-Search query
-Search result
Showing 1 - 50 of 51 items for (author: fabre & l)

EMDB-51273: 
Cryo-EM structure of Vibrio cholerae RNA polymerase holoenzyme bound to an ompU promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51274: 
Cryo-EM structure of Vibrio cholerae RNA polymerase holoenzyme bound to an ompU promoter DNA fragment and 5-mer RNA
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51275: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR transcription factor and ompU promoter DNA
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51276: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with TcpP transcription factor and a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51277: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR and TcpP transcription factors and a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51278: 
Cryo-EM structure of Vibrio cholerae RNA polymerase dimer with ToxR and TcpP transcription factors and a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51774: 
Consensus map of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR transcription factor and bound to an ompU promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51775: 
Focused map #1 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR transcription factor and bound to an ompU promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51776: 
Focused map #2 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR transcription factor and bound to an ompU promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51948: 
Consensus map of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with TcpP transcription factor and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51949: 
Focused map #1 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with TcpP transcription factor and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51950: 
Focused map #2 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with TcpP transcription factor and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51955: 
Consensus map of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR and TcpP transcription factors and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51956: 
Focused map #1 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR and TcpP transcription factors and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-51958: 
Focused map #2 of the cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR and TcpP transcription factors and bound to a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

PDB-9gdo: 
Cryo-EM structure of Vibrio cholerae RNA polymerase holoenzyme bound to an ompU promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

PDB-9gdp: 
Cryo-EM structure of Vibrio cholerae RNA polymerase holoenzyme bound to an ompU promoter DNA fragment and 5-mer RNA
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

PDB-9gdq: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR transcription factor and ompU promoter DNA
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

PDB-9gdr: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with TcpP transcription factor and a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

PDB-9gds: 
Cryo-EM structure of Vibrio cholerae RNA polymerase Transcription Activation Complex with ToxR and TcpP transcription factors and a toxT promoter DNA fragment
Method: single particle / : Alcaide-Jimenez A, Baudin F, Canals A, Machon C, Murciano B, Fabrega-Ferrer M, Bantysh O, Perez-Luque R, Krukonis ES, Muller CW, Coll M

EMDB-19352: 
Cryo-EM structure of the 60S subunit with the RQC complex containing the mutant F340I Rqc2 protein and the Flag tagged Ltn1-Delta_ring
Method: single particle / : Fabret C, Giudice E, Chat S, Gillet R, Namy O

EMDB-21361: 
TriABC triclosan efflux pump from Pseudomonas aeruginosa- No symmetry
Method: single particle / : Rouiller I, Fabre L, Bhattacharyya S, Sygusch J

EMDB-21362: 
Negative stain map of TriABC triclosan efflux pump from Pseudomonas aeruginosa
Method: single particle / : Rouiller I, Fabre L, Sygusch J

EMDB-21363: 
Cryo-EM map of TriABC triclosan efflux pump from Pseudomonas aeruginosa
Method: single particle / : Fabre L, Bhattacharyya S, Ruickoldt J, Rouiller I, Zgurskaya HI, Sygusch J

PDB-6vej: 
TriABC transporter from Pseudomonas aeruginosa
Method: single particle / : Sygusch J, Fabre L, Rouiller I, Bhattacharyya S

EMDB-10010: 
Atomic structure of the Epstein-Barr portal, structure I
Method: single particle / : Machon C, Fabrega-Ferrer M

EMDB-10011: 
Atomic structure of the Epstein-Barr portal, structure II
Method: single particle / : Machon C, Fabrega-Ferrer M

PDB-6rvr: 
Atomic structure of the Epstein-Barr portal, structure I
Method: single particle / : Machon C, Fabrega-Ferrer M, Zhou D, Cuervo A, Carrascosa JL, Stuart DI, Coll M

PDB-6rvs: 
Atomic structure of the Epstein-Barr portal, structure II
Method: single particle / : Machon C, Fabrega-Ferrer M, Zhou D, Cuervo A, Carrascosa JL, Stuart DI, Coll M

EMDB-4667: 
Cryo-EM structure of T7 bacteriophage portal protein, 13mer, closed valve
Method: single particle / : Cuervo A, Fabrega-Ferrer M, Machon C, Fernandez FJ, Perez-Luque R, Pous J, Vega MC, Carrascosa JL, Coll M

EMDB-4669: 
Bacteriophage packaging protein
Method: single particle / : Fabrega-Ferrer M, Cuervo A

EMDB-4706: 
Bacteriophage ejection complex
Method: single particle / : Cuervo A, Fabrega-Ferrer M

PDB-6qxm: 
Cryo-EM structure of T7 bacteriophage portal protein, 12mer, open valve
Method: single particle / : Fabrega-Ferrer M, Cuervo A, Machon C, Fernandez FJ, Perez-Luque R, Pous J, Vega MC, Carrascosa JL, Coll M

PDB-6r21: 
Cryo-EM structure of T7 bacteriophage fiberless tail complex
Method: single particle / : Cuervo A, Fabrega-Ferrer M, Machon C, Conesa JJ, Perez-Ruiz M, Coll M, Carrascosa JL

EMDB-8530: 
MalE-MalFGK2 ADP-V04
Method: single particle / : Fabre L, Bao H, Innes J, Duong F, Rouiller I
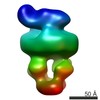
EMDB-8531: 
MalE-x-MalFGK2 ADP
Method: single particle / : Fabre L, Bao H, Innes J, Duong F, Rouiller I

EMDB-8534: 
MalE-x-MalFGK2 Maltose (open conformation)
Method: single particle / : Fabre L, Bao H, Innes J, Duong F, Rouiller I

EMDB-8535: 
MalE-x-MalFGK2 Maltose (semi-closed symmetric conformation)
Method: single particle / : Fabre L, Bao H, Innes J, Duong F, Rouiller I
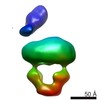
EMDB-8536: 
MalE-MalFGK2 maltose with MalE semi-detached
Method: single particle / : Fabre L, Bao H, Innes J, Duong F, Rouiller I

EMDB-2859: 
3D Reconstruction of Membrane Protein Complex ExbB4-ExbD1-TonB1
Method: single particle / : Sverzhinsky A, Chung JWC, Deme JC, Fabre L, Levey KT, Plesa M, Carter DM, Lypaczewski P, Coulton JW

EMDB-2934: 
3D Reconstruction of Membrane Protein Complex ExbB4-ExbD1-TonB1
Method: single particle / : Sverzhinsky A, Chung JWC, Deme JC, Fabre L, Levey KT, Plesa M, Carter DM, Lypaczewski P, Coulton JW

EMDB-2935: 
3D Reconstruction of Membrane Protein Complex ExbB4-ExbD1-TonB1
Method: single particle / : Sverzhinsky A, Chung JWC, Deme JC, Fabre L, Levey KT, Plesa M, Carter DM, Lypaczewski P, Coulton JW

EMDB-5901: 
3D Reconstruction of Membrane Protein Complex ExbB4-ExbD2
Method: single particle / : Sverzhinsky A, Fabre L, Cottreau AL, Biot-Pelletier DMP, Khalil S, Bostina M, Rouiller I, Coulton JW

EMDB-5902: 
3D Reconstruction of Membrane Protein Complex ExbB4-ExbD2
Method: single particle / : Sverzhinsky A, Fabre L, Cottreau AL, Biot-Pelletier DMP, Khalil S, Bostina M, Rouiller I, Coulton JW
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search







 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
