-Search query
-Search result
Showing 1 - 50 of 63 items for (author: jakobi & aj)

EMDB-19761: 
Cryo-EM structure of Pseudomonas aeruginosa recombinase A (RecA) in complex with ssDNA 72mer and ATPgS
Method: helical / : De Felice S, Vascon F, Huber ST, Grinzato A, Jakobi AJ, Cendron L

EMDB-19771: 
Cryo-EM structure of Pseudomonas aeruginosa Recombinase A (RecA) in complex with LexAS125A mutant
Method: single particle / : De Felice S, Vascon F, Huber ST, Catalano C, Jakobi AJ, Cendron L

EMDB-18968: 
Cryo-EM structure of a coxsackievirus A6 virus-like particle
Method: single particle / : Giannopoulou EA, Jakobi AJ

EMDB-18973: 
Cryo-EM structure of Human SHMT1
Method: single particle / : Spizzichino S, Marabelli C, Bharadwaj A, Jakobi AJ, Chaves-Sanjuan A, Giardina G, Bolognesi M, Cutruzzola F

PDB-8r7h: 
Cryo-EM structure of Human SHMT1
Method: single particle / : Spizzichino S, Marabelli C, Bharadwaj A, Jakobi AJ, Chaves-Sanjuan A, Giardina G, Bolognesi M, Cutruzzola F

EMDB-16813: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ
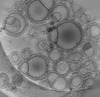
EMDB-16814: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ
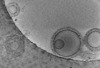
EMDB-16815: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ

EMDB-16794: 
Cryo-EM structure of the human GBP1 dimer bound to GDP-AlF3
Method: single particle / : Kuhm TI, Jakobi AJ

EMDB-15065: 
Cryo-EM structure of the Human SHMT1-RNA complex
Method: single particle / : Spizzichino S, Marabelli C, Bharadwaj A, Jakobi AJ, Chaves-Sanjuan A, Giardina G, Bolognesi M, Cutruzzola F

PDB-8a11: 
Cryo-EM structure of the Human SHMT1-RNA complex
Method: single particle / : Spizzichino S, Marabelli C, Bharadwaj A, Jakobi AJ, Chaves-Sanjuan A, Giardina G, Bolognesi M, Cutruzzola F

EMDB-14106: 
Growing microtubule plus-end in presence of Tip1, Tea2 and Mal3
Method: electron tomography / : Volkov VA

EMDB-14107: 
Growing microtubules in presence of Tip1, Tea2 and Mal3
Method: electron tomography / : Volkov VA

EMDB-14108: 
Microtubule plus-end in presence of Mal3
Method: electron tomography / : Volkov VA

EMDB-14109: 
Growing microtubule minus-end in presence of Mal3
Method: electron tomography / : Volkov VA

EMDB-14110: 
Growing microtubule plus-end in absence of additional proteins
Method: electron tomography / : Volkov VA

EMDB-14111: 
Growing microtubule minus-end in absence of additional proteins
Method: electron tomography / : Volkov VA

EMDB-14112: 
Tip1, Tea2 and Mal3 in presence of PEG without tubulin or microtubules
Method: electron tomography / : Volkov VA

EMDB-14182: 
Tip1, Tea2 and Mal3 in presence of PEG, microtubules and tubulin
Method: electron tomography / : Volkov VA, Dogterom M

EMDB-14238: 
Cryo-EM structure of Bacillus megaterium gas vesicles
Method: helical / : Huber ST, Evers W, Jakobi AJ

EMDB-14340: 
Cryo-EM structure of Bacillus megaterium gas vesicles
Method: helical / : Huber ST, Evers W, Jakobi AJ

EMDB-12901: 
Cryo-EM structure of pyrococcus furiosus apoferritin in nanofluidic channels
Method: single particle / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-12902: 
Electron cryo-tomogram of pyrococcus furiosus apoferritin in nanofluidic channels
Method: electron tomography / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-12903: 
Cryo-EM structure of tobacco mosaic virus in nanofluidic channels
Method: single particle / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-12914: 
Electron cryo-tomogram of tobacco mosaic virus in nanofluidic channels
Method: electron tomography / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-12915: 
Cryo-EM structure of T20S proteasome in nanofluidic channels
Method: single particle / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-12917: 
Electron cryo-tomogram of T20S proteasome in nanofluidic channels
Method: electron tomography / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

PDB-7ohf: 
Cryo-EM structure of pyrococcus furiosus apoferritin in nanofluidic channels
Method: single particle / : Huber ST, Sarajlic E, Huijink R, Evers WH, Jakobi AJ

EMDB-10500: 
Cryo-EM structure of AtNBR1-PB1 filament (S-type)
Method: helical / : Jakobi AJ, Sachse C

EMDB-10499: 
Cryo-EM structure of AtNBR1-PB1 filament (L-type)
Method: helical / : Jakobi AJ, Sachse C

EMDB-10501: 
Cryo-EM structure of p62-PB1 filament (L-type)
Method: helical / : Huber ST, Jakobi AJ, Mortensen SA, Sachse C

EMDB-10502: 
Cryo-EM structure of p62-PB1 filament (L-type)
Method: helical / : Huber ST, Jakobi AJ, Mortensen SA, Sachse C

EMDB-3727: 
RNA polymerase I pre-initiation complex
Method: single particle / : Sadian Y, Tafur L, Kosinski J, Jakobi AJ, Wetzel R, Buczak K, Hagen WJH, Beck M, Sachse C, Muller CW

EMDB-3728: 
RNA polymerase I pre-initiation complex (CF focused refinement)
Method: single particle / : Sadian Y, Tafur L, Kosinski J, Jakobi AJ, Wetzel R, Buczak K, Hagen WJH, Beck M, Sachse C, Muller CW

EMDB-3729: 
RNA polymerase I pre-initiation complex (Pol-Rrn3 focused refinement)
Method: single particle / : Sadian Y, Tafur L, Kosinski J, Jakobi AJ, Wetzel R, Buczak K, Hagen WJH, Beck M, Sachse C, Muller CW

EMDB-3449: 
RNA Polymerase I elongation complex with A49 tandem winged helix domain
Method: single particle / : Tafur L, Sadian Y
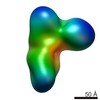
EMDB-8235: 
Negative stain structure of Vps15/Vps34 complex
Method: single particle / : Kirsten ML, Zhang L

EMDB-4016: 
A 4.5 Angstrom structure of HIV-1 CA-SP1
Method: subtomogram averaging / : Schur FKM, Obr M, Hagen WJH, Wan W, Arjen JJ, Kirkpatrick JM, Sachse C, Kraeusslich HG, Briggs JAG

EMDB-4017: 
A 4.2 Angstrom structure of HIV-1 CA-SP1 from immature HIV-1 particles
Method: subtomogram averaging / : Schur FKM, Obr M, Hagen WJH, Wan W, Arjen JJ, Kirkpatrick JM, Sachse C, Kraeusslich HG, Briggs JAG

EMDB-4018: 
Cryo-electron tomogram containing HIV-1 deltaMACANC virus-like particles assembled in vitro in presence of Bevirimat
Method: electron tomography / : Schur FKM, Obr M, Hagen WJH, Wan W, Arjen JJ, Kirkpatrick JM, Sachse C, Kraeusslich HG, Briggs JAG

EMDB-4019: 
Cryo-electron tomogram containing HIV-1 deltaMACANC virus-like particles assembled in vitro
Method: electron tomography / : Schur FKM, Obr M, Hagen WJH, Wan W, Arjen JJ, Kirkpatrick JM, Sachse C, Kraeusslich HG, Briggs JAG

EMDB-4020: 
Cryo-electron tomogram containing immature HIV-1 particles
Method: electron tomography / : Schur FKM, Obr M, Hagen WJH, Wan W, Arjen JJ, Kirkpatrick JM, Sachse C, Kraeusslich HG, Briggs JAG

EMDB-4015: 
A 3.9 Angstrom structure of HIV-1 CA-SP1 assembled in presence of Bevirimat
Method: subtomogram averaging / : Schur FKM, Obr M

EMDB-8166: 
Structure of the S. cerevisiae alpha-mannosidase 1
Method: single particle / : Schneider S, Kosinski J

EMDB-8167: 
Structure of S. cerevesiae mApe1 dodecamer
Method: single particle / : Sachse C, Bertipaglia C

EMDB-3178: 
Cryo-EM structure of yeast RNA polymerase III elongation complex at 3.9 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

EMDB-3179: 
Cryo-EM structure of yeast apo RNA polymerase III at 4.6 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
