-Search query
-Search result
Showing 1 - 50 of 167 items for (author: lewis & c)
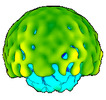
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry

EMDB-19568: 
DtpB hexamer from Streptomyces lividans

PDB-8rwy: 
DtpB hexamer from Streptomyces lividans

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
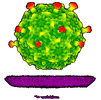
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
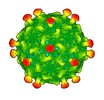
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging

PDB-9ffg: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

PDB-9ffh: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
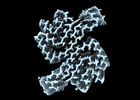
EMDB-19986: 
Structure of recombinant alpha-synuclein fibrils 1B capable of seeding GCIs in vivo

PDB-9euu: 
Structure of recombinant alpha-synuclein fibrils 1B capable of seeding GCIs in vivo

EMDB-17449: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

EMDB-17458: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

EMDB-17459: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-17460: 
S. cerevisiae sCMGE with N-ter Mcm10 density

PDB-8p5e: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

PDB-8p62: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

PDB-8p63: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8qox: 
Two-component assembly of SlaA and SlaB S-layer proteins of Sulfolobus acidocaldarius

PDB-8qp0: 
A hexamer pore in the S-layer of Sulfolobus acidocaldarius formed by SlaA protein

EMDB-18127: 
S-layer of archaeon Sulfolobus acidocaldarius by subtomogram averaging

EMDB-41810: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-41820: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41823: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41838: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to partially open HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

PDB-8u1d: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-15530: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 10.0

EMDB-15531: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 7.0

PDB-8an2: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 10.0

PDB-8an3: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 7.0

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

EMDB-41182: 
Cryo-EM map of the Unmodified nucleosome core particle in 100 mM KCl with local resolution values
EMDB-41183: 
Cryo-EM map of the PARylated nucleosome core particle in 100 mM KCl with local resolution values
EMDB-41184: 
Cryo-EM map of the Unmodified nucleosome core particle in 5 mM KCl with local resolution values
EMDB-41178: 
Cryo-EM map of the PARylated nucleosome core particle in 5 mM KCl with local resolution values

EMDB-16519: 
Slipper limpet hemocyanin didecamer

EMDB-16523: 
Slipper limpet hemocyanin tridecamer

EMDB-14635: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 4.0

PDB-7zcx: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 4.0
EMDB-29101: 
EGFR:Degrader:VHL:Elongin-B/C:Cul2
PDB-7t3h: 
MicroED structure of Dynobactin

EMDB-14242: 
E. coli BAM complex (BamABCDE) bound to dynobactin A

PDB-7r1w: 
E. coli BAM complex (BamABCDE) bound to dynobactin A

EMDB-13164: 
Cryo-EM structure of encapsulin from Mycobacterium tuberculosis

PDB-7p1t: 
Cryo-EM structure of encapsulin from Mycobacterium tuberculosis

EMDB-13978: 
S. cerevisiae CMGE nucleating origin DNA melting
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
