-Search query
-Search result
Showing all 28 items for (author: fischer & ef)

EMDB-48504: 
Cryo-EM structure of VCP bound to p47 UBX domain and VCPIP1 VCPIDs (with stalk region)
Method: single particle / : Shah B, Hunkeler M, Buhrlage SJ, Fischer EF

EMDB-48505: 
Cryo-EM structure of p47 bound to VCP N-domain (with D1 domain)
Method: single particle / : Shah B, Hunkeler M, Buhrlage SJ, Fischer EF

EMDB-48506: 
Cryo-EM structure of three VCPIP1 VCPIDs bound to VCP
Method: single particle / : Shah B, Hunkeler M, Buhrlage SJ, Fischer EF

EMDB-50336: 
Cryo-EM structure of the ternary DARPin NY_1/HLA-A0201/NY-ESO1 complex.
Method: single particle / : Schulte T, Wallden K, Carroni M, Sandalova T, Walser M, Mueller S, Venetz N, Achour A

PDB-9fe1: 
Cryo-EM structure of the ternary DARPin NY_1/HLA-A0201/NY-ESO1 complex.
Method: single particle / : Schulte T, Wallden K, Carroni M, Sandalova T, Walser M, Mueller S, Venetz N, Achour A

EMDB-17539: 
Cryo-EM structure of dimeric UBR5
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

EMDB-17540: 
Cryo-EM structure of full-length human UBR5 (homotetramer)
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

PDB-8p82: 
Cryo-EM structure of dimeric UBR5
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

PDB-8p83: 
Cryo-EM structure of full-length human UBR5 (homotetramer)
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

EMDB-17542: 
Negative stain map of UBR5 (dimer) in complex with RARA/RXRA
Method: single particle / : Aguirre JD, Cavadini S, Kempf G, Kater L, Thoma NH

EMDB-16035: 
Tau Paired Helical Filament from Extracellular Vesicles from Alzheimer's disease brain (Individual 1)
Method: helical / : Behr TS, Ryskeldi-Falcon B

EMDB-16039: 
Tau Paired Helical Filament from Cellular Fraction of Alzheimer's disease brain
Method: helical / : Behr TS, Ryskeldi-Falcon B
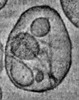
EMDB-16064: 
Cryo-electron tomogram of an extracellular vesicle containing tau filaments isolated from Alzheimer's Disease patient brain
Method: electron tomography / : Behr TS, Ryskeldi-Falcon B

PDB-8bgs: 
Tau Paired Helical Filament from Extracellular Vesicles from Alzheimer's disease brain
Method: helical / : Behr TS, Ryskeldi-Falcon B

PDB-8bgv: 
Tau Paired Helical Filament from Cellular Fraction of Alzheimer's disease brain
Method: helical / : Behr TS, Ryskeldi-Falcon B

EMDB-11610: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with megabody MbNbF3c7HopQ
Method: single particle / : Uchanski T, Masiulis S

EMDB-4542: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with histamine and megabody Mb25
Method: single particle / : Uchanski T, Masiulis S

PDB-6qfa: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with histamine and megabody Mb25
Method: single particle / : Uchanski T, Masiulis S, Fischer B, Kalichuk V, Wohlkoening A, Zoegg T, Remaut H, Vranken W, Aricescu AR, Pardon E, Steyaert J

EMDB-23211: 
Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1
Method: single particle / : Lees JA, Han S

EMDB-23215: 
Cryo-EM structure of protein encoded by vaccine candidate BNT162b2
Method: single particle / : Lees JA, Han S

PDB-7l7f: 
Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1
Method: single particle / : Lees JA, Han S

PDB-7l7k: 
Cryo-EM structure of protein encoded by vaccine candidate BNT162b2
Method: single particle / : Lees JA, Han S

EMDB-22265: 
Cryo-EM structure of BCL6 bound to BI-3802
Method: single particle / : Yoon H, Burman SSR

PDB-6xmx: 
Cryo-EM structure of BCL6 bound to BI-3802
Method: single particle / : Yoon H, Burman SSR, Hunkeler M, Nowak RP, Fischer ES

EMDB-8624: 
Cryo EM structure of anti-CRISPRs, AcrF1 and AcrF2, bound to type I-F crRNA-guided CRISPR surveillance complex
Method: single particle / : Chowdhury S, Carter J

PDB-5uz9: 
Cryo EM structure of anti-CRISPRs, AcrF1 and AcrF2, bound to type I-F crRNA-guided CRISPR surveillance complex
Method: single particle / : Chowdhury S, Carter J, Rollins MF, Jackson RN, Hoffmann C, Nosaka L, Bondy-Denomy J, Maxwell KL, Davidson AR, Fischer ER, Lander GC, Wiedenheft B

PDB-5aef: 
Electron cryo-microscopy of an Abeta(1-42)amyloid fibril
Method: helical / : Schmidt M, Rohou A, Lasker K, Yadav JK, Schiene-Fischer C, Fandrich M, Grigorieff N

EMDB-3132: 
Electron cryo-microscopy of an Abeta(1-42)amyloid fibril
Method: helical / : Schmidt M, Rohou A, Lasker K, Yadav JK, Schiene-Fischer C, Fandrich M, Grigorieff N
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
