-Search query
-Search result
Showing 1 - 50 of 305 items for (author: kater & l)

EMDB-19767: 
Structure of a 2873 Scaffold Base DNA Origami V1

EMDB-19769: 
Structure of a 2873 Scaffold Base DNA Origami V2

EMDB-19770: 
Structure of a 2873 Scaffold Base DNA Origami V3

EMDB-19775: 
Structure of a 1033 Scaffold Base DNA Origami Nanostructure V4 with Desalted Purified Staples

EMDB-19776: 
Structure of a 1033 Scaffold Base DNA Origami Nanostructure V4 with HPLC Purified Staples
EMDB-19867: 
Cryo-EM Structure of a 1033 Scaffold Base DNA Origami Nanostructure V4 and TBA

EMDB-19874: 
Refinement Focused on the 1st Body of a 1033 Scaffold-Based DNA Origami Nanostructure V4 with TBA

EMDB-19875: 
Refinement Focused on the 2nd Body of a 1033 Scaffold-Based DNA Origami Nanostructure V4 with TBA

EMDB-19876: 
Refinement Focused on the 3rd Body of a 1033 Scaffold-Based DNA Origami Nanostructure V4 with TBA

PDB-9eoq: 
Cryo-EM Structure of a 1033 Scaffold Base DNA Origami Nanostructure V4 and TBA
EMDB-19477: 
Saccharomyces cerevisiae FAS type I

EMDB-19489: 
Tobacco mosaic virus from scanning transmission electron microscopy at CSA=2.0 mrad

EMDB-19132: 
Structure of dynein-2 intermediate chain DYNC2I2 (WDR34) in complex with dynein-2 heavy chain DYNC2H1.

EMDB-19133: 
Structure of dynein-2 intermediate chain DYNC2I1 (WDR60) in complex with the dynein-2 heavy chain DYNC2H1.

PDB-8rgg: 
Structure of dynein-2 intermediate chain DYNC2I2 (WDR34) in complex with dynein-2 heavy chain DYNC2H1.

PDB-8rgh: 
Structure of dynein-2 intermediate chain DYNC2I1 (WDR60) in complex with the dynein-2 heavy chain DYNC2H1.
EMDB-18497: 
Human Sirtuin 6 in complex with nucleosome - structure showing H2A tail path

EMDB-40223: 
Cryo-ET reconstruction of WT HSV-1 NEC coat

EMDB-40224: 
Cryo-ET reconstruction of HSV-1 DN/SUP mutant NEC coat
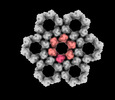

EMDB-40225: 
Cryo-ET reconstruction of HSV-1 SUP mutant NEC coat

EMDB-16552: 
Structure of the RQT-bound 80S ribosome from S. cerevisiae (C2) - composite map

EMDB-16553: 
Structure of RQT (C1) bound to the stalled ribosome in a disome unit from S. cerevisiae - focused Refinement 80S

EMDB-16554: 
Structure of RQT (C1) bound to the stalled ribosome in a disome unit from S. cerevisiae - focused Refinement RQT

EMDB-18809: 
Monomeric E6AP-E6-p53 ternary complex

EMDB-18810: 
Dimeric ternary structure of E6AP-E6-p53

PDB-8r1f: 
Monomeric E6AP-E6-p53 ternary complex

PDB-8r1g: 
Dimeric ternary structure of E6AP-E6-p53

EMDB-17958: 
Structure of DPS determined by cryoEM at 100 keV

EMDB-17959: 
Structure of bacterial ribosome determined by cryoEM at 100 keV

EMDB-17960: 
Structure of GABAAR determined by cryoEM at 100 keV

EMDB-17961: 
Structure of mouse heavy-chain apoferritin determined by cryoEM at 100 keV

EMDB-17962: 
Structure of catalase determined by cryoEM at 100 keV

EMDB-17963: 
Structure of AHIR determined by cryoEM at 100 keV

EMDB-17964: 
Structure of GAPDH determined by cryoEM at 100 keV

EMDB-17965: 
Structure of E. coli glutamine synthetase determined by cryoEM at 100 keV

EMDB-17966: 
Structure of human apo ALDH1A1 determined by cryoEM at 100 keV

EMDB-17967: 
Structure of PaaZ determined by cryoEM at 100 keV

EMDB-17968: 
Structure of lumazine synthase determined by cryoEM at 100 keV

PDB-8pv9: 
Structure of DPS determined by cryoEM at 100 keV

PDB-8pva: 
Structure of bacterial ribosome determined by cryoEM at 100 keV

PDB-8pvb: 
Structure of GABAAR determined by cryoEM at 100 keV

PDB-8pvc: 
Structure of mouse heavy-chain apoferritin determined by cryoEM at 100 keV

PDB-8pvd: 
Structure of catalase determined by cryoEM at 100 keV

PDB-8pve: 
Structure of AHIR determined by cryoEM at 100 keV

PDB-8pvf: 
Structure of GAPDH determined by cryoEM at 100 keV

PDB-8pvg: 
Structure of E. coli glutamine synthetase determined by cryoEM at 100 keV

PDB-8pvh: 
Structure of human apo ALDH1A1 determined by cryoEM at 100 keV

PDB-8pvi: 
Structure of PaaZ determined by cryoEM at 100 keV

PDB-8pvj: 
Structure of lumazine synthase determined by cryoEM at 100 keV

EMDB-17992: 
Structure of K27A mutant E.coli DPS
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
