+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2vwm | ||||||
|---|---|---|---|---|---|---|---|
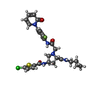
| Title | Aminopyrrolidine Factor Xa inhibitor | ||||||
 Components Components |
| ||||||
 Keywords Keywords | BLOOD CLOTTING / HYDROLASE / CATION / PLASMA / CALCIUM / ZYMOGEN / PROTEASE / INHIBITOR / POLYMORPHISM / GLYCOPROTEIN / GAMMA-CARBOXYGLUTAMIC ACID / COAGULATION FACTOR / HYDROXYLATION / SERINE PROTEASE / EGF-LIKE DOMAIN / CLEAVAGE ON PAIR OF BASIC RESIDUES | ||||||
| Function / homology |  Function and homology information Function and homology informationcoagulation factor Xa / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / : / positive regulation of TOR signaling / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / : / Gamma-carboxylation of protein precursors / Removal of aminoterminal propeptides from gamma-carboxylated proteins ...coagulation factor Xa / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / : / positive regulation of TOR signaling / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / : / Gamma-carboxylation of protein precursors / Removal of aminoterminal propeptides from gamma-carboxylated proteins / : / phospholipid binding / Golgi lumen / blood coagulation / positive regulation of cell migration / endoplasmic reticulum lumen / serine-type endopeptidase activity / external side of plasma membrane / calcium ion binding / proteolysis / : / extracellular region / plasma membrane Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.96 Å MOLECULAR REPLACEMENT / Resolution: 1.96 Å | ||||||
 Authors Authors | Groebke-Zbinden, K. / Banner, D.W. / Benz, J.M. / Blasco, F. / Decoret, G. / Himber, J. / Kuhn, B. / Panday, N. / Ricklin, F. / Risch, P. ...Groebke-Zbinden, K. / Banner, D.W. / Benz, J.M. / Blasco, F. / Decoret, G. / Himber, J. / Kuhn, B. / Panday, N. / Ricklin, F. / Risch, P. / Schlatter, D. / Stahl, M. / Unger, R. / Haap, W. | ||||||
 Citation Citation |  Journal: Eur.J.Med.Chem. / Year: 2009 Journal: Eur.J.Med.Chem. / Year: 2009Title: Design of Novel Aminopyrrolidine Factor Xa Inhibitors from a Screening Hit. Authors: Zbinden, K.G. / Anselm, L. / Banner, D.W. / Benz, J.M. / Blasco, F. / Decoret, G. / Himber, J. / Kuhn, B. / Panday, N. / Ricklin, F. / Risch, P. / Schlatter, D. / Stahl, M. / Thomi, S. / Unger, R. / Haap, W. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW ... SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 8-STRANDED BARREL THIS IS REPRESENTED BY A 9-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. THE SHEETS PRESENTED AS "AB" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 7-STRANDED BARREL THIS IS REPRESENTED BY A 8-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. THE SHEETS PRESENTED AS "BA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 8-STRANDED BARREL THIS IS REPRESENTED BY A 9-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. THE SHEETS PRESENTED AS "BB" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 6-STRANDED BARREL THIS IS REPRESENTED BY A 7-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2vwm.cif.gz 2vwm.cif.gz | 142.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2vwm.ent.gz pdb2vwm.ent.gz | 111.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2vwm.json.gz 2vwm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/vw/2vwm https://data.pdbj.org/pub/pdb/validation_reports/vw/2vwm ftp://data.pdbj.org/pub/pdb/validation_reports/vw/2vwm ftp://data.pdbj.org/pub/pdb/validation_reports/vw/2vwm | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2vvcSC  2vvuC  2vvvC  2vwlC  2vwnC  2vwoC C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| 5 | 
| ||||||||
| 6 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 27172.980 Da / Num. of mol.: 2 / Fragment: PEPTIDASE S1 DOMAIN, RESIDUES 235-475 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Production host: HOMO SAPIENS (human) / Production host:  #2: Protein | Mass: 6060.816 Da / Num. of mol.: 2 / Fragment: EGF2, RESIDUES 126-180 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Production host: HOMO SAPIENS (human) / Production host:  #3: Chemical | #4: Chemical | #5: Water | ChemComp-HOH / | Compound details | ENGINEERED | Has protein modification | Y | Nonpolymer details | HETGROUP LZI : THE CYCLOPROPYL-METHYL GROUP OF THE INHIBITOR HAS NO ELECTRON DENSITY.THE AMIDE ...HETGROUP LZI : THE CYCLOPROPY | Sequence details | ARG-GLU MUTANT.THE RESIDUE NUMBERING IN THE CATALYTIC DOMAIN FOLLOWS THAT OF CHYMOTRYPS | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.25 Å3/Da / Density % sol: 45.3 % / Description: NONE |
|---|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS FR591 / Wavelength: 1.5418 ROTATING ANODE / Type: ENRAF-NONIUS FR591 / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jan 19, 2005 / Details: OSMICS |
| Radiation | Monochromator: OSMICS / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.96→50 Å / Num. obs: 36648 / % possible obs: 91.6 % / Observed criterion σ(I): -3 / Redundancy: 4.01 % / Rmerge(I) obs: 0.04 / Net I/σ(I): 20.7 |
| Reflection shell | Resolution: 1.96→2.07 Å / Redundancy: 3.93 % / Rmerge(I) obs: 0.2 / Mean I/σ(I) obs: 6.57 / % possible all: 99.6 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2VVC Resolution: 1.96→74.74 Å / Cor.coef. Fo:Fc: 0.939 / Cor.coef. Fo:Fc free: 0.889 / SU B: 5.249 / SU ML: 0.153 / Cross valid method: THROUGHOUT / ESU R: 0.244 / ESU R Free: 0.214 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 30.68 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.96→74.74 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller













































































 PDBj
PDBj