-Search query
-Search result
Showing 1 - 50 of 1,737 items for (author: steve & m)

EMDB-41570: 
Cryo-EM structure of the rat P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE

EMDB-41581: 
Cryo-EM structure of the rat P2X7 receptor in complex with the high-affinity agonist BzATP
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE

EMDB-42976: 
Cryo-EM structure of the rat P2X7 receptor in the apo closed state purified in the absence of sodium
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE

PDB-8tr5: 
Cryo-EM structure of the rat P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE

PDB-8trj: 
Cryo-EM structure of the rat P2X7 receptor in complex with the high-affinity agonist BzATP
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE

PDB-8v4s: 
Cryo-EM structure of the rat P2X7 receptor in the apo closed state purified in the absence of sodium
Method: single particle / : Oken AC, Lisi NE, Krishnamurthy I, McCarthy AE, Godsey MH, Glasfeld A, Mansoor SE
EMDB-45078: 
E.coli GroEL + PBZ1587 inhibitor
Method: single particle / : Watson ER, Lander GC
EMDB-45079: 
E.coli GroEL apoenzyme
Method: single particle / : Watson ER, Lander GC
EMDB-45080: 
E.Faecium GroEL
Method: single particle / : Watson ER, Lander GC

EMDB-45098: 
E.coli GroEL + PBZ1587 inhibitor C1 reconstruction
Method: single particle / : Watson ER, Lander GC
EMDB-36897: 
The cryo-EM map of close TIEA-TIC complex
Method: single particle / : Zhang KN, Liu Y, Chen M, Wang Y, Lin W, Li M, Zhang X, Gao Y, Gong Q, Chen H, Steve M, Li S, Zhang K, Liu B

PDB-8k58: 
The cryo-EM map of close TIEA-TIC complex
Method: single particle / : Zhang KN, Liu Y, Chen M, Wang Y, Lin W, Li M, Zhang X, Gao Y, Gong Q, Chen H, Steve M, Li S, Zhang K, Liu B

EMDB-36899: 
The cryo-EM map of open TIEA-TIC complex
Method: single particle / : Zhang KN, Liu Y, Chen M, Wang Y, Lin W, Li M, Zhang X, Gao Y, Gong Q, Chen H, Steve M, Li S, Zhang K, Liu B

PDB-8k5a: 
The cryo-EM map of open TIEA-TIC complex
Method: single particle / : Zhang KN, Liu Y, Chen M, Wang Y, Lin W, Li M, Zhang X, Gao Y, Gong Q, Chen H, Steve M, Li S, Zhang K, Liu B

EMDB-43385: 
Cryo-EM map of close dodecameric CaMKII beta holoenzyme T287A T306A T307A
Method: single particle / : Chien CT, Chiu W, Khan S

EMDB-43296: 
Constituent EM map: Focused refinement S100A1 of mouse RyR1 in complex with S100A1 (EGTA-only dataset)
Method: single particle / : Weninger G, Marks AR

EMDB-40873: 
Cryo-EM structure of tetradecameric hub domain of CaMKII alpha
Method: single particle / : Chien CT, Chiu W, Khan S

EMDB-41070: 
Cryo-EM structure of tetradecameric CaMKII beta holoenzyme T287A T306A T307A
Method: single particle / : Chien CT, Chiu W, Khan S
EMDB-41077: 
Cryo-EM structure of dodecameric CaMKII beta holoenzyme T287A T306A T307A
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8syg: 
Cryo-EM structure of tetradecameric hub domain of CaMKII alpha
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8t6k: 
Cryo-EM structure of tetradecameric CaMKII beta holoenzyme T287A T306A T307A
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8t6q: 
Cryo-EM structure of dodecameric CaMKII beta holoenzyme T287A T306A T307A
Method: single particle / : Chien CT, Chiu W, Khan S

EMDB-40955: 
Cryo-EM structure of dodecameric hub domain of CaMKII alpha
Method: single particle / : Chien CT, Chiu W, Khan S

EMDB-40956: 
Cryo-EM structure of tetradecameric hub domain of CaMKII beta
Method: single particle / : Chien CT, Chiu W, Khan S

EMDB-40957: 
Cryo-EM structure of dodecameric hub domain of CaMKII beta
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8t15: 
Cryo-EM structure of dodecameric hub domain of CaMKII alpha
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8t17: 
Cryo-EM structure of tetradecameric hub domain of CaMKII beta
Method: single particle / : Chien CT, Chiu W, Khan S

PDB-8t18: 
Cryo-EM structure of dodecameric hub domain of CaMKII beta
Method: single particle / : Chien CT, Chiu W, Khan S
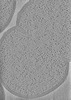

EMDB-42074: 
Representative tomogram of Enterococcus faecium WT Com15
Method: electron tomography / : Hang HC, Park D
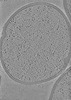

EMDB-42086: 
Representative tomogram of Enterococcus faecium SagA complementation strain
Method: electron tomography / : Hang HC, Park D

EMDB-42087: 
Representative tomogram of Enterococcus faecium SagA deletion strain
Method: electron tomography / : Hang HC, Park D

EMDB-16543: 
Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16544: 
Structure of human mitochondrial RNase Z in complex with mitochondrial pre-tRNA-His(0,Ser)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, YU W, LUISI B, TISNE C

EMDB-16545: 
Structure of human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNA-Ile
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C
EMDB-16547: 
Structure of human mitochondrial MRPP1-MRPP2 in complex with mitochondrial pre-tRNA-Ile
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16667: 
Consensus Refinement - Map 1
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16668: 
Local refinement - Map 2
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16669: 
Cryo-EM structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16673: 
Consensus Refinement - Map 1
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, YU W, LUISI B, TISNE C

EMDB-16674: 
Local refinement - Map 2 (mask 1)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, YU W, LUISI B, TISNE C

EMDB-16675: 
Local refinement - map 3 (Mask 2)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, YU W, LUISI B, TISNE C

EMDB-16690: 
Consensus Refinement - Map 1
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16691: 
Local refinement - Map 2 (Mask 1)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

EMDB-16692: 
Global Refinement - Map 1
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

PDB-8cbk: 
Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C

PDB-8cbl: 
Structure of human mitochondrial RNase Z in complex with mitochondrial pre-tRNA-His(0,Ser)
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, YU W, LUISI B, TISNE C

PDB-8cbm: 
Structure of human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNA-Ile
Method: single particle / : MEYNIER V, HARDWICK S, CATALA M, ROSKE J, OERUM S, CHIRGADZE D, BARRAUD P, LUISI B, TISNE C
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
