-検索条件
-検索結果
検索 (著者・登録者: manajit & hayer-hartl)の結果全39件を表示しています

EMDB-12730: 
SSUL-gCAL mediating Synechococcus elongatus PCC 7942 M58 homo-demixing

EMDB-12731: 
Structure of the repeat unit in the network formed by CcmM full length isoform and Rubisco from Synechococcus elongatus

EMDB-12732: 
CryoEM structure of the interaction between CcmM full length isoform (SSUL) domain with RbcL8 core from Synechococcus elongatus PCC 7942

EMDB-11028: 
CryoEM structure of Rubisco Activase with its substrate Rubisco from Nostoc sp. (strain PCC7120)

EMDB-11029: 
CryoEM structure of the interaction between Rubisco Activase small-subunit-like (SSUL) domain with Rubisco from Nostoc sp. (strain PCC7120)

EMDB-11575: 
CryoEM Local map of Rubisco Activase from the complex with its substrate Rubisco from Nostoc sp. (strain PCC7120)

PDB-6z1f: 
CryoEM structure of Rubisco Activase with its substrate Rubisco from Nostoc sp. (strain PCC7120)

PDB-6z1g: 
CryoEM structure of the interaction between Rubisco Activase small-subunit-like (SSUL) domain with Rubisco from Nostoc sp. (strain PCC7120)

EMDB-10528: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in the apo state

EMDB-10529: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ADP

EMDB-10530: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ATPgammaS

PDB-6tmv: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in the apo state

PDB-6tmw: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ADP

PDB-6tmx: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ATPgammaS

EMDB-0149: 
PTC3 holotoxin complex from Photorhabdus luminecens in prepore state (TcdA1, TcdB2, TccC3)

EMDB-0150: 
PTC3 holotoxin complex from Photorhabdus luminiscens - Mutant TcC-D651A

PDB-6h6e: 
PTC3 holotoxin complex from Photorhabdus luminecens in prepore state (TcdA1, TcdB2, TccC3)

PDB-6h6f: 
PTC3 holotoxin complex from Photorhabdus luminiscens - Mutant TcC-D651A

EMDB-0015: 
Symmetry-free cryo-EM map of GroEL-actin

EMDB-0016: 
Symmetry-free cryo-EM map of TRiC in apo state (nucleotide free)

EMDB-0017: 
Symmetry-free cryo-EM map of TRiC-actin

EMDB-0018: 
Symmetry-free cryo-EM map of TRiC-actin-alpha_CCT1

EMDB-0022: 
Symmetry-free cryo-EM map of TRiC-ADP-BeFx

EMDB-3699: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP

EMDB-3700: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP

EMDB-3701: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP and RcaCC

EMDB-3702: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP and RcaCC

PDB-5nv3: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP
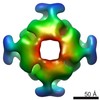
EMDB-3051: 
Structure and mechanism of the Rubisco assembly chaperone Raf1
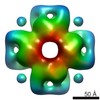
EMDB-3052: 
Structure and mechanism of the Rubisco assembly chaperone Raf1

EMDB-3053: 
Structure and mechanism of the Rubisco assembly chaperone Raf1

EMDB-1940: 
Negative stain EM density of green-type rubisco activase (R294V) from tobacco

PDB-3zw6: 
MODEL OF HEXAMERIC AAA DOMAIN ARRANGEMENT OF GREEN-TYPE RUBISCO ACTIVASE FROM TOBACCO.

PDB-3zuh: 
Negative stain EM Map of the AAA protein CbbX, a red-type Rubisco activase from R. sphaeroides

EMDB-1932: 
Negative stain EM Map of the AAA protein CbbX, a red-type Rubisco activase from R. sphaeroides

EMDB-1654: 
Rubisco RbcL8-RbcX2-8 complex

EMDB-1655: 
Coupled chaperone action in folding and assembly of hexadecameric Rubisco

EMDB-1656: 
Control structure of the RbcL8 octamer

PDB-2wvw: 
Cryo-EM structure of the RbcL-RbcX complex
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
