[English] 日本語
 Yorodumi
Yorodumi- EMDB-41791: In situ cryo-EM structure of bacteriophage P22 gp1:gp4:gp5:gp10:g... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In situ cryo-EM structure of bacteriophage P22 gp1:gp4:gp5:gp10:gp9 N-term complex in conformation 1 at 3.2A resolution | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phage / bacteriophage / tail spike protein / TSP / gene product 9 (gp9) / Packaged DNA stabilization protein / gene product 10 (gp10) / STRUCTURAL PROTEIN / VIRAL PROTEIN / head-to-tail protein / gene product 4 (gp4) / Tail hub protein / gene product (1) / Portal protein / Coat protein / Major capsid protein / gene product 5 (gp5) | |||||||||
| Function / homology |  Function and homology information Function and homology informationviral DNA genome packaging, headful / symbiont entry into host cell via disruption of host cell wall peptidoglycan / endo-1,3-alpha-L-rhamnosidase activity / symbiont entry into host cell via disruption of host cell envelope lipopolysaccharide / viral procapsid / viral portal complex / viral procapsid maturation / symbiont genome ejection through host cell envelope, short tail mechanism / viral DNA genome packaging / virus tail, fiber ...viral DNA genome packaging, headful / symbiont entry into host cell via disruption of host cell wall peptidoglycan / endo-1,3-alpha-L-rhamnosidase activity / symbiont entry into host cell via disruption of host cell envelope lipopolysaccharide / viral procapsid / viral portal complex / viral procapsid maturation / symbiont genome ejection through host cell envelope, short tail mechanism / viral DNA genome packaging / virus tail, fiber / symbiont entry into host cell via disruption of host cell envelope / virus tail / symbiont entry into host / T=7 icosahedral viral capsid / Hydrolases; Glycosylases; Glycosidases, i.e. enzymes that hydrolyse O- and S-glycosyl compounds / adhesion receptor-mediated virion attachment to host cell / virion assembly / virion component / viral capsid / lyase activity / hydrolase activity / virion attachment to host cell / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Salmonella phage P22 (virus) Salmonella phage P22 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Iglesias S / Feng-Hou C / Cingolani G | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2023 Journal: J Mol Biol / Year: 2023Title: Molecular Architecture of Salmonella Typhimurium Virus P22 Genome Ejection Machinery. Authors: Stephano M Iglesias / Ravi K Lokareddy / Ruoyu Yang / Fenglin Li / Daniel P Yeggoni / Chun-Feng David Hou / Makayla N Leroux / Juliana R Cortines / Justin C Leavitt / Mary Bird / Sherwood R ...Authors: Stephano M Iglesias / Ravi K Lokareddy / Ruoyu Yang / Fenglin Li / Daniel P Yeggoni / Chun-Feng David Hou / Makayla N Leroux / Juliana R Cortines / Justin C Leavitt / Mary Bird / Sherwood R Casjens / Simon White / Carolyn M Teschke / Gino Cingolani /   Abstract: Bacteriophage P22 is a prototypical member of the Podoviridae superfamily. Since its discovery in 1952, P22 has become a paradigm for phage transduction and a model for icosahedral viral capsid ...Bacteriophage P22 is a prototypical member of the Podoviridae superfamily. Since its discovery in 1952, P22 has become a paradigm for phage transduction and a model for icosahedral viral capsid assembly. Here, we describe the complete architecture of the P22 tail apparatus (gp1, gp4, gp10, gp9, and gp26) and the potential location and organization of P22 ejection proteins (gp7, gp20, and gp16), determined using cryo-EM localized reconstruction, genetic knockouts, and biochemical analysis. We found that the tail apparatus exists in two equivalent conformations, rotated by ∼6° relative to the capsid. Portal protomers make unique contacts with coat subunits in both conformations, explaining the 12:5 symmetry mismatch. The tail assembles around the hexameric tail hub (gp10), which folds into an interrupted β-propeller characterized by an apical insertion domain. The tail hub connects proximally to the dodecameric portal protein and head-to-tail adapter (gp4), distally to the trimeric tail needle (gp26), and laterally to six trimeric tailspikes (gp9) that attach asymmetrically to gp10 insertion domain. Cryo-EM analysis of P22 mutants lacking the ejection proteins gp7 or gp20 and biochemical analysis of purified recombinant proteins suggest that gp7 and gp20 form a molecular complex associated with the tail apparatus via the portal protein barrel. We identified a putative signal transduction pathway from the tailspike to the tail needle, mediated by three flexible loops in the tail hub, that explains how lipopolysaccharide (LPS) is sufficient to trigger the ejection of the P22 DNA in vitro. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41791.map.gz emd_41791.map.gz | 227.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41791-v30.xml emd-41791-v30.xml emd-41791.xml emd-41791.xml | 22.3 KB 22.3 KB | Display Display |  EMDB header EMDB header |
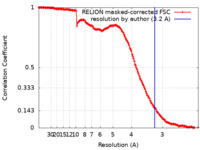
| FSC (resolution estimation) |  emd_41791_fsc.xml emd_41791_fsc.xml | 15.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_41791.png emd_41791.png | 59.4 KB | ||
| Masks |  emd_41791_msk_1.map emd_41791_msk_1.map | 325 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-41791.cif.gz emd-41791.cif.gz | 7.5 KB | ||
| Others |  emd_41791_half_map_1.map.gz emd_41791_half_map_1.map.gz emd_41791_half_map_2.map.gz emd_41791_half_map_2.map.gz | 259.5 MB 258.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41791 http://ftp.pdbj.org/pub/emdb/structures/EMD-41791 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41791 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41791 | HTTPS FTP |
-Related structure data
| Related structure data |  8u10MC  8tvrC  8tvuC  8u11C  8u1oC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41791.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41791.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.16582 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_41791_msk_1.map emd_41791_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
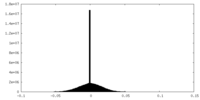
| Density Histograms |
-Half map: #2
| File | emd_41791_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
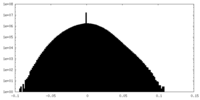
| Density Histograms |
-Half map: #1
| File | emd_41791_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Salmonella phage P22
| Entire | Name:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Salmonella phage P22
| Supramolecule | Name: Salmonella phage P22 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2908168 / Sci species name: Salmonella phage P22 / Virus type: VIRION / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Salmonella enterica (bacteria) Salmonella enterica (bacteria) |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
| Molecular weight | Theoretical: 82.829375 KDa |
| Sequence | String: MADNENRLES ILSRFDADWT ASDEARREAK NDLFFSRVSQ WDDWLSQYTT LQYRGQFDVV RPVVRKLVSE MRQNPIDVLY RPKDGARPD AADVLMGMYR TDMRHNTAKI AVNIAVREQI EAGVGAWRLV TDYEDQSPTS NNQVIRREPI HSACSHVIWD S NSKLMDKS ...String: MADNENRLES ILSRFDADWT ASDEARREAK NDLFFSRVSQ WDDWLSQYTT LQYRGQFDVV RPVVRKLVSE MRQNPIDVLY RPKDGARPD AADVLMGMYR TDMRHNTAKI AVNIAVREQI EAGVGAWRLV TDYEDQSPTS NNQVIRREPI HSACSHVIWD S NSKLMDKS DARHCTVIHS MSQNGWEDFA EKYDLDADDI PSFQNPNDWV FPWLTQDTIQ IAEFYEVVEK KETAFIYQDP VT GEPVSYF KRDIKDVIDD LADSGFIKIA ERQIKRRRVY KSIITCTAVL KDKQLIAGEH IPIVPVFGEW GFVEDKEVYE GVV RLTKDG QRLRNMIMSF NADIVARTPK KKPFFWPEQI AGFEHMYDGN DDYPYYLLNR TDENSGDLPT QPLAYYENPE VPQA NAYML EAATSAVKEV ATLGVDTEAV NGGQVAFDTV NQLNMRADLE TYVFQDNLAT AMRRDGEIYQ SIVNDIYDVP RNVTI TLED GSEKDVQLMA EVVDLATGEK QVLNDIRGRY ECYTDVGPSF QSMKQQNRAE ILELLGKTPQ GTPEYQLLLL QYFTLL DGK GVEMMRDYAN KQLIQMGVKK PETPEEQQWL VEAQQAKQGQ QDPAMVQAQG VLLQGQAELA KAQNQTLSLQ IDAAKVE AQ NQLNAARIAE IFNNMDLSKQ SEFREFLKTV ASFQQDRSED ARANAELLLK GDEQTHKQRM DIANILQSQR QNQPSGSV A ETPQ UniProtKB: Portal protein |
-Macromolecule #2: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 2 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
| Molecular weight | Theoretical: 46.795613 KDa |
| Sequence | String: MALNEGQIVT LAVDEIIETI SAITPMAQKA KKYTPPAASM QRSSNTIWMP VEQESPTQEG WDLTDKATGL LELNVAVNMG EPDNDFFQL RADDLRDETA YRRRIQSAAR KLANNVELKV ANMAAEMGSL VITSPDAIGT NTADAWNFVA DAEEIMFSRE L NRDMGTSY ...String: MALNEGQIVT LAVDEIIETI SAITPMAQKA KKYTPPAASM QRSSNTIWMP VEQESPTQEG WDLTDKATGL LELNVAVNMG EPDNDFFQL RADDLRDETA YRRRIQSAAR KLANNVELKV ANMAAEMGSL VITSPDAIGT NTADAWNFVA DAEEIMFSRE L NRDMGTSY FFNPQDYKKA GYDLTKRDIF GRIPEEAYRD GTIQRQVAGF DDVLRSPKLP VLTKSTATGI TVSGAQSFKP VA WQLDNDG NKVNVDNRFA TVTLSATTGM KRGDKISFAG VKFLGQMAKN VLAQDATFSV VRVVDGTHVE ITPKPVALDD VSL SPEQRA YANVNTSLAD AMAVNILNVK DARTNVFWAD DAIRIVSQPI PANHELFAGM KTTSFSIPDV GLNGIFATQG DIST LSGLC RIALWYGVNA TRPEAIGVGL PGQTA UniProtKB: Major capsid protein |
-Macromolecule #3: Packaged DNA stabilization protein gp10
| Macromolecule | Name: Packaged DNA stabilization protein gp10 / type: protein_or_peptide / ID: 3 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
| Molecular weight | Theoretical: 52.51202 KDa |
| Sequence | String: MPIQQLPMMK GMGKDFKNAD YIDYLPVNML ATPKEILNSS GYLRSFPGIT KRYDMNGVSR GVEYNTAQNA VYRVCGGKLY KGESEVGDV AGSGRVSMAH GRTSQAVGVN GQLVEYRYDG TVKTVSNWPA DSGFTQYELG SVRDITRLRG RYAWSKDGTD S WFITDLED ...String: MPIQQLPMMK GMGKDFKNAD YIDYLPVNML ATPKEILNSS GYLRSFPGIT KRYDMNGVSR GVEYNTAQNA VYRVCGGKLY KGESEVGDV AGSGRVSMAH GRTSQAVGVN GQLVEYRYDG TVKTVSNWPA DSGFTQYELG SVRDITRLRG RYAWSKDGTD S WFITDLED ESHPDRYSAQ YRAESQPDGI IGIGTWRDFI VCFGSSTIEY FSLTGATTAG AALYVAQPSL MVQKGIAGTY CK TPFADSY AFISHPATGA PSVYIIGSGQ ASPIATASIE KIIRSYTAEE MATGVMETLR FDSHELLIIH LPRHVLVYDA SSS QNGPQW CVLKTGLYDD VYRGVDFMYE GNQITCGDKS EAVVGQLQFD ISSQYDKQQE HLLFTPLFKA DNARCFDLEV ESST GVAQY ADRLFLSATT DGINYGREQM IEQNEPFVYD KRVLWKRVGR IRRLIGFKLR VITKSPVTLS GCQIRLE UniProtKB: Tail hub protein gp10 |
-Macromolecule #4: Tail spike protein
| Macromolecule | Name: Tail spike protein / type: protein_or_peptide / ID: 4 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
| Molecular weight | Theoretical: 71.923523 KDa |
| Sequence | String: MTDITANVVV SNPRPIFTES RSFKAVANGK IYIGQIDTDP VNPANQIPVY IENEDGSHVQ ITQPLIINAA GKIVYNGQLV KIVTVQGHS MAIYDANGSQ VDYIANVLKY DPDQYSIEAD KKFKYSVKLS DYPTLQDAAS AAVDGLLIDR DYNFYGGETV D FGGKVLTI ...String: MTDITANVVV SNPRPIFTES RSFKAVANGK IYIGQIDTDP VNPANQIPVY IENEDGSHVQ ITQPLIINAA GKIVYNGQLV KIVTVQGHS MAIYDANGSQ VDYIANVLKY DPDQYSIEAD KKFKYSVKLS DYPTLQDAAS AAVDGLLIDR DYNFYGGETV D FGGKVLTI ECKAKFIGDG NLIFTKLGKG SRIAGVFMES TTTPWVIKPW TDDNQWLTDA AAVVATLKQS KTDGYQPTVS DY VKFPGIE TLLPPNAKGQ NITSTLEIRE CIGVEVHRAS GLMAGFLFRG CHFCKMVDAN NPSGGKDGII TFENLSGDWG KGN YVIGGR TSYGSVSSAQ FLRNNGGFER DGGVIGFTSY RAGESGVKTW QGTVGSTTSR NYNLQFRDSV VIYPVWDGFD LGAD TDMNP ELDRPGDYPI TQYPLHQLPL NHLIDNLLVR GALGVGFGMD GKGMYVSNIT VEDCAGSGAY LLTHESVFTN IAIID TNTK DFQANQIYIS GACRVNGLRL IGIRSTDGQG LTIDAPNSTV SGITGMVDPS RINVANLAEE GLGNIRANSF GYDSAA IKL RIHKLSKTLD SGALYSHING GAGSGSAYTQ LTAISGSTPD AVSLKVNHKD CRGAEIPFVP DIASDDFIKD SSCFLPY WE NNSTSLKALV KKPNGELVRL TLATL UniProtKB: Tail spike protein |
-Macromolecule #5: Peptidoglycan hydrolase gp4
| Macromolecule | Name: Peptidoglycan hydrolase gp4 / type: protein_or_peptide / ID: 5 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella phage P22 (virus) Salmonella phage P22 (virus) |
| Molecular weight | Theoretical: 18.044959 KDa |
| Sequence | String: MQIKTKGDLV RAALRKLGVA SDATLTDVEP QSMQDAVDDL EAMMAEWYQD GKGIITGYVF SDDENPPAEG DDHGLRSSAV SAVFHNLAC RIAPDYALEA TAKIIATAKY GKELLYKQTA ISRAKRAPYP SRMPTGSGNS FANLNEWHYF PGEQNADSTT P HDEGNG UniProtKB: Head-to-tail adapter gp4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 11520 pixel / Digitization - Dimensions - Height: 8184 pixel / Average electron dose: 1.08 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 2.1 µm / Calibrated defocus min: 0.8 µm / Calibrated magnification: 29000 / Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8u10: |
 Movie
Movie Controller
Controller








 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)