-Search query
-Search result
Showing 1 - 50 of 79 items for (author: chang & lf)

EMDB-66358: 
Cryo-EM structure of TMEM63A-digitonin-cholesterol
Method: single particle / : Lin Y, Zhou Z, Han Y, Cheng D, Wang H, Ju L, Zhang Y, Cox DC, Corry B

PDB-9wxv: 
Cryo-EM structure of TMEM63A-digitonin-cholesterol
Method: single particle / : Lin Y, Zhou Z, Han Y, Cheng D, Wang H, Ju L, Zhang Y, Cox DC, Corry B

EMDB-48548: 
SARS-CoV-2 S2 monomer in complex with R125-61 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-48549: 
SARS-CoV-2 S2 monomer in complex with NICA01B-1113 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-48550: 
SARS-CoV-2 S2 monomer in complex with NICA01A-1401 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

PDB-9mr1: 
SARS-CoV-2 S2 monomer in complex with R125-61 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

PDB-9mr2: 
SARS-CoV-2 S2 monomer in complex with NICA01A-1401 Fab
Method: single particle / : Park S, Bangaru B, Ward AB

EMDB-18667: 
Structure of human SPNS2 in LMNG
Method: single particle / : Li HZ, Pike ACW, McKinley G, Mukhopadhyay SMM, Moreau C, Scacioc A, Abrusci P, Borkowska O, Chalk R, Stefanic S, Burgess-Brown N, Duerr KL, Sauer DB

EMDB-18668: 
Structure of human SPNS2 in DDM
Method: single particle / : Li HZ, Pike ACW, McKinley G, Mukhopadhyay SMM, Moreau C, Scacioc A, Abrusci P, Borkowska O, Chalk R, Stefanic S, Burgess-Brown N, Duerr KL, Sauer DB

PDB-8qv5: 
Structure of human SPNS2 in LMNG
Method: single particle / : Li HZ, Pike ACW, McKinley G, Mukhopadhyay SMM, Moreau C, Scacioc A, Abrusci P, Borkowska O, Chalk R, Stefanic S, Burgess-Brown N, Duerr KL, Sauer DB

PDB-8qv6: 
Structure of human SPNS2 in DDM
Method: single particle / : Li HZ, Pike ACW, McKinley G, Mukhopadhyay SMM, Moreau C, Scacioc A, Abrusci P, Borkowska O, Chalk R, Stefanic S, Burgess-Brown N, Duerr KL, Sauer DB

EMDB-45492: 
Structure of the TSC:WIPI3 lysosomal recruitment complex
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45510: 
The WIPI3:TSC lysosomal docking complex (consensus reconstruction)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45511: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; core)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45512: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; TSC1 N-terminus)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45513: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; TBC1D7)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45514: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; TBC1D7/TSC2)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45515: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; WIPI3)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-45529: 
The WIPI3:TSC lysosomal docking complex (focused reconstruction; WIPI3 TSC2)
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

PDB-9ce3: 
Structure of the TSC:WIPI3 lysosomal recruitment complex
Method: single particle / : Bayly-Jones C, Lupton CJ, D'Andrea L, Ellisdon AM

EMDB-47150: 
Mitochondrial fission site in neurons at the constriction stage
Method: electron tomography / : Peng R, Chang YW

EMDB-47151: 
Mitochondrial fission site in neurons at the post fission state 1
Method: electron tomography / : Peng R, Chang YW

EMDB-47152: 
Mitochondrial fission site in neurons at the post fission state 2
Method: electron tomography / : Peng R, Chang YW

EMDB-36241: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (consensus map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36242: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (Masked refinement of the cap domain)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36243: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (masked refinement of the transmembrane domain)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36244: 
Cryo-EM structure of mouse Piezo1-MDFIC(C240A) complex
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-35577: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (composite map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

PDB-8imz: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (composite map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-27112: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (global refinement)
Method: single particle / : Ozorowski G, Torres JL, Turner HL, Ward AB

EMDB-27113: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)
Method: single particle / : Ozorowski G, Torres JL, Turner HL, Ward AB

PDB-8d0z: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-25524: 
Reconstruction of full-length Prex-1 (PtdIns(3,4,5)P3-dependent Rac Exchanger 1)
Method: single particle / : Lupton CJ, Bayly-Jones C

EMDB-25525: 
Localised reconstruction of the N-terminal half of P-Rex1 (PI(3,4,5)P3-dependent Rac Exchanger 1)
Method: single particle / : Lupton CJ, Bayly-Jones C, Ellisdon AM

EMDB-25526: 
Localised reconstruction of the C-terminal half of P-Rex 1 (PI(3,4,5)P3-dependent Rac Exchanger 1)
Method: single particle / : Lupton CJ, Bayly-Jones C, Ellisdon AM

PDB-7syf: 
Reconstruction of full-length Prex-1 (PtdIns(3,4,5)P3-dependent Rac Exchanger 1)
Method: single particle / : Lupton CJ, Bayly-Jones C, Ellisdon AM

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
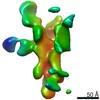
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
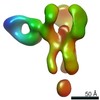
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
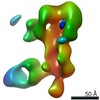
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
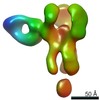
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25646: 
Negative stain map of polyclonal Fab 241.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25655: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
