-Search query
-Search result
Showing 1 - 50 of 1,281 items for (author: rai & j)

EMDB-38931: 
Cryo-EM structure of artificial protein nanocage mTIP120-Ba

EMDB-40812: 
Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 1

EMDB-40813: 
Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 2

EMDB-40814: 
Local refinement of SARS-CoV-2 (HP-GSAS-Mut7) spike NTD in complex with TXG-0078 Fab

PDB-8swh: 
Local refinement of SARS-CoV-2 (HP-GSAS-Mut7) spike NTD in complex with TXG-0078 Fab

EMDB-19426: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api137

EMDB-19427: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation I

EMDB-19428: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation II

EMDB-19429: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation III

PDB-8rpy: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api137

PDB-8rpz: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation I

PDB-8rq0: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation II

PDB-8rq2: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation III

EMDB-17296: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

PDB-8oyu: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

EMDB-18594: 
Cryo-EM structure of E. coli cytochrome bo3 quinol oxidase assembled in peptidiscs

PDB-8qqk: 
Cryo-EM structure of E. coli cytochrome bo3 quinol oxidase assembled in peptidiscs

EMDB-17125: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions

EMDB-17131: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions

PDB-8orh: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions

PDB-8ors: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions

EMDB-19735: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, helical reconstruction

EMDB-19736: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, single particle reconstruction

EMDB-19737: 
Full-length human cystathionine beta-synthase, basal state, helical reconstruction

EMDB-19738: 
Full-length human cystathionine beta-synthase, basal state, single particle reconstruction

EMDB-19739: 
Full-length human cystathionine beta-synthase, basal state, partially degraded tetramer
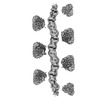
EMDB-19740: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, helical reconstruction

EMDB-19741: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, local helical reconstruction

EMDB-19742: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, local single particle reconstruction

PDB-8s5h: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, helical reconstruction

PDB-8s5i: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, single particle reconstruction

PDB-8s5j: 
Full-length human cystathionine beta-synthase, basal state, helical reconstruction

PDB-8s5k: 
Full-length human cystathionine beta-synthase, basal state, single particle reconstruction

PDB-8s5l: 
Full-length human cystathionine beta-synthase, basal state, partially degraded tetramer

PDB-8s5m: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, helical reconstruction
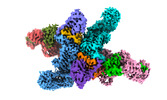
EMDB-37736: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 1)
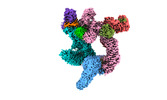
EMDB-37737: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 2)

EMDB-37739: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CDK5R1
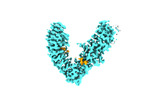
EMDB-37740: 
Local refinement of FEM1B bound with the C-degron of CCC89
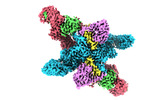
EMDB-37742: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 1)
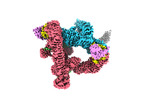
EMDB-37743: 
cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 2)
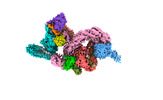
EMDB-37744: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 1)
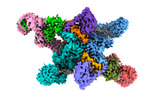
EMDB-37745: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 2)

EMDB-37746: 
Local refinement of FEM1B bound with the C-degron of CUX1

PDB-8wqa: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 1)

PDB-8wqb: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 2)

PDB-8wqc: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CDK5R1

PDB-8wqd: 
Local refinement of FEM1B bound with the C-degron of CCC89

PDB-8wqe: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 1)

PDB-8wqf: 
cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 2)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
