[English] 日本語
 Yorodumi
Yorodumi- EMDB-19428: Escherichia coli 50S subunit in complex with the antimicrobial pe... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
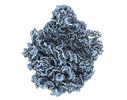
| Title | Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation II | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | ribosome / antimicrobial peptide / RNA / ribosomal protein / PrAMP / proline-rich peptide / antibiotics / 50S / Api88 | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of cytoplasmic translational initiation / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity ...negative regulation of cytoplasmic translational initiation / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / mRNA 5'-UTR binding / large ribosomal subunit / transferase activity / ribosome binding / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / mRNA binding / DNA binding / RNA binding / zinc ion binding / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |   | ||||||||||||||||||
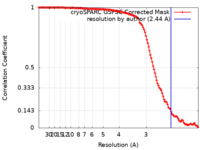
| Method | single particle reconstruction / cryo EM / Resolution: 2.44 Å | ||||||||||||||||||
 Authors Authors | Lauer S / Nikolay R / Spahn C | ||||||||||||||||||
| Funding support |  Germany, 5 items Germany, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Multimodal binding and inhibition of bacterial ribosomes by the antimicrobial peptides Api137 and Api88. Authors: Simon M Lauer / Maren Reepmeyer / Ole Berendes / Dorota Klepacki / Jakob Gasse / Sara Gabrielli / Helmut Grubmüller / Lars V Bock / Andor Krizsan / Rainer Nikolay / Christian M T Spahn / Ralf Hoffmann /   Abstract: Proline-rich antimicrobial peptides (PrAMPs) inhibit bacterial protein biosynthesis by binding to the polypeptide exit tunnel (PET) near the peptidyl transferase center. Api137, an optimized ...Proline-rich antimicrobial peptides (PrAMPs) inhibit bacterial protein biosynthesis by binding to the polypeptide exit tunnel (PET) near the peptidyl transferase center. Api137, an optimized derivative of honeybee PrAMP apidaecin, inhibits protein expression by trapping release factors (RFs), which interact with stop codons on ribosomes to terminate translation. This study uses cryo-EM, functional assays and molecular dynamic (MD) simulations to show that Api137 additionally occupies a second binding site near the exit of the PET and can repress translation independently of RF-trapping. Api88, a C-terminally amidated (-CONH) analog of Api137 (-COOH), binds to the same sites, occupies a third binding pocket and interferes with the translation process presumably without RF-trapping. In conclusion, apidaecin-derived PrAMPs inhibit bacterial ribosomes by multimodal mechanisms caused by minor structural changes and thus represent a promising pool for drug development efforts. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19428.map.gz emd_19428.map.gz | 89.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19428-v30.xml emd-19428-v30.xml emd-19428.xml emd-19428.xml | 52.1 KB 52.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19428_fsc.xml emd_19428_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_19428.png emd_19428.png | 121.7 KB | ||
| Masks |  emd_19428_msk_1.map emd_19428_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19428.cif.gz emd-19428.cif.gz | 10.3 KB | ||
| Others |  emd_19428_half_map_1.map.gz emd_19428_half_map_1.map.gz emd_19428_half_map_2.map.gz emd_19428_half_map_2.map.gz | 165 MB 165 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19428 http://ftp.pdbj.org/pub/emdb/structures/EMD-19428 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19428 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19428 | HTTPS FTP |
-Related structure data
| Related structure data |  8rq0MC  8rpyC  8rpzC  8rq2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19428.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19428.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.998 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19428_msk_1.map emd_19428_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19428_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19428_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Api88 in complex with the large ribosomal subunit - conformation II
+Supramolecule #1: Api88 in complex with the large ribosomal subunit - conformation II
+Supramolecule #2: Large ribosomal subunit
+Supramolecule #3: Api88
+Macromolecule #1: Large ribosomal subunit protein bL32
+Macromolecule #2: Large ribosomal subunit protein bL33
+Macromolecule #3: Large ribosomal subunit protein bL34
+Macromolecule #4: Large ribosomal subunit protein bL35
+Macromolecule #5: Large ribosomal subunit protein bL36A
+Macromolecule #6: Large ribosomal subunit protein bL31
+Macromolecule #9: Large ribosomal subunit protein uL2
+Macromolecule #10: Large ribosomal subunit protein uL3
+Macromolecule #11: Large ribosomal subunit protein uL4
+Macromolecule #12: Large ribosomal subunit protein uL5
+Macromolecule #13: Large ribosomal subunit protein uL6
+Macromolecule #14: Large ribosomal subunit protein bL9
+Macromolecule #15: Large ribosomal subunit protein uL13
+Macromolecule #16: Large ribosomal subunit protein uL14
+Macromolecule #17: Large ribosomal subunit protein uL15
+Macromolecule #18: Large ribosomal subunit protein uL16
+Macromolecule #19: Large ribosomal subunit protein bL17
+Macromolecule #20: Large ribosomal subunit protein uL18
+Macromolecule #21: Large ribosomal subunit protein bL19
+Macromolecule #22: Large ribosomal subunit protein bL20
+Macromolecule #23: Large ribosomal subunit protein bL21
+Macromolecule #24: Large ribosomal subunit protein uL22
+Macromolecule #25: Large ribosomal subunit protein uL23
+Macromolecule #26: Large ribosomal subunit protein uL24
+Macromolecule #27: Large ribosomal subunit protein bL25
+Macromolecule #28: Large ribosomal subunit protein bL27
+Macromolecule #29: Large ribosomal subunit protein bL28
+Macromolecule #30: Large ribosomal subunit protein uL29
+Macromolecule #31: Large ribosomal subunit protein uL30
+Macromolecule #32: Apidaecins type 88
+Macromolecule #7: 23S ribosomal RNA
+Macromolecule #8: 5S ribosomal RNA
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 26.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8rq0: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)