+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Plasmodium falciparum Glutamine synthetase (PfGs) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 2.8 Å | |||||||||
 Authors Authors | Makumire S / Mlelu AE / Gqunu S / Weber WB / Sewell BT | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The structure of glutamine synthetase from Plasmodium falciparum Authors: Makumire S / Gqunu S / Mlelu AE / Weber WB / Sewell BT | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12589.map.gz emd_12589.map.gz | 28.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12589-v30.xml emd-12589-v30.xml emd-12589.xml emd-12589.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12589.png emd_12589.png | 113 KB | ||
| Masks |  emd_12589_msk_1.map emd_12589_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Others |  emd_12589_half_map_1.map.gz emd_12589_half_map_1.map.gz emd_12589_half_map_2.map.gz emd_12589_half_map_2.map.gz | 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12589 http://ftp.pdbj.org/pub/emdb/structures/EMD-12589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12589 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12589.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12589.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_12589_msk_1.map emd_12589_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
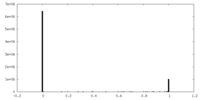
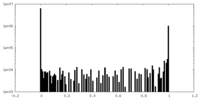
| Density Histograms |
-Half map: #2
| File | emd_12589_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
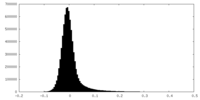
| Density Histograms |
-Half map: #1
| File | emd_12589_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
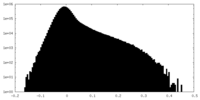
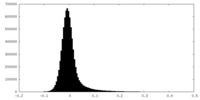
| Density Histograms |
- Sample components
Sample components
-Entire : P falciparum Glutamine synthetase
| Entire | Name: P falciparum Glutamine synthetase |
|---|---|
| Components |
|
-Supramolecule #1: P falciparum Glutamine synthetase
| Supramolecule | Name: P falciparum Glutamine synthetase / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 756 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #1: Glutamine synthetase
| Macromolecule | Name: Glutamine synthetase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MLIKLFFSNN AELYEYIKDK KNDVEIVACI ITNLLGTYFK CFFYVKEITL NKLESGFSFD ASSIKLCSD TEVSDFFIKV DHSTCYLEEC DGKNILNIMC DIKRYNGFDY YKCPRTILKK T CEFVKNEG IADKVCIGNE LEFFIFDKVN YSLDEYNTYL KVYDRESFSC ...String: MLIKLFFSNN AELYEYIKDK KNDVEIVACI ITNLLGTYFK CFFYVKEITL NKLESGFSFD ASSIKLCSD TEVSDFFIKV DHSTCYLEEC DGKNILNIMC DIKRYNGFDY YKCPRTILKK T CEFVKNEG IADKVCIGNE LEFFIFDKVN YSLDEYNTYL KVYDRESFSC KNDLSSIYGN HV VNKVEPH KDHFNNPNNE YLINDDSKKV KKKSGYFTTD PYDTSNIIKL RICRALNDMN INV QRYHHE VSTSQHEISL KYFDALTNAD FLLITKQIIK TTVSSFNRTA TFMPKPLVND NGNG LHCNI SLWKNNKNIF YHNDPSTFFL SKESFYFMYG IVKHAKALQA FCNATMNSYK RLVPG FETC QKLFYSFGSR SAVIRLSLIN YSNPSEKRIE FRLPDCANSP HLVMAAIILA GYDGIK SKE QPLVPFESKD NHFYISSIFS KYVQHPENFN ILTHALEGYE SLHTINESPE FKNFFKC EE PQGISFSLVE SLDALEKDHA FLTVNNIFTE EMIQEYIKFK REEIDAYNKY VNAYDYHL Y YEC |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Staining | Type: NEGATIVE / Material: uranyl acetate |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.34 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)