-Search query
-Search result
Showing 1 - 50 of 127 items for (author: doudna & j)

EMDB-48856: 
70S Ribosome of Goslar infected WT E. coli
Method: subtomogram averaging / : Klusch N, Villa E

EMDB-48875: 
70S Ribosome of Goslar infected chmA KD E. coli
Method: subtomogram averaging / : Hutchings J, Rodriguez ZK, Klusch N, Villa E

EMDB-48876: 
70S Ribosome of Goslar infected chmA KD E. coli
Method: subtomogram averaging / : Hutchings J, Rodriguez ZK, Klusch N, Villa E

EMDB-49120: 
In situ cryoET of an EPI vesicle in a Goslar infected chmA KD E. coli cell 90 mpi
Method: electron tomography / : Hutchings J, Rodriguez ZK, Klusch N, Villa E

EMDB-49121: 
In situ cryoET of an EPI vesicle in a Goslar infected chmA KD E. coli cell 90 mpi
Method: electron tomography / : Hutchings J, Rodriguez ZK, Klusch N, Villa E

EMDB-49122: 
In situ cryoET of an EPI vesicle in a Goslar infected chmA KD E. coli cell 30 mpi
Method: electron tomography / : Klusch N, Villa E

EMDB-49123: 
In situ cryoET of an EPI vesicle in a Goslar infected WT E. coli cell 1 mpi
Method: electron tomography / : Klusch N, Villa E
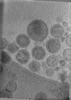
EMDB-47705: 
Tomogram of Cas9 containing Enveloped Delivery Vehicles (EDVs)
Method: electron tomography / : Peukes J, Ngo W, Nogales E, Doudna J
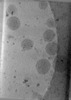
EMDB-47741: 
Tomogram of Enveloped Delivery Vehicles (EDVs) without Cas9
Method: electron tomography / : Peukes J, Ngo W, Nogales E, Doudna J

EMDB-47743: 
Subtomogram average of capsid (CA) from immature Enveloped Delivery Vehicles (EDVs) without Cas9
Method: subtomogram averaging / : Peukes J, Ngo W, Nogales E, Doudna J

EMDB-47745: 
Subtomogram average of capsid (CA) from immature Enveloped Delivery Vehicles (EDVs) containing Cas9
Method: subtomogram averaging / : Peukes J, Ngo W, Nogales E, Doudna J
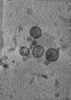
EMDB-47858: 
Tomogram of a minimized Enveloped Delivery Vehicle (miniEDV)
Method: electron tomography / : Peukes J, Ngo W, Nogales E, Doudna J

EMDB-45631: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 1
Method: single particle / : Soczek KM, Doudna JA

EMDB-45632: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 2
Method: single particle / : Soczek KM, Doudna JA

EMDB-45633: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 3
Method: single particle / : Soczek KM, Doudna JA

PDB-9cjh: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 1
Method: single particle / : Soczek KM, Doudna JA

PDB-9cji: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 2
Method: single particle / : Soczek KM, Doudna JA

PDB-9cjj: 
Cas12a:gRNA:DNA (Acidaminococcus sp.) with 0 RNA:DNA base pairs, structure 3
Method: single particle / : Soczek KM, Doudna JA

EMDB-42837: 
Cryo-EM structure of GeoCas9 in complex with sgRNA and target DNA
Method: single particle / : Eggers AR, Soczek KM, Tuck OT, Doudna JA

EMDB-42838: 
Cryo-EM structure of iGeoCas9 in complex with sgRNA and target DNA
Method: single particle / : Eggers AR, Soczek KM, Tuck OT, Doudna JA

PDB-8uza: 
Cryo-EM structure of GeoCas9 in complex with sgRNA and target DNA
Method: single particle / : Eggers AR, Soczek KM, Tuck OT, Doudna JA

PDB-8uzb: 
Cryo-EM structure of iGeoCas9 in complex with sgRNA and target DNA
Method: single particle / : Eggers AR, Soczek KM, Tuck OT, Doudna JA

EMDB-43613: 
Structure of HamAB apo complex from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

EMDB-43615: 
Structure of HamB-DNA complex, conformation 1, from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

EMDB-43616: 
Structure of HamB-DNA complex, conformation 2, from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

EMDB-43643: 
Structure of HamA(E138A,K140A)B-plasmid DNA complex from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Hu JJ, Doudna JA

PDB-8vx9: 
Structure of HamAB apo complex from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

PDB-8vxa: 
Structure of HamB-DNA complex, conformation 1, from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

PDB-8vxc: 
Structure of HamB-DNA complex, conformation 2, from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Doudna JA

PDB-8vxy: 
Structure of HamA(E138A,K140A)B-plasmid DNA complex from the Escherichia coli Hachiman defense system
Method: single particle / : Tuck OT, Hu JJ, Doudna JA

EMDB-29561: 
Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-deficient prespacer complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

EMDB-29562: 
Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-containing prespacer complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

EMDB-29563: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

EMDB-29564: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex linear CRISPR repeat conformation
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

EMDB-29565: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex bent CRISPR repeat conformation
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

PDB-8fy9: 
Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-deficient prespacer complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

PDB-8fya: 
Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-containing prespacer complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

PDB-8fyb: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

PDB-8fyc: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex linear CRISPR repeat conformation
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

PDB-8fyd: 
Cryo-EM structure of Cas1:Cas2-DEDDh:half-site integration complex bent CRISPR repeat conformation
Method: single particle / : Skopintsev P, Tuck OT, Soczek KM, Doudna J

EMDB-27320: 
Cryo-EM structure of CasLambda (Cas12l) bound to crRNA and DNA
Method: single particle / : Al-Shayeb B, Skopintsev P, Soczek K, Doudna J

PDB-8dc2: 
Cryo-EM structure of CasLambda (Cas12l) bound to crRNA and DNA
Method: single particle / : Al-Shayeb B, Skopintsev P, Soczek K, Doudna J

EMDB-24817: 
Cas9:sgRNA:DNA (S. pyogenes) with 0 RNA:DNA base pairs, closed-protein/bent-DNA conformation
Method: single particle / : Cofsky JC, Soczek KM

EMDB-24818: 
Cas9:sgRNA (S. pyogenes) in the open-protein conformation
Method: single particle / : Cofsky JC, Soczek KM

EMDB-24819: 
Cas9:sgRNA:DNA (S. pyogenes) forming a 3-base-pair R-loop
Method: single particle / : Cofsky JC, Soczek KM

EMDB-24823: 
Cas9:sgRNA:DNA (S. pyogenes) with 0 RNA:DNA base pairs, open-protein/linear-DNA conformation
Method: single particle / : Cofsky JC, Soczek KM

PDB-7s36: 
Cas9:sgRNA:DNA (S. pyogenes) with 0 RNA:DNA base pairs, closed-protein/bent-DNA conformation
Method: single particle / : Cofsky JC, Soczek KM, Knott GJ, Nogales E, Doudna JA

PDB-7s37: 
Cas9:sgRNA (S. pyogenes) in the open-protein conformation
Method: single particle / : Cofsky JC, Soczek KM, Knott GJ, Nogales E, Doudna JA

PDB-7s38: 
Cas9:sgRNA:DNA (S. pyogenes) forming a 3-base-pair R-loop
Method: single particle / : Cofsky JC, Soczek KM, Knott GJ, Nogales E, Doudna JA

PDB-7s3h: 
Cas9:sgRNA:DNA (S. pyogenes) with 0 RNA:DNA base pairs, open-protein/linear-DNA conformation
Method: single particle / : Cofsky JC, Soczek KM, Knott GJ, Nogales E, Doudna JA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
