-Search query
-Search result
Showing 1 - 50 of 350 items for (author: wang & mm)
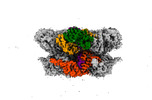
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
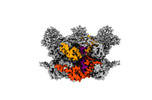
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
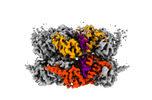
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
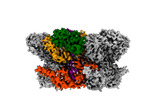
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
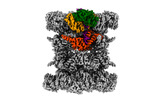
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)
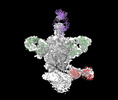
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.

EMDB-44635: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM

PDB-9bjk: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM
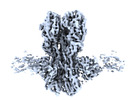
EMDB-41874: 
CryoEM structure of A/Solomon Islands/3/2006 H1 HA in complex with 05.GC.w2.3C10-H1_SI06

EMDB-17377: 
Structure of human SIT1 (focussed map / refinement)

EMDB-17378: 
Structure of human SIT1:ACE2 complex (open PD conformation)

EMDB-17379: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

EMDB-17380: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

EMDB-17381: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

EMDB-17382: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

PDB-8p2w: 
Structure of human SIT1 (focussed map / refinement)

PDB-8p2x: 
Structure of human SIT1:ACE2 complex (open PD conformation)

PDB-8p2y: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

PDB-8p2z: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

PDB-8p30: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

PDB-8p31: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

EMDB-17197: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-19108: 
Human TPC2 in Complex withAntagonist (R)-SG-094

PDB-8ouo: 
Human TPC2 in Complex with Antagonist (S)-SG-094
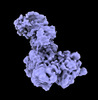
EMDB-40261: 
DDB1/CRBN in complex with ARV-471 and the ER ligand-binding domain

EMDB-42676: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

EMDB-42999: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

PDB-8uwl: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

PDB-8v6u: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

EMDB-19798: 
human PLD3 homodimer structure

PDB-8s86: 
human PLD3 homodimer structure

EMDB-38453: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)

EMDB-38454: 
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2

EMDB-43593: 
Langya Virus attachment (G) glycoprotein with K85L/L86K mutation

EMDB-37648: 
SARS-CoV-2 EG.5.1 spike glycoprotein (1-up state)

EMDB-37650: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-2 state)

EMDB-37651: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-1 state)

EMDB-18792: 
Capsid structure of Giardiavirus (GLV) CAT strain

EMDB-18791: 
Capsid structure of Giardiavirus (GLV) HP strain

EMDB-41235: 
VMAT1 dimer with MPP+ and reserpine

EMDB-41236: 
VMAT1 dimer with amphetamine and reserpine

EMDB-41237: 
VMAT1 dimer with dopamine and reserpine
EMDB-41238: 
VMAT1 dimer in unbound form and with reserpine

EMDB-41239: 
VMAT1 dimer with histamine and reserpine

EMDB-41240: 
VMAT1 dimer with norepinephrine and reserpine

EMDB-41241: 
VMAT1 dimer with reserpine

EMDB-41242: 
VMAT1 dimer with serotonin and reserpine

PDB-8tgg: 
VMAT1 dimer with MPP+ and reserpine
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
