+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10010 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
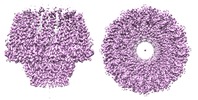
| Title | Atomic structure of the Epstein-Barr portal, structure I | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | viral protein / DNA packaging | ||||||||||||||||||||||||
| Function / homology | Herpesvirus portal protein / Herpesvirus UL6 like / chromosome organization / virion component / host cell nucleus / BBRF1 / Portal protein Function and homology information Function and homology information | ||||||||||||||||||||||||
| Biological species |  Human gammaherpesvirus 4 (Epstein-Barr virus) / Human gammaherpesvirus 4 (Epstein-Barr virus) /  Epstein-Barr virus (strain GD1) (Epstein-Barr virus) Epstein-Barr virus (strain GD1) (Epstein-Barr virus) | ||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.46 Å | ||||||||||||||||||||||||
 Authors Authors | Machon C / Fabrega-Ferrer M | ||||||||||||||||||||||||
| Funding support |  Spain, 7 items Spain, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Atomic structure of the Epstein-Barr virus portal. Authors: Cristina Machón / Montserrat Fàbrega-Ferrer / Daming Zhou / Ana Cuervo / José L Carrascosa / David I Stuart / Miquel Coll /   Abstract: Herpesviridae is a vast family of enveloped DNA viruses that includes eight distinct human pathogens, responsible for diseases that range from almost asymptomatic to severe and life-threatening. ...Herpesviridae is a vast family of enveloped DNA viruses that includes eight distinct human pathogens, responsible for diseases that range from almost asymptomatic to severe and life-threatening. Epstein-Barr virus infects B-cells and epithelial cells, causing infectious mononucleosis, as well as a number of cancers. Epstein-Barr infection cannot be cured since neither vaccine nor antiviral drug treatments are available. All herpesviruses contain a linear double-stranded DNA genome, enclosed within an icosahedral capsid. Viral portal protein plays a key role in the procapsid assembly and DNA packaging. The portal is the entrance and exit pore for the viral genome, making it an attractive pharmacological target for the development of new antivirals. Here we present the atomic structure of the portal protein of Epstein-Barr virus, solved by cryo-electron microscopy at 3.5 Å resolution. The detailed architecture of this protein suggests that it plays a functional role in DNA retention during packaging. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10010.map.gz emd_10010.map.gz | 38 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10010-v30.xml emd-10010-v30.xml emd-10010.xml emd-10010.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10010.png emd_10010.png | 467.2 KB | ||
| Filedesc metadata |  emd-10010.cif.gz emd-10010.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10010 http://ftp.pdbj.org/pub/emdb/structures/EMD-10010 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10010 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10010 | HTTPS FTP |
-Related structure data
| Related structure data |  6rvrMC  6rvsC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10010.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10010.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : DNA packaging viral protein
| Entire | Name: DNA packaging viral protein |
|---|---|
| Components |
|
-Supramolecule #1: DNA packaging viral protein
| Supramolecule | Name: DNA packaging viral protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Human gammaherpesvirus 4 (Epstein-Barr virus) Human gammaherpesvirus 4 (Epstein-Barr virus) |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Epstein-Barr virus (strain GD1) (Epstein-Barr virus) Epstein-Barr virus (strain GD1) (Epstein-Barr virus)Strain: GD1 |
| Molecular weight | Theoretical: 68.539641 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MFNMNVDESA SGALGSSAIP VHPTPASVRL FEILQGKYAY VQGQTIYANL RNPGVFSRQV FTHLFKRAIS HCTYDDVLHD WNKFEACIQ KRWPSDDSCA SRFRESTFES WSTTMKLTVR DLLTTNIYRV LHSRSVLSYE RYVDWICATG MVPAVKKPIT Q ELHSKIKS ...String: MFNMNVDESA SGALGSSAIP VHPTPASVRL FEILQGKYAY VQGQTIYANL RNPGVFSRQV FTHLFKRAIS HCTYDDVLHD WNKFEACIQ KRWPSDDSCA SRFRESTFES WSTTMKLTVR DLLTTNIYRV LHSRSVLSYE RYVDWICATG MVPAVKKPIT Q ELHSKIKS LRDRCVCREL GHERTIRSIG TELYEATKEI IESLNSTFIP QFTEVTIEYL PRSDEYVAYY CGRRIRLHVL FP PAIFAGT VTFDSPVQRL YQNIFMCYRT LEHAKICQLL NTAPLKAIVG HGGRDMYKDI LAHLEQNSQR KDPKKELLNL LVK LSENKT ISGVTDVVEE FITDASNNLV DRNRLFGQPG ETAAQGLKKK VSNTVVKCLT DQINEQFDQI NGLEKERELY LKKI RSMES QLQASLGPGG NNPAASAPAA VAAEAASVDI LTGSTASAIE KLFNSPSASL GARVSGHNES ILNSFVSQYI PPSRE MTKD LTELWESELF NTFKLTPVVD NQGQRLYVRY SSDTISILLG PFTYLVAELS PVELVTDVYA TLGIVEIIDE LYRSSR LAI YIEDLGRKYC PASATGGDHG IRQAPSARGD TEPDHAKSKP ARDPPPGAGS UniProtKB: BBRF1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C12 (12 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.46 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 73395 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















