-検索条件
-検索結果
検索 (著者・登録者: sieber & s)の結果102件中、1から50件目までを表示しています
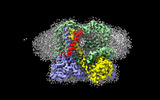
EMDB-15616: 
Four subunit cytochrome b-c1 complex from Rhodobacter sphaeroides in native nanodiscs - consensus refinement in the b-b conformation
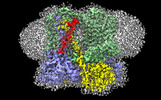
EMDB-15617: 
Four subunit cytochrome b-c1 complex from Rhodobacter sphaeroides in native nanodiscs - focussed refinement in the b-c conformation

PDB-8asi: 
Four subunit cytochrome b-c1 complex from Rhodobacter sphaeroides in native nanodiscs - consensus refinement in the b-b conformation

PDB-8asj: 
Four subunit cytochrome b-c1 complex from Rhodobacter sphaeroides in native nanodiscs - focussed refinement in the b-c conformation

EMDB-27070: 
apo form Cryo-EM structure of Campylobacter jejune ketol-acid reductoisommerase crosslinked by Glutaraldehyde

EMDB-14224: 
2.8 Angstrom cryo-EM structure of the dimeric cytochrome b6f-PetP complex from Synechocystis sp. PCC 6803 with natively bound lipids and plastoquinone molecules
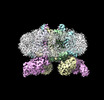
EMDB-15017: 
3.15 Angstrom cryo-EM structure of the dimeric cytochrome b6f complex from Synechocystis sp. PCC 6803 with natively bound plastoquinone and lipid molecules.

PDB-7r0w: 
2.8 Angstrom cryo-EM structure of the dimeric cytochrome b6f-PetP complex from Synechocystis sp. PCC 6803 with natively bound lipids and plastoquinone molecules

PDB-7zxy: 
3.15 Angstrom cryo-EM structure of the dimeric cytochrome b6f complex from Synechocystis sp. PCC 6803 with natively bound plastoquinone and lipid molecules.

EMDB-14332: 
1.58 A STRUCTURE OF HUMAN APOFERRITIN OBTAINED FROM TITAN KRIOS 2 AT eBIC, DLS UNDER COMMISSIONING SESSION CM26464-2

PDB-7r5o: 
1.58 A STRUCTURE OF HUMAN APOFERRITIN OBTAINED FROM TITAN KRIOS 2 AT eBIC, DLS UNDER COMMISSIONING SESSION CM26464-2

EMDB-31081: 
Subtomogram average of a meiotic triple helix from a pachytene S. cerevisiae cell.

EMDB-12679: 
Cryo-EM structure (model_1a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.4 A

EMDB-12680: 
Cryo-EM structure (model_2a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.5 A

EMDB-12681: 
Cryo-EM structure of the RC-dLH complex (model_1b) from Gemmatimonas phototrophica at 2.47 A

EMDB-12682: 
Cryo-EM structure (model_2b) of the RC-dLH complex from Gemmatimonas phototrophica at 2.44 A

PDB-7o0u: 
Cryo-EM structure (model_1a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.4 A

PDB-7o0v: 
Cryo-EM structure (model_2a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.5 A

PDB-7o0w: 
Cryo-EM structure of the RC-dLH complex (model_1b) from Gemmatimonas phototrophica at 2.47 A

PDB-7o0x: 
Cryo-EM structure (model_2b) of the RC-dLH complex from Gemmatimonas phototrophica at 2.44 A

EMDB-12274: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

EMDB-12275: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-88 Fab

EMDB-12276: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-150 Fab

EMDB-12277: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

EMDB-12278: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-316 Fab

EMDB-12279: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-384 Fab

EMDB-12280: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-253H55L Fab

EMDB-12281: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-253H55L Fab

EMDB-12282: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-253H165L Fab

EMDB-12283: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-159

EMDB-12284: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-159

PDB-7nd3: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

PDB-7nd4: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-88 Fab

PDB-7nd5: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-150 Fab

PDB-7nd6: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

PDB-7nd7: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-316 Fab

PDB-7nd8: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-384 Fab

PDB-7nd9: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-253H55L Fab

PDB-7nda: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-253H55L Fab

PDB-7ndb: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-253H165L Fab

PDB-7ndc: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-159

PDB-7ndd: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-159

EMDB-10316: 
Hepatitis B virus core protein virus-like particle displaying the antigen: extra cellular adhesion domain NadA from Neisseria meningitidis.

PDB-6tik: 
Hepatitis B virus core shell--virus-like particle with NadA epitope

EMDB-10162: 
Cryo-EM structure of the ClpXP1/2 protein degradation machinery

PDB-6sfw: 
Cryo-EM Structure of the ClpX component of the ClpXP1/2 degradation machinery.

PDB-6sfx: 
Cryo-EM structure of ClpP1/2 in the LmClpXP1/2 complex

PDB-6q81: 
Structure of P-glycoprotein(ABCB1) in the post-hydrolytic state

EMDB-3882: 
Thermus thermophilus PilF ATPase (apoprotein form)

EMDB-4194: 
Thermus thermophilus PilF ATPase (AMPPNP-bound form)
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
