-Search query
-Search result
Showing all 45 items for (author: miller & dj)

EMDB-46692: 
S. thermophilus class III ribonucleotide reductase signal subtracted cone domains and core
Method: single particle / : Andree GA, Drennan CL

EMDB-46693: 
S. thermophilus class III ribonucleotide reductase focused refined core
Method: single particle / : Andree GA, Drennan CL

EMDB-46696: 
S. thermophilus class III ribonucleotide reductase consensus
Method: single particle / : Andree GA, Drennan CL

EMDB-46698: 
S. thermophilus class III ribonucleotide reductase with dATP and TTP
Method: single particle / : Andree GA, Drennan CL

EMDB-46712: 
S. thermophilus class III ribonucleotide reductase signal subtracted cone domains and core
Method: single particle / : Andree GA, Drennan CL

EMDB-46713: 
S. thermophilus class III ribonucleotide reductase focused refined core
Method: single particle / : Andree GA, Drennan CL

EMDB-46746: 
S. thermophilus class III ribonucleotide reductase consensus
Method: single particle / : Andree GA, Drennan CL

EMDB-46747: 
S. thermophilus class III ribonucleotide reductase with ATP and TTP
Method: single particle / : Andree GA, Drennan CL

EMDB-47447: 
Glucagon Like Peptide Receptor-1 (GLP1R) A316T mutant with GLP-1 peptide. Dominant negative Gs complex.
Method: single particle / : Deane-Alder K, Belousoff MJ, Wootten DL

EMDB-47927: 
CryoEM Structure Of Respiratory Syncytial Virus Polymerase in complex with Novel Non-Nucleoside Inhibitor Compound 16
Method: single particle / : Yin Y, Tran MT, Yu X, Jonckers T, Carney C

EMDB-47931: 
CryoEM map of Respiratory Syncytial Virus Polymerase with Non-Nucleoside Inhibitor compound 21
Method: single particle / : Yin Y, Tran MT, Yu X, Jonckers T, Carney S

EMDB-46644: 
Cryo-EM structure of a trapped ARIH1-diUB-CRL2-KLHDC10 complex - consensus map
Method: single particle / : Chittori S, Scott DC, Schulman BA

EMDB-46645: 
Focused map of Cryo-EM structure of Ubiquitin C-degron bound to KLHDC10-EloB/C
Method: single particle / : Chittori S, Scott DC, Schulman BA

EMDB-40180: 
MsbA bound to cerastecin C
Method: single particle / : Chen Y, Klein D

EMDB-41097: 
cryoEM structure of Smc5/6 5mer
Method: single particle / : Yu Y, Patel DJ

EMDB-41098: 
Smc5/6 8mer
Method: single particle / : Yu Y, Patel DJ

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
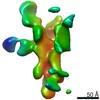
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
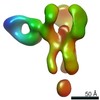
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
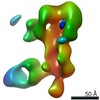
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
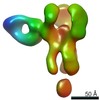
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25646: 
Negative stain map of polyclonal Fab 241.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25655: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB

EMDB-24095: 
Anaplastic lymphoma kinase (ALK) extracellular fragment of ligand binding region 673-1025 in complex with AUG-alpha
Method: single particle / : Myasnikov AG, Reshetnyak AV, Kalodimos CG

EMDB-23792: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A
EMDB-23793: 
Negative stain map of monoclonal Fab SFV009 2G01 binding the RBS of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23794: 
Negative stain map of monoclonal Fab 045-09 2B05 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23795: 
Negative stain map of monoclonal Fab SFV019 2A06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
EMDB-23796: 
Negative stain map of monoclonal Fab SFV015 2F02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23797: 
Negative stain map of monoclonal Fab 047-09 4G02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
EMDB-23798: 
Negative stain map of monoclonal Fab 047-09 4B06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-10072: 
cryo EM map of human APC/CCDH1 bound to the avid UbVW_dim trap
Method: single particle / : Watson ER, Prabu JR, Schulman BA

EMDB-8978: 
Cryo-EM structure of the active, Gs-protein complexed, human CGRP receptor
Method: single particle / : Liang YL, Khoshouei M

PDB-6e3y: 
Cryo-EM structure of the active, Gs-protein complexed, human CGRP receptor
Method: single particle / : Liang YL, Khoshouei M, Deganutti G, Glukhova A, Koole C, Peat TS, Radjainia M, Plitzko JM, Baumeister W, Miller LJ, Hay DL, Christopoulos A, Reynolds CA, Wootten D, Sexton PM

EMDB-7039: 
3.3 angstrom phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex.
Method: single particle / : Liang YL, Khoshouei M

PDB-6b3j: 
3.3 angstrom phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex
Method: single particle / : Liang YL, Khoshouei M, Glukhova A, Furness SGB, Koole C, Zhao P, Clydesdale L, Thal DM, Radjainia M, Danev R, Baumeister W, Wang MW, Miller LJ, Christopoulos A, Sexton PM, Wootten D

EMDB-8623: 
Volta phase plate cryo-electron microscopy structure of a calcitonin receptor-heterotrimeric Gs protein complex
Method: single particle / : Liang YL, Khoshouei M

PDB-5uz7: 
Volta phase plate cryo-electron microscopy structure of a calcitonin receptor-heterotrimeric Gs protein complex
Method: single particle / : Liang YL, Khoshouei M, Radjainia M, Zhang Y, Glukhova A, Tarrasch J, Thal DM, Furness SGB, Christopoulos G, Coudrat T, Danev R, Baumeister W, Miller LJ, Christopoulos A, Kobilka BK, Wootten D, Skiniotis G, Sexton PM
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
