-Search query
-Search result
Showing all 31 items for (author: sewell & bt)

EMDB-19910: 
Cyanide dihydratase from Bacillus pumilus C1 E155R variant with altered helical twist.
Method: single particle / : Dlamini LS, Woodward JD, Sewell BT

EMDB-17409: 
Cyanide dihydratase from Bacillus pumilus C1
Method: helical / : Mulelu AE, Reitz J, van Rooyen JM, Scheffer M, Frangakis AS, Dlamini LS, Woodward JD, Benedik MJ, Sewell BT

EMDB-16437: 
Cyanide dihydratase from Bacillus pumilus C1 variant - Q86R,H305K,H308K,H323K
Method: helical / : Mulelu AE, Reitz J, van Rooyen J, Scheffer M, Frangakis AS, Dlamini LS, Woodward JD, Benedik MJ, Sewell BT

EMDB-13797: 
Structure of full-length, monomeric, soluble somatic angiotensin I-converting enzyme showing the N- and C-terminal ellipsoid domains
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-13799: 
Local refinement structure of the N-domain of full-length, monomeric, soluble somatic angiotensin I-converting enzyme
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-13801: 
Local refinement structure of the C-domain of full-length, monomeric, soluble somatic angiotensin I-converting enzyme
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-13802: 
Structure of full-length, dimeric, soluble somatic angiotensin I-converting enzyme
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-13803: 
Local refinement structure of the two interacting N-domains of full-length, dimeric, soluble somatic angiotensin I-converting enzyme
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-13804: 
Local refinement structure of a single N-domain of full-length, dimeric, soluble somatic angiotensin I-converting enzyme
Method: single particle / : Lubbe L, Sewell BT, Sturrock ED

EMDB-12589: 
Plasmodium falciparum Glutamine synthetase (PfGs)
Method: single particle / : Makumire S, Mlelu AE, Gqunu S, Weber WB, Sewell BT

EMDB-11158: 
Cryo-EM structure of the nitrilase from Pseudomonas fluorescens EBC191 at 3.3 Angstroms
Method: helical / : Eppinger E, Stolz A

EMDB-3486: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M
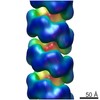
EMDB-3496: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3497: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3498: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3499: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3500: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3501: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3503: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3504: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-3505: 
Substrate specificity in plant nitrilase helical assemblies is determined by their twist.
Method: helical / : Woodward JD, Trompetter I, Sewell BT, Piotrowski M

EMDB-5412: 
CryoEM icosahedral reconstruction of AHSV7
Method: single particle / : Manole V, Laurinmaki P, Wyngaardt WV, Potgieter CA, Wright IM, Venter GJ, Dijk AAV, Sewell BT, Butcher SJ

EMDB-2075: 
CryoEM icosahedral reconstruction of AHSV7
Method: single particle / : Manole V, Laurinmaki P, Wyngaardt WV, Potgieter CA, Wright IM, Venter GJ, Dijk AAV, Sewell BT, Butcher SJ

EMDB-2076: 
CryoEM icosahedral reconstruction of AHSV4
Method: single particle / : Manole V, Laurinmaki P, Wyngaardt WV, Potgieter CA, Wright IM, Venter GJ, Dijk AAV, Sewell BT, Butcher SJ

EMDB-1870: 
Three-Dimensional Reconstruction of Heterocapsa circularisquama RNA Virus (HcRNAV-109) by Cryo-Electron Microscopy
Method: single particle / : Miller JL, Woodward JD, Chen S, Jaffer MA, Weber BW, Nagasaki K, Tomaru Y, Wepf R, Roseman A, Sewell BT, Varsani A

EMDB-1469: 
Iterative Helical Real Space Reconstruction of Histone Octamer Helical Tubes
Method: helical / : Frouws TD, Patterton HG, Sewell BT

EMDB-1313: 
Post-translational cleavage of recombinantly expressed nitrilase from Rhodococcus rhodochrous J1 yields a stable, active helical form.
Method: helical / : Thuku RN, Weber BW, Varsani A, Sewell BT

EMDB-1204: 
Three-dimensional structure of a type III glutamine synthetase by single-particle reconstruction.
Method: single particle / : van Rooyen JM, Abratt VR, Sewell BT

EMDB-1205: 
Three-dimensional structure of a type III glutamine synthetase by single-particle reconstruction.
Method: single particle / : van Rooyen JM, Abratt VR, Sewell BT

EMDB-1095: 
A mutant chaperonin with rearranged inter-ring electrostatic contacts and temperature-sensitive dissociation.
Method: single particle / : Sewell BT, Best RB, Chen S, Roseman AM, Farr GW, Horwich AR, Saibil HR

EMDB-1050: 
The cyanide degrading nitrilase from Pseudomonas stutzeri AK61 is a two-fold symmetric, 14-subunit spiral.
Method: single particle / : Sewell BT, Berman MN, Meyers PR, Jandhyala D, Benedik MJ
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
