[English] 日本語
 Yorodumi
Yorodumi- EMDB-19741: Full-length human cystathionine beta-synthase with C-terminal 6xH... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, local helical reconstruction | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Filament / Allostery / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationCysteine formation from homocysteine / L-homocysteine catabolic process / modified amino acid binding / cystathionine beta-synthase / cystathionine beta-synthase activity / L-serine catabolic process / : / Metabolism of ingested SeMet, Sec, MeSec into H2Se / : / carbon monoxide binding ...Cysteine formation from homocysteine / L-homocysteine catabolic process / modified amino acid binding / cystathionine beta-synthase / cystathionine beta-synthase activity / L-serine catabolic process / : / Metabolism of ingested SeMet, Sec, MeSec into H2Se / : / carbon monoxide binding / cartilage development involved in endochondral bone morphogenesis / hydrogen sulfide biosynthetic process / homocysteine metabolic process / cerebellum morphogenesis / L-cysteine catabolic process / L-serine metabolic process / L-cysteine biosynthetic process / endochondral ossification / response to folic acid / transsulfuration / nitric oxide binding / : / DNA protection / S-adenosyl-L-methionine binding / nitrite reductase (NO-forming) activity / regulation of JNK cascade / superoxide metabolic process / maternal process involved in female pregnancy / blood vessel remodeling / blood vessel diameter maintenance / oxygen binding / pyridoxal phosphate binding / cellular response to hypoxia / heme binding / ubiquitin protein ligase binding / negative regulation of apoptotic process / enzyme binding / protein homodimerization activity / metal ion binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
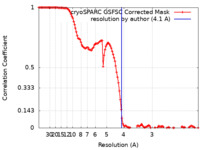
| Method | helical reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | McCorvie TJ / Yue WW | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Architecture and regulation of filamentous human cystathionine beta-synthase. Authors: Thomas J McCorvie / Douglas Adamoski / Raquel A C Machado / Jiazhi Tang / Henry J Bailey / Douglas S M Ferreira / Claire Strain-Damerell / Arnaud Baslé / Andre L B Ambrosio / Sandra M G Dias / Wyatt W Yue /    Abstract: Cystathionine beta-synthase (CBS) is an essential metabolic enzyme across all domains of life for the production of glutathione, cysteine, and hydrogen sulfide. Appended to the conserved catalytic ...Cystathionine beta-synthase (CBS) is an essential metabolic enzyme across all domains of life for the production of glutathione, cysteine, and hydrogen sulfide. Appended to the conserved catalytic domain of human CBS is a regulatory domain that modulates activity by S-adenosyl-L-methionine (SAM) and promotes oligomerisation. Here we show using cryo-electron microscopy that full-length human CBS in the basal and SAM-bound activated states polymerises as filaments mediated by a conserved regulatory domain loop. In the basal state, CBS regulatory domains sterically block the catalytic domain active site, resulting in a low-activity filament with three CBS dimers per turn. This steric block is removed when in the activated state, one SAM molecule binds to the regulatory domain, forming a high-activity filament with two CBS dimers per turn. These large conformational changes result in a central filament of SAM-stabilised regulatory domains at the core, decorated with highly flexible catalytic domains. Polymerisation stabilises CBS and reduces thermal denaturation. In PC-3 cells, we observed nutrient-responsive CBS filamentation that disassembles when methionine is depleted and reversed in the presence of SAM. Together our findings extend our understanding of CBS enzyme regulation, and open new avenues for investigating the pathogenic mechanism and therapeutic opportunities for CBS-associated disorders. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19741.map.gz emd_19741.map.gz | 91.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19741-v30.xml emd-19741-v30.xml emd-19741.xml emd-19741.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19741_fsc.xml emd_19741_fsc.xml | 12 KB | Display |  FSC data file FSC data file |
| Images |  emd_19741.png emd_19741.png | 37.5 KB | ||
| Masks |  emd_19741_msk_1.map emd_19741_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19741.cif.gz emd-19741.cif.gz | 5.9 KB | ||
| Others |  emd_19741_half_map_1.map.gz emd_19741_half_map_1.map.gz emd_19741_half_map_2.map.gz emd_19741_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19741 http://ftp.pdbj.org/pub/emdb/structures/EMD-19741 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19741 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19741 | HTTPS FTP |
-Related structure data
| Related structure data |  8s5hC  8s5iC  8s5jC  8s5kC  8s5lC  8s5mC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19741.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19741.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19741_msk_1.map emd_19741_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
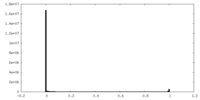
| Density Histograms |
-Half map: #1
| File | emd_19741_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19741_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Helical assembly of full-length human cystathionine beta-synthase...
| Entire | Name: Helical assembly of full-length human cystathionine beta-synthase bound to SAM in the activated state |
|---|---|
| Components |
|
-Supramolecule #1: Helical assembly of full-length human cystathionine beta-synthase...
| Supramolecule | Name: Helical assembly of full-length human cystathionine beta-synthase bound to SAM in the activated state type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: cystathionine beta-synthase
| Macromolecule | Name: cystathionine beta-synthase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: cystathionine beta-synthase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MPSETPQAEV GPTGCPHRSG PHSAKGSLEK GSPEDKEAKE PLWIRPDAPS RCTWQLGRPA SESPHHHTA PAKSPKILPD ILKKIGDTPM VRINKIGKKF GLKCELLAKC EFFNAGGSVK D RISLRMIE DAERDGTLKP GDTIIEPTSG NTGIGLALAA AVRGYRCIIV ...String: MPSETPQAEV GPTGCPHRSG PHSAKGSLEK GSPEDKEAKE PLWIRPDAPS RCTWQLGRPA SESPHHHTA PAKSPKILPD ILKKIGDTPM VRINKIGKKF GLKCELLAKC EFFNAGGSVK D RISLRMIE DAERDGTLKP GDTIIEPTSG NTGIGLALAA AVRGYRCIIV MPEKMSSEKV DV LRALGAE IVRTPTNARF DSPESHVGVA WRLKNEIPNS HILDQYRNAS NPLAHYDTTA DEI LQQCDG KLDMLVASVG TGGTITGIAR KLKEKCPGCR IIGVDPEGSI LAEPEELNQT EQTT YEVEG IGYDFIPTVL DRTVVDKWFK SNDEEAFTFA RMLIAQEGLL CGGSAGSTVA VAVKA AQEL QEGQRCVVIL PDSVRNYMTK FLSDRWMLQK GFLKEEDLTE KKPWWWHLRV QELGLS APL TVLPTITCGH TIEILREKGF DQAPVVDEAG VILGMVTLGN MLSSLLAGKV QPSDQVG KV IYKQFKQIRL TDTLGRLSHI LEMDHFALVV HEQIQYHSTG KSSQRQMVFG VVTAIDLL N FVAAQERDQK AAHHHHHH UniProtKB: Cystathionine beta-synthase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: filter sterile 25 mM HEPES, pH 7.5, 200 mM NaCl, 2.0 mM TCEP, 0.005% (v/v) tween-20, 5 mM SAM | ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 11220 / Average exposure time: 3.53 sec. / Average electron dose: 39.96 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)