[English] 日本語
 Yorodumi
Yorodumi- EMDB-14224: 2.8 Angstrom cryo-EM structure of the dimeric cytochrome b6f-PetP... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 2.8 Angstrom cryo-EM structure of the dimeric cytochrome b6f-PetP complex from Synechocystis sp. PCC 6803 with natively bound lipids and plastoquinone molecules | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Complex / cytochrome / membrane protein / electron transport / PHOTOSYNTHESIS | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcytochrome b6f complex / cytochrome complex assembly / photosynthetic electron transport chain / plasma membrane-derived thylakoid membrane / membrane => GO:0016020 / photosynthesis / respiratory electron transport chain / electron transfer activity / oxidoreductase activity / iron ion binding ...cytochrome b6f complex / cytochrome complex assembly / photosynthetic electron transport chain / plasma membrane-derived thylakoid membrane / membrane => GO:0016020 / photosynthesis / respiratory electron transport chain / electron transfer activity / oxidoreductase activity / iron ion binding / heme binding / membrane / metal ion binding Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||||||||
 Authors Authors | Farmer DF / Proctor MS / Malone LA / Swainsbury DPK / Hawkings FR / Hitchcock A / Johnson MP | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Biochem J / Year: 2022 Journal: Biochem J / Year: 2022Title: Cryo-EM structures of the Synechocystis sp. PCC 6803 cytochrome b6f complex with and without the regulatory PetP subunit. Authors: Matthew S Proctor / Lorna A Malone / David A Farmer / David J K Swainsbury / Frederick R Hawkings / Federica Pastorelli / Thomas Z Emrich-Mills / C Alistair Siebert / C Neil Hunter / Matthew ...Authors: Matthew S Proctor / Lorna A Malone / David A Farmer / David J K Swainsbury / Frederick R Hawkings / Federica Pastorelli / Thomas Z Emrich-Mills / C Alistair Siebert / C Neil Hunter / Matthew P Johnson / Andrew Hitchcock /  Abstract: In oxygenic photosynthesis, the cytochrome b6f (cytb6f) complex links the linear electron transfer (LET) reactions occurring at photosystems I and II and generates a transmembrane proton gradient via ...In oxygenic photosynthesis, the cytochrome b6f (cytb6f) complex links the linear electron transfer (LET) reactions occurring at photosystems I and II and generates a transmembrane proton gradient via the Q-cycle. In addition to this central role in LET, cytb6f also participates in a range of processes including cyclic electron transfer (CET), state transitions and photosynthetic control. Many of the regulatory roles of cytb6f are facilitated by auxiliary proteins that differ depending upon the species, yet because of their weak and transient nature the structural details of these interactions remain unknown. An apparent key player in the regulatory balance between LET and CET in cyanobacteria is PetP, a ∼10 kDa protein that is also found in red algae but not in green algae and plants. Here, we used cryogenic electron microscopy to determine the structure of the Synechocystis sp. PCC 6803 cytb6f complex in the presence and absence of PetP. Our structures show that PetP interacts with the cytoplasmic side of cytb6f, displacing the C-terminus of the PetG subunit and shielding the C-terminus of cytochrome b6, which binds the heme cn cofactor that is suggested to mediate CET. The structures also highlight key differences in the mode of plastoquinone binding between cyanobacterial and plant cytb6f complexes, which we suggest may reflect the unique combination of photosynthetic and respiratory electron transfer in cyanobacterial thylakoid membranes. The structure of cytb6f from a model cyanobacterial species amenable to genetic engineering will enhance future site-directed mutagenesis studies of structure-function relationships in this crucial ET complex. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14224.map.gz emd_14224.map.gz | 15.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14224-v30.xml emd-14224-v30.xml emd-14224.xml emd-14224.xml | 37.3 KB 37.3 KB | Display Display |  EMDB header EMDB header |
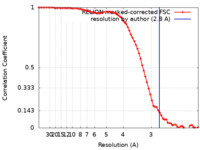
| FSC (resolution estimation) |  emd_14224_fsc.xml emd_14224_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_14224.png emd_14224.png | 65.6 KB | ||
| Filedesc metadata |  emd-14224.cif.gz emd-14224.cif.gz | 9.4 KB | ||
| Others |  emd_14224_half_map_1.map.gz emd_14224_half_map_1.map.gz emd_14224_half_map_2.map.gz emd_14224_half_map_2.map.gz | 23.4 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14224 http://ftp.pdbj.org/pub/emdb/structures/EMD-14224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14224 | HTTPS FTP |
-Related structure data
| Related structure data |  7r0wMC  7zxyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14224.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14224.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_14224_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14224_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Cytochrome b6f complex in association with PetP
+Supramolecule #1: Cytochrome b6f complex in association with PetP
+Macromolecule #1: Uncharacterized protein Cp097, conserved in cyanobacteria
+Macromolecule #2: Cytochrome b6
+Macromolecule #3: Cytochrome b6-f complex subunit 4
+Macromolecule #4: Cytochrome f
+Macromolecule #5: Cytochrome B6
+Macromolecule #6: Cytochrome b6-f complex subunit 7
+Macromolecule #7: Cytochrome b6-f complex subunit 5
+Macromolecule #8: Cytochrome b6-f complex subunit 8
+Macromolecule #9: Rieske domain, PetC
+Macromolecule #10: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #11: (1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-...
+Macromolecule #12: beta,beta-caroten-4-one
+Macromolecule #13: 2,3-DIMETHYL-5-(3,7,11,15,19,23,27,31,35-NONAMETHYL-2,6,10,14,18,...
+Macromolecule #14: CHLOROPHYLL A
+Macromolecule #15: HEME C
+Macromolecule #16: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #17: (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-...
+Macromolecule #18: (1S,8E)-1-{[(2S)-1-hydroxy-3-{[(1S)-1-hydroxypentadecyl]oxy}propa...
+Macromolecule #19: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #20: 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL
+Macromolecule #21: EICOSANE
+Macromolecule #22: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 70 % / Chamber temperature: 288 K / Instrument: LEICA EM GP | |||||||||||||||
| Details | Sample was monodisperse but not present in every hole |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 20133 / Average exposure time: 3.5 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID:  7ppw Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-7r0w: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)