-Search query
-Search result
Showing 1 - 50 of 90 items for (author: tampe & r)

EMDB-53326: 
Consensus Map of the Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-54377: 
Heterodimeric ABC exporter TmrAB (EQ mutant) in ATP-bound outward-facing occluded conformation in the absence of Mg2+
Method: single particle / : Susac L, Nocker C, Tampe R

EMDB-54378: 
Heterodimeric ABC exporter TmrAB (wild type) in ATP-bound outward-facing occluded conformation in the absence of Mg2+
Method: single particle / : Susac L, Nocker C, Tampe R

PDB-9rye: 
Heterodimeric ABC exporter TmrAB (EQ mutant) in ATP-bound outward-facing occluded conformation in the absence of Mg2+
Method: single particle / : Susac L, Nocker C, Tampe R

PDB-9ryf: 
Heterodimeric ABC exporter TmrAB (wild type) in ATP-bound outward-facing occluded conformation in the absence of Mg2+
Method: single particle / : Susac L, Nocker C, Tampe R

EMDB-53330: 
Central Tapasin Scaffold of the Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-53331: 
Editing Module 2 of the Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-53332: 
Translocation Module of the Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-53334: 
Editing Module 1 of the Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-53923: 
Structure of the Human Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

PDB-9rcv: 
Structure of the Human Peptide-Loading Complex Arrested by HCMV US6
Method: single particle / : Stolz M, Susac L, Trowitzsch S, Tampe R

EMDB-48548: 
SARS-CoV-2 S2 monomer in complex with R125-61 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-48549: 
SARS-CoV-2 S2 monomer in complex with NICA01B-1113 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-48550: 
SARS-CoV-2 S2 monomer in complex with NICA01A-1401 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

PDB-9mr1: 
SARS-CoV-2 S2 monomer in complex with R125-61 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

PDB-9mr2: 
SARS-CoV-2 S2 monomer in complex with NICA01A-1401 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-16976: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

PDB-8omr: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

EMDB-14923: 
Structure of the human tRNA splicing endonuclease defines substrate recognition
Method: single particle / : Sekulovski S, Trowitzsch S

EMDB-14925: 
Human TSEN with pre-tRNA-Tyr-GTA
Method: single particle / : Sekulovski S, Trowitzsch S

EMDB-13427: 
Structure of a fully assembled T-cell receptor engaging a tumor-associated peptide-MHC I
Method: single particle / : Susac L, Thomas C

PDB-7phr: 
Structure of a fully assembled T-cell receptor engaging a tumor-associated peptide-MHC I
Method: single particle / : Susac L, Thomas C, Tampe R

EMDB-14119: 
Structure of the human MHC I peptide-loading complex editing module
Method: single particle / : Domnick A, Susac L

PDB-7qpd: 
Structure of the human MHC I peptide-loading complex editing module
Method: single particle / : Domnick A, Susac L, Trowitzsch S, Thomas C, Tampe R

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
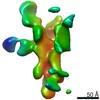
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
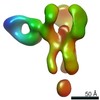
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
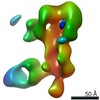
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
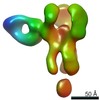
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25646: 
Negative stain map of polyclonal Fab 241.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25655: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB

PDB-7t3d: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23792: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A
EMDB-23793: 
Negative stain map of monoclonal Fab SFV009 2G01 binding the RBS of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23794: 
Negative stain map of monoclonal Fab 045-09 2B05 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23795: 
Negative stain map of monoclonal Fab SFV019 2A06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
EMDB-23796: 
Negative stain map of monoclonal Fab SFV015 2F02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23797: 
Negative stain map of monoclonal Fab 047-09 4G02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
EMDB-23798: 
Negative stain map of monoclonal Fab 047-09 4B06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

PDB-7mem: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A

EMDB-10519: 
Structure of an archaeal ABCE1-bound ribosomal post-splitting complex
Method: single particle / : Kratzat H, Becker T

PDB-6tmf: 
Structure of an archaeal ABCE1-bound ribosomal post-splitting complex
Method: single particle / : Kratzat H, Becker T, Tampe R, Beckmann R

EMDB-4773: 
Heterodimeric ABC exporter TmrAB in inward-facing narrow conformation under turnover conditions
Method: single particle / : Januliene D, Hofmann S, Medhdipour AR, Thomas C, Hummer G, Tampe R, Moeller A
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
