-Search query
-Search result
Showing 1 - 50 of 501 items for (author: yang & oo)

EMDB-41409: 
Cryo-EM structure of PCSK9 mimic HIT01-K21Q-R218E with AMG145 Fab
Method: single particle / : Cheng J, Kwong PD

EMDB-43931: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB-43932: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axa: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axc: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X
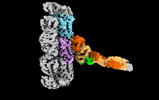
EMDB-18304: 
Outer kinetochore Ndc80-Dam1 alpha/beta-tubulin complex
Method: single particle / : Muir KW, Barford D

EMDB-18485: 
Ndc80c microtubule complex
Method: single particle / : Muir KW, Barford D

EMDB-38313: 
Structure of yeast replisome associated with FACT and histone hexamer, the region of FACT-Histones optimized local map
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

EMDB-38314: 
Structure of yeast replisome associated with FACT and histone hexamer,Conformation-2
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

EMDB-38315: 
Structure of yeast replisome associated with FACT and histone hexamer, the region of polymerase epsilon optimized local map
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

EMDB-38316: 
Structure of yeast replisome associated with FACT and histone hexamer, Conformation-1
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

EMDB-38317: 
Structure of yeast replisome associated with FACT and histone hexamer, Composite map
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

PDB-8xgc: 
Structure of yeast replisome associated with FACT and histone hexamer, Composite map
Method: single particle / : Li N, Gao Y, Yu D, Gao N, Zhai Y

EMDB-17557: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on mother filament
Method: single particle / : Liu T, Moores CA

EMDB-17553: 
Cryo-EM structure of cortactin-stabilized Arp2/3 complex nucleated actin branches-Daughter filament consensus map
Method: single particle / : Liu T, Moores CA

EMDB-17554: 
Cryo-EM structure of cortactin-stabilized Arp2/3 nucleated actin branches-Local refined map on Arp2/3 complex
Method: single particle / : Liu T, Moores CA

EMDB-17555: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on the daughter filament and cortactin density
Method: single particle / : Liu T, Moores CA

EMDB-17556: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on capping protein
Method: single particle / : Liu T, Moores CA

EMDB-17558: 
Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleated actin branches
Method: single particle / : Liu T, Moores CA

PDB-8p94: 
Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleated actin branches
Method: single particle / : Liu T, Moores CA
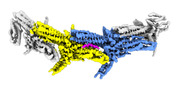
EMDB-18246: 
Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex
Method: single particle / : Muir KW, Barford D
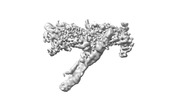
EMDB-18247: 
Outer kinetochore Dam1 protomer monomer Ndc80-Nuf2 coiled-coil complex
Method: single particle / : Muir KW, Barford D

PDB-8q84: 
Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex
Method: single particle / : Muir KW, Barford D

PDB-8q85: 
Outer kinetochore Dam1 protomer monomer Ndc80-Nuf2 coiled-coil complex
Method: single particle / : Muir KW, Barford D
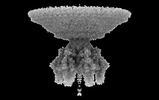
EMDB-41649: 
P22 Mature Virion tail - C6 Localized Reconstruction
Method: single particle / : Iglesias S, Cingolani G, Feng-Hou C

EMDB-41651: 
In situ cryo-EM structure of bacteriophage P22 portal protein: head-to-tail protein complex at 3.0A resolution
Method: single particle / : Iglesias SM, Cingolani G, Feng-Hou C

EMDB-41819: 
In situ cryo-EM structure of bacteriophage P22 tailspike protein complex at 3.4A resolution
Method: single particle / : Iglesias SM, Feng-Hou C, Cingolani G

PDB-8tvr: 
In situ cryo-EM structure of bacteriophage P22 tail hub protein: tailspike protein complex at 2.8A resolution
Method: single particle / : Iglesias S, Cingolani G, Feng-Hou C

PDB-8tvu: 
In situ cryo-EM structure of bacteriophage P22 portal protein: head-to-tail protein complex at 3.0A resolution
Method: single particle / : Iglesias SM, Cingolani G, Feng-Hou C

PDB-8u1o: 
In situ cryo-EM structure of bacteriophage P22 tailspike protein complex at 3.4A resolution
Method: single particle / : Iglesias SM, Feng-Hou C, Cingolani G

EMDB-41791: 
In situ cryo-EM structure of bacteriophage P22 gp1:gp4:gp5:gp10:gp9 N-term complex in conformation 1 at 3.2A resolution
Method: single particle / : Iglesias S, Feng-Hou C, Cingolani G

EMDB-41792: 
In situ cryo-EM structure of bacteriophage P22 gp1:gp5:gp4: gp10: gp9 N-term complex in conformation 2 at 3.1A resolution
Method: single particle / : Iglesias S, Feng-Hou C, Cingolani G

PDB-8u10: 
In situ cryo-EM structure of bacteriophage P22 gp1:gp4:gp5:gp10:gp9 N-term complex in conformation 1 at 3.2A resolution
Method: single particle / : Iglesias S, Feng-Hou C, Cingolani G

PDB-8u11: 
In situ cryo-EM structure of bacteriophage P22 gp1:gp5:gp4: gp10: gp9 N-term complex in conformation 2 at 3.1A resolution
Method: single particle / : Iglesias S, Feng-Hou C, Cingolani G

EMDB-28728: 
Structure of 3A10 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

EMDB-28729: 
Structure of 1F04 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

EMDB-28730: 
Structure of 3C08 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

PDB-8ez3: 
Structure of 3A10 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

PDB-8ez7: 
Structure of 1F04 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

PDB-8ez8: 
Structure of 3C08 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase
Method: single particle / : Mou Z, Lei R, Wu NC, Dai X

EMDB-28115: 
Western Equine Encephalitis Virus-Like Particle in Complex with SKW19 Fab
Method: single particle / : Pletnev S, Tsybovsky Y, Verardi R, Roedeger M, Kwong P

EMDB-28116: 
Western Equine Encephalitis Virus-Like Particle in Complex with SKW24 Fab
Method: single particle / : Pletnev S, Tsybovsky Y, Verardi R, Roedeger M, Kwong PD

EMDB-28117: 
Eastern Equine Encephalitis Virus-Like Particle in Complex with SKE26 Fab
Method: single particle / : Pletnev S, Verardi R, Roedeger M, Kwong P

PDB-8c5v: 
Chemotaxis core signalling unit from E protein lysed E. coli cells
Method: subtomogram averaging / : Cassidy CK, Qin Z, Zhang P

EMDB-34530: 
Membrane protein A
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-34531: 
Membrane protein B
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-35713: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Nakamura S, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8h86: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8h87: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR2 in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8iu0: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Nakamura S, Yamashita K, Fukuda M, Deisseroth K, Kato HE
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
