-Search query
-Search result
Showing 1 - 50 of 642 items for (author: hay & id)
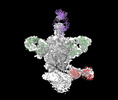
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.

EMDB-42394: 
Single particle analysis of recombinant human MFAP4

EMDB-42398: 
MFAP4 after treatment with EDTA/without Ca2+

PDB-8un7: 
Single particle analysis of recombinant human MFAP4

EMDB-44644: 
SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052
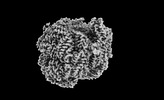
EMDB-41434: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)
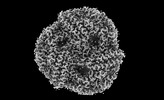
EMDB-41435: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
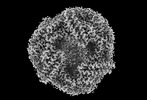
EMDB-41436: 
Central rod disk in D3 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
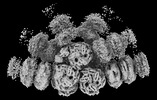
EMDB-41463: 
Synechocystis PCC 6803 Phycobilisome quenched by OCP, high resolution

EMDB-41475: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)
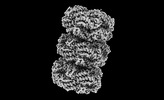
EMDB-41585: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8to2: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8to5: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8tpj: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8tro: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)

EMDB-18674: 
Structure of nPBD of human SRP68/72

EMDB-43226: 
DNA origami colloid for self-assembly of tubules: (6,0) monomer

EMDB-43227: 
DNA origami colloid for self-assembly of tubules: (10,0) monomer

EMDB-41810: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-41820: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41823: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41838: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to partially open HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

PDB-8u1d: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-18350: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 7.5

EMDB-18351: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 5.5

EMDB-18352: 
S. cerevisia Niemann-Pick type C protein NCR1 in LMNG at pH 5.5

EMDB-18353: 
S. cerevisia Niemann-Pick type C protein NCR1 in Peptidisc at pH 7.5

PDB-8qeb: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 7.5

PDB-8qec: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 5.5

PDB-8qed: 
S. cerevisia Niemann-Pick type C protein NCR1 in LMNG at pH 5.5

PDB-8qee: 
S. cerevisia Niemann-Pick type C protein NCR1 in Peptidisc at pH 7.5

EMDB-18448: 
CTE type II (tau intermediate amyloid)

PDB-8qjj: 
CTE type II (tau intermediate amyloid)

EMDB-28209: 
Apo rat TRPV2 in nanodiscs, state 1

EMDB-28210: 
Apo rat TRPV2 in nanodiscs, state 2

EMDB-28211: 
Apo rat TRPV2 in nanodiscs, state 3

EMDB-28212: 
rat TRPV2 in nanodiscs in the presence of weak acid at pH 5

PDB-8ekp: 
Apo rat TRPV2 in nanodiscs, state 1

PDB-8ekq: 
Apo rat TRPV2 in nanodiscs, state 2

PDB-8ekr: 
Apo rat TRPV2 in nanodiscs, state 3

PDB-8eks: 
rat TRPV2 in nanodiscs in the presence of weak acid at pH 5

EMDB-18112: 
Tau - AD-MIA4

EMDB-18252: 
Tau - AD-LIA4 (tau intermediate amyloid)

EMDB-18253: 
Tau - AD-LIA5 (tau intermediate amyloid)

EMDB-18254: 
Tau - AD-LIA7 (tau intermediate amyloid)

EMDB-18255: 
Tau - AD-PHF long crossover

EMDB-18258: 
Tau - AD-PHF

EMDB-18259: 
Tau - AD-THF

EMDB-18261: 
Tau - CTE-MIA1 (tau intermediate amyloid)

EMDB-18262: 
Tau - CTE-MIA3
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
