[English] 日本語
 Yorodumi
Yorodumi- EMDB-40812: Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 1 | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / spike / COVID / monoclonal antibody / complex / NTD / VIRAL PROTEIN | |||||||||
| Biological species |  | |||||||||
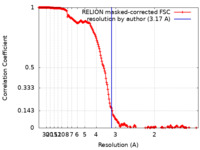
| Method | single particle reconstruction / cryo EM / Resolution: 3.17 Å | |||||||||
 Authors Authors | Bangaru S / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Deep repertoire mining uncovers ultra-broad coronavirus neutralizing antibodies targeting multiple spike epitopes. Authors: Jonathan Hurtado / Thomas F Rogers / David B Jaffe / Bruce A Adams / Sandhya Bangaru / Elijah Garcia / Tazio Capozzola / Terrence Messmer / Pragati Sharma / Ge Song / Nathan Beutler / ...Authors: Jonathan Hurtado / Thomas F Rogers / David B Jaffe / Bruce A Adams / Sandhya Bangaru / Elijah Garcia / Tazio Capozzola / Terrence Messmer / Pragati Sharma / Ge Song / Nathan Beutler / Wanting He / Katharina Dueker / Rami Musharrafieh / Sarah Burbach / Alina Truong / Michael J T Stubbington / Dennis R Burton / Raiees Andrabi / Andrew B Ward / Wyatt J McDonnell / Bryan Briney /  Abstract: The development of vaccines and therapeutics that are broadly effective against known and emergent coronaviruses is an urgent priority. We screened the circulating B cell repertoires of COVID-19 ...The development of vaccines and therapeutics that are broadly effective against known and emergent coronaviruses is an urgent priority. We screened the circulating B cell repertoires of COVID-19 survivors and vaccinees to isolate over 9,000 severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-specific monoclonal antibodies (mAbs), providing an expansive view of the SARS-CoV-2-specific Ab repertoire. Among the recovered antibodies was TXG-0078, an N-terminal domain (NTD)-specific neutralizing mAb that recognizes diverse alpha- and beta-coronaviruses. TXG-0078 achieves its exceptional binding breadth while utilizing the same VH1-24 variable gene signature and heavy-chain-dominant binding pattern seen in other NTD-supersite-specific neutralizing Abs with much narrower specificity. We also report CC24.2, a pan-sarbecovirus neutralizing antibody that targets a unique receptor-binding domain (RBD) epitope and shows similar neutralization potency against all tested SARS-CoV-2 variants, including BQ.1.1 and XBB.1.5. A cocktail of TXG-0078 and CC24.2 shows protection in vivo, suggesting their potential use in variant-resistant therapeutic Ab cocktails and as templates for pan-coronavirus vaccine design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40812.map.gz emd_40812.map.gz | 502.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40812-v30.xml emd-40812-v30.xml emd-40812.xml emd-40812.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40812_fsc.xml emd_40812_fsc.xml | 18.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_40812.png emd_40812.png | 73.5 KB | ||
| Masks |  emd_40812_msk_1.map emd_40812_msk_1.map | 536.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40812.cif.gz emd-40812.cif.gz | 6.4 KB | ||
| Others |  emd_40812_half_map_1.map.gz emd_40812_half_map_1.map.gz emd_40812_half_map_2.map.gz emd_40812_half_map_2.map.gz | 430.9 MB 430.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40812 http://ftp.pdbj.org/pub/emdb/structures/EMD-40812 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40812 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40812 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40812.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40812.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.725 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40812_msk_1.map emd_40812_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
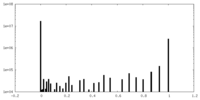
| Density Histograms |
-Half map: Half-map 2
| File | emd_40812_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
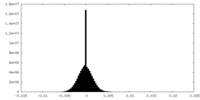
| Density Histograms |
-Half map: Half-map 1
| File | emd_40812_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
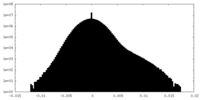
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-...
| Entire | Name: Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 1 |
|---|---|
| Components |
|
-Supramolecule #1: Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-...
| Supramolecule | Name: Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: SARS-CoV-2 spike (HP-GSAS-Mut7)
| Macromolecule | Name: SARS-CoV-2 spike (HP-GSAS-Mut7) / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF LGVYYHKNNK SWMESEFRVY SSANNCTFEY ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF LGVYYHKNNK SWMESEFRVY SSANNCTFEY VSQPFLMDLE GKQGNFKNLR EFVFKNIDGY FKIYSKHTPI NLVRDLPQGF SALEPLVDLP IGINITRFQT LLALHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIYQTSNFRV QPTESIVRFP NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADSF VIRGDEVRQI APGQTGKIAD YNYKLPDDFT GCVIAWNSNN LDSKVGGNYN YLYRLFRKSN LKPFERDIST EIYQAGSTPC NGVEGFNCYF PLQSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCGPKKS TNLVKNKCVN FNFNGLTGTG VLTESNKKFL PFQQFGRDIA DTTDAVRDPQ TLEILDITPC SFGGVSVITP GTNTSNQVAV LYQDVNCTEV PVAIHADQLT PTWRVYSTGS NVFQTRAGCL IGAEHVNNSY ECDIPIGAGI CASYQTQTNS PGSASSVASQ SIIAYTMSLG AENSCAYSNN SIAIPTNFTI SVTTEILPVS MTKTSVDCTM YICGDSTECS NLLLQYGSFC TQLNRALTGI AVEQDKNTQE VFAQVKQIYK TPPIKDFGGF NFSQILPDPS KPSKRSPIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TICSGWTFGA GPALQIPFPM QMAYRFNGIG VTQNVLYENQ KLIANQFNSA IGKIQDSLSS TPSALGKLQD VVNQNAQALN TLVKQLSSNF GAISSVLNDI LSRLDPPEAE VQIDRLITGR LQSLQTYVTQ QLIRAAEIRA SANLAATKMS ECVLGQSKRV DFCGKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVSGNCDVVI GIVNNTVYDP LQPELDSFKE ELDKYFKNHT SPDVDLGDIS GINASVVNIQ KEIDRLNEVA KNLNESLIDL QELGKYEQGS GYIPEAPRDG QAYVRKDGEW VLLSTFLGRS LEVLFQGPGS AWSHPQFEKG GGSGGGGSGG SAWSHPQFEK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: 1X TBS | |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 10 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | The spike protein was incubated with a 3-fold molar excess of TXG-0078 Fab for 30 minutes and mixed with 0.04 mM Lauryl maltose neopentyl glycol solution immediately prior to sample deposition. |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4257 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 190000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)