+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of artificial protein nanocage mTIP120-Ba | |||||||||||||||||||||||||||
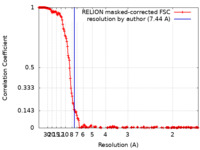
 Map data Map data | main map, job066, postprocess, C1, 7.44A resolution | |||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||
 Keywords Keywords | Artificial designed protein complex / Protein cage / Protein nanoparticle / Metal-induced protein assembly / Protein metal complex / DE NOVO PROTEIN | |||||||||||||||||||||||||||
| Biological species | synthetic construct (others) | |||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.44 Å | |||||||||||||||||||||||||||
 Authors Authors | Ohara N / Kawakami N / Arai R / Adachi N / Ikeda A / Senda T / Miyamoto K | |||||||||||||||||||||||||||
| Funding support |  Japan, 8 items Japan, 8 items
| |||||||||||||||||||||||||||
 Citation Citation | Journal: Angew Chem Int Ed Engl / Year: 2018 Title: Design of Hollow Protein Nanoparticles with Modifiable Interior and Exterior Surfaces. Authors: Norifumi Kawakami / Hiroki Kondo / Yuki Matsuzawa / Kaoru Hayasaka / Erika Nasu / Kenji Sasahara / Ryoichi Arai / Kenji Miyamoto /  Abstract: Protein-based nanoparticles hold promise for a broad range of applications. Here, we report the production of a uniform anionic hollow protein nanoparticle, designated TIP60, which spontaneously ...Protein-based nanoparticles hold promise for a broad range of applications. Here, we report the production of a uniform anionic hollow protein nanoparticle, designated TIP60, which spontaneously assembles from a designed fusion protein subunit based on the geometric features of polyhedra. We show that TIP60 tolerates mutation and both its interior and exterior surfaces can be chemically modified. Moreover, TIP60 forms larger structures upon the addition of a cationic protein. Therefore, TIP60 can be used as a modifiable nano-building block for further molecular assembly. | |||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38931.map.gz emd_38931.map.gz | 561.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38931-v30.xml emd-38931-v30.xml emd-38931.xml emd-38931.xml | 28.3 KB 28.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38931_fsc.xml emd_38931_fsc.xml | 19.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_38931.png emd_38931.png | 109.7 KB | ||
| Masks |  emd_38931_msk_1.map emd_38931_msk_1.map | 600.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-38931.cif.gz emd-38931.cif.gz | 6.7 KB | ||
| Others |  emd_38931_additional_1.map.gz emd_38931_additional_1.map.gz emd_38931_half_map_1.map.gz emd_38931_half_map_1.map.gz emd_38931_half_map_2.map.gz emd_38931_half_map_2.map.gz | 485.1 MB 487.8 MB 487.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38931 http://ftp.pdbj.org/pub/emdb/structures/EMD-38931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38931 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38931.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38931.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map, job066, postprocess, C1, 7.44A resolution | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_38931_msk_1.map emd_38931_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: full map, job053, refine3d, C1, 8.40A resolution
| File | emd_38931_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | full map, job053, refine3d, C1, 8.40A resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2, job053, refine3d, C1, 8.40A resolution
| File | emd_38931_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2, job053, refine3d, C1, 8.40A resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1, job053, refine3d, C1, 8.40A resolution
| File | emd_38931_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1, job053, refine3d, C1, 8.40A resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : artificial protein cage, mTIP120-Ba
| Entire | Name: artificial protein cage, mTIP120-Ba |
|---|---|
| Components |
|
-Supramolecule #1: artificial protein cage, mTIP120-Ba
| Supramolecule | Name: artificial protein cage, mTIP120-Ba / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: metal-ion induced TIP120 (truncated icosahedral protein composed of 120-mer fusion proteins) complexed with barium ions |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 1.074 MDa |
-Macromolecule #1: CoreN domain of artificial protein nanocage, mTIP120-Ba
| Macromolecule | Name: CoreN domain of artificial protein nanocage, mTIP120-Ba type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SQDPKNIKIM RLVTGEDIIG NISESQGLIT IKKAFVIIPM QATP |
-Macromolecule #2: CoreC domain of artificial protein nanocage, mTIP120-Ba
| Macromolecule | Name: CoreC domain of artificial protein nanocage, mTIP120-Ba type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SQDPLEVLFQ GPGKPVQLVL SPWQPYTDDK EIVIDDSKVI TITSPKDDII KSYESHTRVL ENKQVEEILR LEKEIEDLQR MKEQQELSLT EASLQKLQER RDQELRRLEE E |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: The grid was washed by acetone prior to use. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291 K / Instrument: FEI VITROBOT MARK IV / Details: Blotting time was 5 seconds (blot force 15). |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 6191 / Average exposure time: 4.66 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 120000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 582 / Target criteria: Correlation coefficient |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)