-Search query
-Search result
Showing 1 - 50 of 177 items for (author: nicholas & ci)
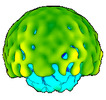
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry

EMDB-19075: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K composite map

EMDB-19282: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K Cas12k domain local-refinement map

EMDB-19283: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K TnsC domain local-refinement map

EMDB-19284: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K TnsB domain local-refinement map

EMDB-19286: 
consensus map of the V-K CRISPR-associated Transposon Integration Assembly

PDB-8rdu: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K composite map

PDB-8rkt: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K Cas12k domain local-refinement map

PDB-8rku: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K TnsC domain local-refinement map

PDB-8rkv: 
Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K TnsB domain local-refinement map

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
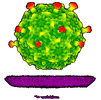
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
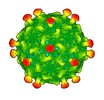
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging

EMDB-15697: 
Cryo-EM structure of shCas12k-sgRNA-dsDNA ternary complex (type V-K CRISPR-associated transposon)

PDB-8axa: 
Cryo-EM structure of shCas12k-sgRNA-dsDNA ternary complex (type V-K CRISPR-associated transposon)

EMDB-29530: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-29531: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-40240: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8fxb: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8fxc: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8s9g: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-41140: 
APC/C-CDH1-UBE2C-Ubiquitin-CyclinB-NTD

EMDB-41142: 
APC/C-CDH1-UBE2C-UBE2S-Ubiquitin-CyclinB

PDB-8tar: 
APC/C-CDH1-UBE2C-Ubiquitin-CyclinB-NTD

PDB-8tau: 
APC/C-CDH1-UBE2C-UBE2S-Ubiquitin-CyclinB
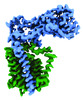
EMDB-41299: 
Structural basis of peptidoglycan synthesis by E. coli RodA-PBP2 complex
EMDB-41303: 
Transmembrane map
EMDB-41304: 
Periplasmic map

PDB-8tj3: 
Structural basis of peptidoglycan synthesis by E. coli RodA-PBP2 complex

EMDB-16087: 
Structure of the human UBR5 Dimer.

EMDB-14778: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the half-closed conformation

EMDB-14801: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C2 symmetry

EMDB-14852: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C1 symmetry

EMDB-14960: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the open conformation

PDB-7zla: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the half-closed conformation

PDB-7zn5: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C2 symmetry.

PDB-7zpa: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C1 symmetry

PDB-7zth: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the open conformation

EMDB-28999: 
Engineered human dynein motor domain in microtubule-bound state

EMDB-29003: 
Engineered human dynein motor domain in the microtubule-unbound state in the buffer containing ATP-Vi

EMDB-29012: 
Engineered human dynein motor domain in the microtubule-unbound state with LIS1 complex in the buffer containing ATP-Vi

EMDB-29014: 
Engineered human dynein motor domain in the microtubule-unbound state with LIS1 complex in the buffer containing ATP-Vi (local refined on AAA3-AAA5 and LIS1)

PDB-8fcy: 
Engineered human dynein motor domain in microtubule-bound state

PDB-8fd6: 
Engineered human dynein motor domain in the microtubule-unbound state in the buffer containing ATP-Vi

PDB-8fdt: 
Engineered human dynein motor domain in the microtubule-unbound state with LIS1 complex in the buffer containing ATP-Vi

PDB-8fdu: 
Engineered human dynein motor domain in the microtubule-unbound state with LIS1 complex in the buffer containing ATP-Vi (local refined on AAA3-AAA5 and LIS1)

EMDB-15831: 
CryoEM structure of the pointy tip (proteins pIII/pVI/pVIII) from the f1 filamentous bacteriophage

EMDB-15832: 
CryoEM structure of the round tip (proteins pVII/pVIII/pIX) from the f1 filamentous bacteriophage
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
