+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4y5g | ||||||
|---|---|---|---|---|---|---|---|
| Title | Endothiapepsin in complex with fragment 272 | ||||||
 Components Components | Endothiapepsin | ||||||
 Keywords Keywords | HYDROLASE / fragment screening / aspartic protease / inhibition | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Cryphonectria parasitica (chestnut blight fungus) Cryphonectria parasitica (chestnut blight fungus) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.47 Å MOLECULAR REPLACEMENT / Resolution: 1.47 Å | ||||||
 Authors Authors | Wang, X. / Heine, A. / Klebe, G. | ||||||
| Funding support |  Germany, 1items Germany, 1items
| ||||||
 Citation Citation |  Journal: to be published Journal: to be publishedTitle: Crystallographic Fragment Sreening of an Entire Library Authors: Wang, X. / Heine, A. / Klebe, G. #1:  Journal: J. Med. Chem. / Year: 2011 Journal: J. Med. Chem. / Year: 2011Title: A small nonrule of 3 compatible fragment library provides high hit rate of endothiapepsin crystal structures with various fragment chemotypes. Authors: Koester, H. / Craan, T. / Brass, S. / Herhaus, C. / Zentgraf, M. / Neumann, L. / Heine, A. / Klebe, G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4y5g.cif.gz 4y5g.cif.gz | 194.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4y5g.ent.gz pdb4y5g.ent.gz | 156.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4y5g.json.gz 4y5g.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/y5/4y5g https://data.pdbj.org/pub/pdb/validation_reports/y5/4y5g ftp://data.pdbj.org/pub/pdb/validation_reports/y5/4y5g ftp://data.pdbj.org/pub/pdb/validation_reports/y5/4y5g | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4y3aC  4y3hC  4y3lC  4y3tC  4y58C  4y5aC  4y5bC  4y5cC  4y5eC  4y5kC  3pcwS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 33813.855 Da / Num. of mol.: 1 / Source method: isolated from a natural source Source: (natural)  Cryphonectria parasitica (chestnut blight fungus) Cryphonectria parasitica (chestnut blight fungus)References: UniProt: P11838, endothiapepsin |
|---|
-Non-polymers , 5 types, 286 molecules 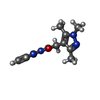








| #2: Chemical | ChemComp-47U / |
|---|---|
| #3: Chemical | ChemComp-GOL / |
| #4: Chemical | ChemComp-ACT / |
| #5: Chemical | ChemComp-DMS / |
| #6: Water | ChemComp-HOH / |
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.91 Å3/Da / Density % sol: 35.61 % |
|---|---|
| Crystal grow | Temperature: 290 K / Method: vapor diffusion, sitting drop / pH: 4.6 Details: 0.1 M ammonium acetate, 0.1 M sodium acetate, 24-30% PEG 4000. crystals obtained by streak-seeding |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  BESSY BESSY  / Beamline: 14.1 / Wavelength: 0.918409 Å / Beamline: 14.1 / Wavelength: 0.918409 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jul 19, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.918409 Å / Relative weight: 1 |
| Reflection | Resolution: 1.47→42.759 Å / Num. obs: 51878 / % possible obs: 93.5 % / Observed criterion σ(I): -3 / Redundancy: 2.3 % / Rsym value: 0.049 / Net I/σ(I): 12.9 |
| Reflection shell | Resolution: 1.47→1.56 Å / Redundancy: 2.31 % / Rmerge(I) obs: 0.277 / Mean I/σ(I) obs: 3.5 / % possible all: 91.7 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3PCW Resolution: 1.47→42.759 Å / SU ML: 0.13 / Cross valid method: FREE R-VALUE / σ(F): 1.36 / Phase error: 14.84 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.47→42.759 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller















































































 PDBj
PDBj




