[English] 日本語
 Yorodumi
Yorodumi- EMDB-12903: Cryo-EM structure of tobacco mosaic virus in nanofluidic channels -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12903 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of tobacco mosaic virus in nanofluidic channels | |||||||||
 Map data Map data | Sharpened and symmetrized cryo-EM density | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Tobacco mosaic virus-like, coat protein / Tobacco mosaic virus-like, coat protein superfamily / Virus coat protein (TMV like) / helical viral capsid / structural molecule activity / identical protein binding / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |   Tobacco mosaic virus Tobacco mosaic virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Huber ST / Sarajlic E / Huijink R / Evers WH / Jakobi AJ | |||||||||
| Funding support | European Union,  Netherlands, 2 items Netherlands, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Nanofluidic chips for cryo-EM structure determination from picoliter sample volumes. Authors: Stefan T Huber / Edin Sarajlic / Roeland Huijink / Felix Weis / Wiel H Evers / Arjen J Jakobi /   Abstract: Cryogenic electron microscopy has become an essential tool for structure determination of biological macromolecules. In practice, the difficulty to reliably prepare samples with uniform ice thickness ...Cryogenic electron microscopy has become an essential tool for structure determination of biological macromolecules. In practice, the difficulty to reliably prepare samples with uniform ice thickness still represents a barrier for routine high-resolution imaging and limits the current throughput of the technique. We show that a nanofluidic sample support with well-defined geometry can be used to prepare cryo-EM specimens with reproducible ice thickness from picoliter sample volumes. The sample solution is contained in electron-transparent nanochannels that provide uniform thickness gradients without further optimisation and eliminate the potentially destructive air-water interface. We demonstrate the possibility to perform high-resolution structure determination with three standard protein specimens. Nanofabricated sample supports bear potential to automate the cryo-EM workflow, and to explore new frontiers for cryo-EM applications such as time-resolved imaging and high-throughput screening. #1:  Journal: Biorxiv / Year: 2021 Journal: Biorxiv / Year: 2021Title: Nanofluidic chips for cryo-EM structure determination from picoliter sample volumes Authors: Huber ST / Sarajlic E / Huijink R / Weis F / Evers WH / Jakobi AJ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12903.map.gz emd_12903.map.gz | 18.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12903-v30.xml emd-12903-v30.xml emd-12903.xml emd-12903.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12903_fsc.xml emd_12903_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_12903.png emd_12903.png | 177.4 KB | ||
| Masks |  emd_12903_msk_1.map emd_12903_msk_1.map | 64 MB |  Mask map Mask map | |
| Others |  emd_12903_additional_1.map.gz emd_12903_additional_1.map.gz emd_12903_additional_2.map.gz emd_12903_additional_2.map.gz emd_12903_half_map_1.map.gz emd_12903_half_map_1.map.gz emd_12903_half_map_2.map.gz emd_12903_half_map_2.map.gz | 18.6 MB 12.3 MB 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12903 http://ftp.pdbj.org/pub/emdb/structures/EMD-12903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12903 | HTTPS FTP |
-Related structure data
| Related structure data |  7ohfC C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10708 (Title: Apoferritin, TMV and T20S proteasome in nanofluidic channels EMPIAR-10708 (Title: Apoferritin, TMV and T20S proteasome in nanofluidic channelsData size: 308.3 Data #1: Unaligned multi-frame movies of pyrococcus furiosus apoferritin in silicon nitride nanochannels [micrographs - multiframe] Data #2: Multi-frame movies of T20S proteasome in silicon nitride nanochannels [micrographs - multiframe] Data #3: Multi-frame movies of TMV in silicon nitride nanochannels [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12903.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12903.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened and symmetrized cryo-EM density | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.288 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12903_msk_1.map emd_12903_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
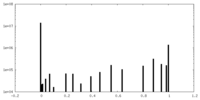
| Density Histograms |
-Additional map: Unsharpened and symmetrized cryo-EM density
| File | emd_12903_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened and symmetrized cryo-EM density | ||||||||||||
| Projections & Slices |
| ||||||||||||
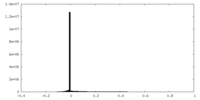
| Density Histograms |
-Additional map: Local resolution map
| File | emd_12903_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution map | ||||||||||||
| Projections & Slices |
| ||||||||||||
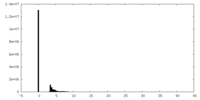
| Density Histograms |
-Half map: Half map A
| File | emd_12903_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
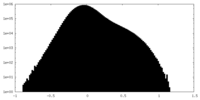
| Density Histograms |
-Half map: Half map B
| File | emd_12903_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tobacco mosaic virus in nanofluidic channels
| Entire | Name: Tobacco mosaic virus in nanofluidic channels |
|---|---|
| Components |
|
-Supramolecule #1: Tobacco mosaic virus in nanofluidic channels
| Supramolecule | Name: Tobacco mosaic virus in nanofluidic channels / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Tobacco mosaic virus Tobacco mosaic virus |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.1 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: Homemade / Material: SILICON NITRIDE |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA PLUNGER Details: The sample was filled into cryoChips through the cantilever and then transferred within ~10 seconds to the Leica plunger for freezing.. |
| Details | The sample was filled into nanofluidic channels. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 82 / Average exposure time: 11.0 sec. / Average electron dose: 59.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.1 mm |
| Sample stage | Specimen holder model: JEOL 3200FSC CRYOHOLDER |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)