 | | Release date: | 2024-11-20 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (3.63 Å) | | Cite: | The chemoreceptor controlling the Wsp-like transduction pathway in Halomonas titanicae KHS3 binds and responds to purine derivatives.
Febs J., 292, 2025
|
|
9BA3
 
 | | High-resolution crystal structure of the ligand binding domain of the Halomonas titanicae chemoreceptor Htc10 in complex with guanine | | Descriptor: | Chemotaxis protein, GUANINE | | Authors: | Ramos Ricciuti, F.E, Herrera Seitz, M.K, Gasperotti, A.F, Boyko, A, Jung, K, Bellinzoni, M, Lisa, M.N, Studdert, C.A. | | Deposit date: | 2024-04-03 | | Release date: | 2024-11-20 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The chemoreceptor controlling the Wsp-like transduction pathway in Halomonas titanicae KHS3 binds and responds to purine derivatives.
Febs J., 292, 2025
|
|
1DYZ
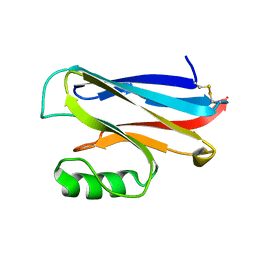 
 | | OXIDISED AZURIN II FROM ALCALIGENES XYLOSOXIDANS | | Descriptor: | AZURIN II, COPPER (II) ION | | Authors: | Dodd, F.E, Abraham, Z.H.L, Eady, R.R, Hasnain, S.S. | | Deposit date: | 2000-02-11 | | Release date: | 2000-07-10 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structures of Oxidised and Reduced Azurin II from Alcaligenes Xylosoxidans at 1.75 Angstoms Resolution
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1DO6
 
 | | CRYSTAL STRUCTURE OF SUPEROXIDE REDUCTASE IN THE OXIDIZED STATE AT 2.0 ANGSTROM RESOLUTION | | Descriptor: | FE (III) ION, SUPEROXIDE REDUCTASE | | Authors: | Yeh, A.P, Hu, Y, Jenney Junior, F.E, Adams, M.W, Rees, D.C. | | Deposit date: | 1999-12-19 | | Release date: | 2000-03-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of the superoxide reductase from Pyrococcus furiosus in the oxidized and reduced states.
Biochemistry, 39, 2000
|
|
3SD3
 
 | | The structure of the tetrahydrofolate riboswitch containing a U25C mutation | | Descriptor: | IRIDIUM HEXAMMINE ION, N-[4-({[(6S)-2-amino-5-formyl-4-oxo-3,4,5,6,7,8-hexahydropteridin-6-yl]methyl}amino)benzoyl]-L-glutamic acid, Tetrahydrofolate riboswitch | | Authors: | Reyes, F.E, Trausch, J.J, Ceres, P, Batey, R.T. | | Deposit date: | 2011-06-08 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The structure of a tetrahydrofolate-sensing riboswitch reveals two ligand binding sites in a single aptamer.
Structure, 19, 2011
|
|
1DZ0
 
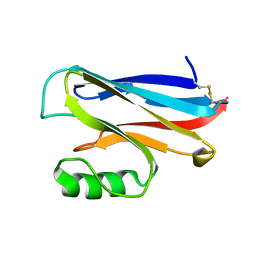 | | REDUCED AZURIN II FROM ALCALIGENES XYLOSOXIDANS | | Descriptor: | AZURIN II, COPPER (I) ION | | Authors: | Dodd, F.E, Abraham, Z.H.L, Eady, R.R, Hasnain, S.S. | | Deposit date: | 2000-02-11 | | Release date: | 2000-07-04 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structures of Oxidised and Reduced Azurin II from Alcaligenes Xylosoxidans at 1.75 Angstoms Resolution
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1DQI
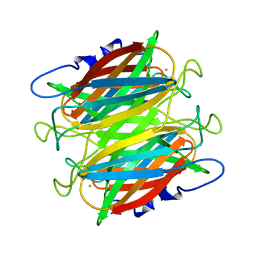 
 | | CRYSTAL STRUCTURE OF SUPEROXIDE REDUCTASE FROM P. FURIOSUS IN THE OXIDIZED STATE AT 1.7 ANGSTROMS RESOLUTION | | Descriptor: | FE (III) ION, SUPEROXIDE REDUCTASE | | Authors: | Yeh, A.P, Hu, Y, Jenney Jr, F.E, Adams, M.W.W, Rees, D.C. | | Deposit date: | 2000-01-04 | | Release date: | 2000-05-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of the superoxide reductase from Pyrococcus furiosus in the oxidized and reduced states.
Biochemistry, 39, 2000
|
|
1DQK
 
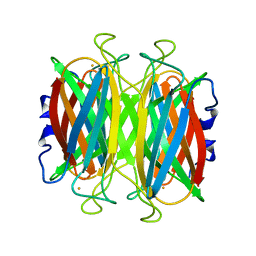 | | CRYSTAL STRUCTURE OF SUPEROXIDE REDUCTASE IN THE REDUCED STATE AT 2.0 ANGSTROMS RESOLUTION | | Descriptor: | FE (II) ION, SUPEROXIDE REDUCTASE | | Authors: | Yeh, A.P, Hu, Y, Jenney Jr, F.E, Adams, M.W.W, Rees, D.C. | | Deposit date: | 2000-01-04 | | Release date: | 2000-05-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of the superoxide reductase from Pyrococcus furiosus in the oxidized and reduced states.
Biochemistry, 39, 2000
|
|
5O0U
 
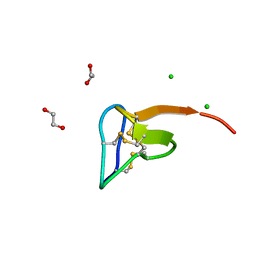 | | Crystal structure of tarantula venom peptide Protoxin-II | | Descriptor: | 1,2-ETHANEDIOL, Beta/omega-theraphotoxin-Tp2a, CHLORIDE ION | | Authors: | Tabor, A, McCarthy, S, Reyes, F.E. | | Deposit date: | 2017-05-17 | | Release date: | 2017-09-13 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | The Role of Disulfide Bond Replacements in Analogues of the Tarantula Toxin ProTx-II and Their Effects on Inhibition of the Voltage-Gated Sodium Ion Channel Nav1.7.
J.Am.Chem.Soc., 139, 2017
|
|
5NGZ
 
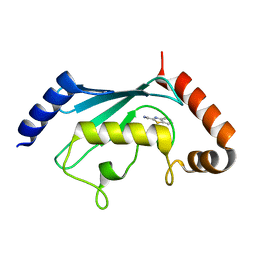 | | Ube2T in complex with fragment EM04 | | Descriptor: | 1-(1,3-benzothiazol-2-yl)methanamine, Ubiquitin-conjugating enzyme E2 T | | Authors: | Morreale, F.E, Bortoluzzi, A, Chaugule, V.K, Arkinson, C, Walden, H, Ciulli, A. | | Deposit date: | 2017-03-21 | | Release date: | 2017-05-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Allosteric Targeting of the Fanconi Anemia Ubiquitin-Conjugating Enzyme Ube2T by Fragment Screening.
J. Med. Chem., 60, 2017
|
|
5OJJ
 
 | | Crystal structure of the Zn-bound ubiquitin-conjugating enzyme Ube2T | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ACETATE ION, Ubiquitin-conjugating enzyme E2 T, ... | | Authors: | Morreale, F.E, Testa, A, Chaugule, V.K, Bortoluzzi, A, Ciulli, A, Walden, H. | | Deposit date: | 2017-07-21 | | Release date: | 2017-10-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mind the Metal: A Fragment Library-Derived Zinc Impurity Binds the E2 Ubiquitin-Conjugating Enzyme Ube2T and Induces Structural Rearrangements.
J. Med. Chem., 60, 2017
|
|
4ER4
 
 | | HIGH-RESOLUTION X-RAY ANALYSES OF RENIN INHIBITOR-ASPARTIC PROTEINASE COMPLEXES | | Descriptor: | ENDOTHIAPEPSIN, H-142 | | Authors: | Foundling, S.I, Watson, F.E, Szelke, M, Blundell, T.L. | | Deposit date: | 1991-01-05 | | Release date: | 1991-04-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | High resolution X-ray analyses of renin inhibitor-aspartic proteinase complexes.
Nature, 327, 1987
|
|
2XO0
 
 | | xpt-pbuX C74U Riboswitch from B. subtilis bound to 24-diamino-1,3,5- triazine identified by virtual screening | | Descriptor: | 1,3,5-TRIAZINE-2,4-DIAMINE, ACETATE ION, COBALT HEXAMMINE(III), ... | | Authors: | Daldrop, P, Reyes, F.E, Robinson, D.A, Hammond, C.M, Lilley, D.M.J, Batey, R.T, Brenk, R. | | Deposit date: | 2010-08-09 | | Release date: | 2011-04-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Novel ligands for a purine riboswitch discovered by RNA-ligand docking.
Chem. Biol., 18, 2011
|
|
