[English] 日本語
 Yorodumi
Yorodumi- EMDB-23211: Cryo-EM structure of human ACE2 receptor bound to protein encoded... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23211 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1 | |||||||||
 Map data Map data | ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / COVID19 / BNT162b1 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction ...positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction / peptidyl-dipeptidase activity / transporter activator activity / regulation of vasoconstriction / Metabolism of Angiotensinogen to Angiotensins / carboxypeptidase activity / angiotensin maturation / viral life cycle / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / blood vessel diameter maintenance / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / positive regulation of reactive oxygen species metabolic process / metallopeptidase activity / endocytic vesicle membrane / regulation of cell population proliferation / virus receptor activity / regulation of inflammatory response / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / endopeptidase activity / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / Potential therapeutics for SARS / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / cilium / membrane raft / endocytosis involved in viral entry into host cell / endoplasmic reticulum lumen / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / cell surface / negative regulation of transcription by RNA polymerase II / : / extracellular exosome / extracellular region / zinc ion binding / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /    Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.24 Å | |||||||||
 Authors Authors | Lees JA / Han S | |||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: BNT162b vaccines protect rhesus macaques from SARS-CoV-2. Authors: Annette B Vogel / Isis Kanevsky / Ye Che / Kena A Swanson / Alexander Muik / Mathias Vormehr / Lena M Kranz / Kerstin C Walzer / Stephanie Hein / Alptekin Güler / Jakob Loschko / Mohan S ...Authors: Annette B Vogel / Isis Kanevsky / Ye Che / Kena A Swanson / Alexander Muik / Mathias Vormehr / Lena M Kranz / Kerstin C Walzer / Stephanie Hein / Alptekin Güler / Jakob Loschko / Mohan S Maddur / Ayuko Ota-Setlik / Kristin Tompkins / Journey Cole / Bonny G Lui / Thomas Ziegenhals / Arianne Plaschke / David Eisel / Sarah C Dany / Stephanie Fesser / Stephanie Erbar / Ferdia Bates / Diana Schneider / Bernadette Jesionek / Bianca Sänger / Ann-Kathrin Wallisch / Yvonne Feuchter / Hanna Junginger / Stefanie A Krumm / André P Heinen / Petra Adams-Quack / Julia Schlereth / Stefan Schille / Christoph Kröner / Ramón de la Caridad Güimil Garcia / Thomas Hiller / Leyla Fischer / Rani S Sellers / Shambhunath Choudhary / Olga Gonzalez / Fulvia Vascotto / Matthew R Gutman / Jane A Fontenot / Shannan Hall-Ursone / Kathleen Brasky / Matthew C Griffor / Seungil Han / Andreas A H Su / Joshua A Lees / Nicole L Nedoma / Ellene H Mashalidis / Parag V Sahasrabudhe / Charles Y Tan / Danka Pavliakova / Guy Singh / Camila Fontes-Garfias / Michael Pride / Ingrid L Scully / Tara Ciolino / Jennifer Obregon / Michal Gazi / Ricardo Carrion / Kendra J Alfson / Warren V Kalina / Deepak Kaushal / Pei-Yong Shi / Thorsten Klamp / Corinna Rosenbaum / Andreas N Kuhn / Özlem Türeci / Philip R Dormitzer / Kathrin U Jansen / Ugur Sahin /   Abstract: A safe and effective vaccine against COVID-19 is urgently needed in quantities that are sufficient to immunize large populations. Here we report the preclinical development of two vaccine candidates ...A safe and effective vaccine against COVID-19 is urgently needed in quantities that are sufficient to immunize large populations. Here we report the preclinical development of two vaccine candidates (BNT162b1 and BNT162b2) that contain nucleoside-modified messenger RNA that encodes immunogens derived from the spike glycoprotein (S) of SARS-CoV-2, formulated in lipid nanoparticles. BNT162b1 encodes a soluble, secreted trimerized receptor-binding domain (known as the RBD-foldon). BNT162b2 encodes the full-length transmembrane S glycoprotein, locked in its prefusion conformation by the substitution of two residues with proline (S(K986P/V987P); hereafter, S(P2) (also known as P2 S)). The flexibly tethered RBDs of the RBD-foldon bind to human ACE2 with high avidity. Approximately 20% of the S(P2) trimers are in the two-RBD 'down', one-RBD 'up' state. In mice, one intramuscular dose of either candidate vaccine elicits a dose-dependent antibody response with high virus-entry inhibition titres and strong T-helper-1 CD4 and IFNγCD8 T cell responses. Prime-boost vaccination of rhesus macaques (Macaca mulatta) with the BNT162b candidates elicits SARS-CoV-2-neutralizing geometric mean titres that are 8.2-18.2× that of a panel of SARS-CoV-2-convalescent human sera. The vaccine candidates protect macaques against challenge with SARS-CoV-2; in particular, BNT162b2 protects the lower respiratory tract against the presence of viral RNA and shows no evidence of disease enhancement. Both candidates are being evaluated in phase I trials in Germany and the USA, and BNT162b2 is being evaluated in an ongoing global phase II/III trial (NCT04380701 and NCT04368728). | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23211.map.gz emd_23211.map.gz | 325.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23211-v30.xml emd-23211-v30.xml emd-23211.xml emd-23211.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23211.png emd_23211.png | 190.3 KB | ||
| Filedesc metadata |  emd-23211.cif.gz emd-23211.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23211 http://ftp.pdbj.org/pub/emdb/structures/EMD-23211 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23211 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23211 | HTTPS FTP |
-Related structure data
| Related structure data |  7l7fMC  7l7kC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23211.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23211.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.87 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
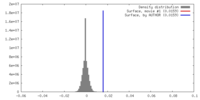
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human ACE2 receptor complexed with protein encoded by vaccine can...
| Entire | Name: Human ACE2 receptor complexed with protein encoded by vaccine candidate BNT162b1 |
|---|---|
| Components |
|
-Supramolecule #1: Human ACE2 receptor complexed with protein encoded by vaccine can...
| Supramolecule | Name: Human ACE2 receptor complexed with protein encoded by vaccine candidate BNT162b1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Angiotensin-converting enzyme 2
| Supramolecule | Name: Angiotensin-converting enzyme 2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Spike glycoprotein, Envelope glycoprotein fusion
| Supramolecule | Name: Spike glycoprotein, Envelope glycoprotein fusion / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Angiotensin-converting enzyme 2
| Macromolecule | Name: Angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: angiotensin-converting enzyme 2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 92.556695 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSSSSWLLLS LVAVTAAQST IEEQAKTFLD KFNHEAEDLF YQSSLASWNY NTNITEENVQ NMNNAGDKWS AFLKEQSTLA QMYPLQEIQ NLTVKLQLQA LQQNGSSVLS EDKSKRLNTI LNTMSTIYST GKVCNPDNPQ ECLLLEPGLN EIMANSLDYN E RLWAWESW ...String: MSSSSWLLLS LVAVTAAQST IEEQAKTFLD KFNHEAEDLF YQSSLASWNY NTNITEENVQ NMNNAGDKWS AFLKEQSTLA QMYPLQEIQ NLTVKLQLQA LQQNGSSVLS EDKSKRLNTI LNTMSTIYST GKVCNPDNPQ ECLLLEPGLN EIMANSLDYN E RLWAWESW RSEVGKQLRP LYEEYVVLKN EMARANHYED YGDYWRGDYE VNGVDGYDYS RGQLIEDVEH TFEEIKPLYE HL HAYVRAK LMNAYPSYIS PIGCLPAHLL GDMWGRFWTN LYSLTVPFGQ KPNIDVTDAM VDQAWDAQRI FKEAEKFFVS VGL PNMTQG FWENSMLTDP GNVQKAVCHP TAWDLGKGDF RILMCTKVTM DDFLTAHHEM GHIQYDMAYA AQPFLLRNGA NEGF HEAVG EIMSLSAATP KHLKSIGLLS PDFQEDNETE INFLLKQALT IVGTLPFTYM LEKWRWMVFK GEIPKDQWMK KWWEM KREI VGVVEPVPHD ETYCDPASLF HVSNDYSFIR YYTRTLYQFQ FQEALCQAAK HEGPLHKCDI SNSTEAGQKL FNMLRL GKS EPWTLALENV VGAKNMNVRP LLNYFEPLFT WLKDQNKNSF VGWSTDWSPY ADQSIKVRIS LKSALGDKAY EWNDNEM YL FRSSVAYAMR QYFLKVKNQM ILFGEEDVRV ANLKPRISFN FFVTAPKNVS DIIPRTEVEK AIRMSRSRIN DAFRLNDN S LEFLGIQPTL GPPNQPPVSI WLIVFGVVMG VIVVGIVILI FTGIRDRKKK NKARSGENPY ASIDISKGEN NPGFQNTDD VQTSF UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #2: Spike glycoprotein, Envelope glycoprotein fusion
| Macromolecule | Name: Spike glycoprotein, Envelope glycoprotein fusion / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 29.490281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVVRFP NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADS FVIRGDEVRQ IAPGQTGKIA DYNYKLPDDF TGCVIAWNSN NLDSKVGGNY NYLYRLFRKS NLKPFERDIS T EIYQAGST ...String: MFVFLVLLPL VSSQCVVRFP NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADS FVIRGDEVRQ IAPGQTGKIA DYNYKLPDDF TGCVIAWNSN NLDSKVGGNY NYLYRLFRKS NLKPFERDIS T EIYQAGST PCNGVEGFNC YFPLQSYGFQ PTNGVGYQPY RVVVLSFELL HAPATVCGPK GSPGSGSGSG YIPEAPRDGQ AY VRKDGEW VLLSTFLGRS LEVLFQGPG UniProtKB: Spike glycoprotein, Spike glycoprotein, Envelope glycoprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 52.51 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.24 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 74784 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)