[English] 日本語
 Yorodumi
Yorodumi- EMDB-16437: Cyanide dihydratase from Bacillus pumilus C1 variant - Q86R,H305K... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cyanide dihydratase from Bacillus pumilus C1 variant - Q86R,H305K,H308K,H323K | |||||||||
 Map data Map data | Map of cyanide dihydratase from Bacillus pumilus C1 variant (Q86R/H305K/H308K/H323K) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Helical / homo-oligomeric / cyanide dihydratase / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationnitrilase activity / detoxification of nitrogen compound / nitrile hydratase activity Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Mulelu AE / Reitz J / van Rooyen J / Scheffer M / Frangakis AS / Dlamini LS / Woodward JD / Benedik MJ / Sewell BT | |||||||||
| Funding support |  South Africa, 1 items South Africa, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The Role of Histidine Residues in the Oligomerization of Cyanide Dihydratase from Bacillus pumilus C1 Authors: Mulelu AE / Reitz J / van Rooyen J / Scheffer M / Frangakis AS / Dlamini LS / Woodward JD / Benedik MJ / Sewell BT | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16437.map.gz emd_16437.map.gz | 95.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16437-v30.xml emd-16437-v30.xml emd-16437.xml emd-16437.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16437.png emd_16437.png | 39.3 KB | ||
| Filedesc metadata |  emd-16437.cif.gz emd-16437.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16437 http://ftp.pdbj.org/pub/emdb/structures/EMD-16437 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16437 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16437 | HTTPS FTP |
-Related structure data
| Related structure data |  8c5iMC  8p4iC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16437.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16437.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map of cyanide dihydratase from Bacillus pumilus C1 variant (Q86R/H305K/H308K/H323K) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
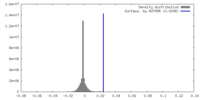
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Active helical nitrilase homo-oligomer of cyanide dihydratase fro...
| Entire | Name: Active helical nitrilase homo-oligomer of cyanide dihydratase from Bacillus pumilus C1 variant (Q86R/H305K/H308K/H323K) |
|---|---|
| Components |
|
-Supramolecule #1: Active helical nitrilase homo-oligomer of cyanide dihydratase fro...
| Supramolecule | Name: Active helical nitrilase homo-oligomer of cyanide dihydratase from Bacillus pumilus C1 variant (Q86R/H305K/H308K/H323K) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Cyanide dihydratase from Bacillus pumilus C1 variant generated by site-directed mutagenesis. |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Cyanide dihydratase
| Macromolecule | Name: Cyanide dihydratase / type: protein_or_peptide / ID: 1 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 37.503363 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTSIYPKFRA AAVQAAPIYL NLEASVEKSC ELIDEAASNG AKLVAFPEAF LPGYPWFAFI GHPEYTRKFY HELYKNAVEI PSLAIRKIS EAAKRNETYV CISCSEKDGG SLYLAQLWFN PNGDLIGKHR KMRASVAERL IWGDGSGSMM PVFQTEIGNL G GLMCWEHQ ...String: MTSIYPKFRA AAVQAAPIYL NLEASVEKSC ELIDEAASNG AKLVAFPEAF LPGYPWFAFI GHPEYTRKFY HELYKNAVEI PSLAIRKIS EAAKRNETYV CISCSEKDGG SLYLAQLWFN PNGDLIGKHR KMRASVAERL IWGDGSGSMM PVFQTEIGNL G GLMCWEHQ VPLDLMAMNA QNEQVHVASW PGYFDDEISS RYYAIATQTF VLMTSSIYTE EMKEMICLTQ EQRDYFETFK SG HTCIYGP DGEPISDMVP AETEGIAYAE IDVERVIDYK YYIDPAGHYS NQSLSMNFNQ QPTPVVKKLN KQKNEVFTYE DIQ YQKGIL EEKV UniProtKB: Cyanide dihydratase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 5.4 Component:
Details: 150 mM NaCl, 50 mM Tris-HCl pH 5.4 | |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 20.0 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: A 2.5 microlitre sample was applied onto a glow-discharged grid, blotted and plunged without incubation.. | |||||||||
| Details | Homogeneous protein sample. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number classes used: 92000 Applied symmetry - Helical parameters - Δz: 16.7 Å Applied symmetry - Helical parameters - Δ&Phi: -77 ° Applied symmetry - Helical parameters - Axial symmetry: C2 (2 fold cyclic) Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: RELION (ver. 2.1) / Number images used: 103000 |
|---|---|
| Startup model | Type of model: INSILICO MODEL / In silico model: Featureless cylinder |
| Final angle assignment | Type: NOT APPLICABLE |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8c5i: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)