-Search query
-Search result
Showing 1 - 50 of 62 items for (author: cambillau & c)

EMDB-70849: 
C33 reconstruction for the PorKN (single) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70850: 
C34 reconstruction for the PorKN (single) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70851: 
C32 reconstruction for the PorKN (single) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70853: 
C33 reconstruction for the PorKN (double) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70854: 
C34 reconstruction for the PorKN (double) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70856: 
C32 reconstruction for the PorKN (double) ring complex of Type IX Secretion System (T9SS)
Method: single particle / : Liu X, Song L, Hu B

EMDB-70857: 
Cryo-EM structure of the T9SS PORkN ring complex of P. Gingivalis
Method: single particle / : Liu X, Song L, Zheng L, Hu B

PDB-9ots: 
Cryo-EM structure of the T9SS PORkN ring complex of P. Gingivalis
Method: single particle / : Liu X, Song L, Zheng L, Hu B

EMDB-11224: 
negative staining 3D reconstruction of p2 virion baseplate in activated conformation (3D class with open Tal trimer)
Method: single particle / : Spinelli S, Cambillau C, Goulet A

EMDB-11225: 
negative staining 3D reconstruction of p2 virion baseplate in activated conformation (3D class with closed Tal trimer)
Method: single particle / : Spinelli S, Cambillau C

EMDB-11226: 
Negative staining 3D reconstruction of p2 virion baseplate in activated conformation
Method: single particle / : Spinelli S, Cambillau C

PDB-6zig: 
Topological model of the p2 virion baseplate in activated conformation (closed Tal trimer)
Method: single particle / : Spinelli S, Cambillau C, Goulet A

PDB-6zih: 
Topological model of p2 virion baseplate in activated conformation
Method: single particle / : Spinelli S, Cambillau C, Goulet A

PDB-6zjj: 
Topological model of p2 virion baseplate in resting conformation
Method: single particle / : Spinelli S, Cambillau C, Goulet A

EMDB-10913: 
Structure of the phage STP1 host adhesion device (baseplate) by negative staining
Method: single particle / : Lavelle K, Goulet A, McDonnell B, Spinelli S, van Sinderen D, Mahony J, Cambillau C

EMDB-4900: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA20-AcrIIA6 monomeric assembly.
Method: single particle / : Swuec P

EMDB-4901: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA20-AcrIIA6 dimeric assembly.
Method: single particle / : Swuec P

EMDB-4902: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex.
Method: single particle / : Swuec P

EMDB-4904: 
Cryo-EM structure of St1Cas9-sgRNA-AcrIIA6-tDNA59-ntPAM complex.
Method: single particle / : Swuec P

PDB-6rj9: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA20-AcrIIA6 monomeric assembly.
Method: single particle / : Goulet A, Chaves-Sanjuan A, Cambillau C

PDB-6rja: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA20-AcrIIA6 dimeric assembly.
Method: single particle / : Goulet A, Cambillau C, Chaves-Sanjuan A

PDB-6rjd: 
Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex.
Method: single particle / : Goulet A, Chaves-Sanjuan A, Cambillau C

PDB-6rjg: 
Cryo-EM structure of St1Cas9-sgRNA-AcrIIA6-tDNA59-ntPAM complex.
Method: single particle / : Goulet A, Chaves-Sanjuan A, Cambillau C

EMDB-9341: 
Structure of the type VI secretion system TssK-TssF-TssG baseplate subcomplex revealed by cryo-electron microscopy - full map sharpened
Method: single particle / : Park YJ, Lacourse KD, Cambillau C, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

EMDB-9342: 
Structure of the type VI secretion system TssK-TssF-TssG baseplate subcomplex revealed by cryo-electron microscopy - TssK focused map
Method: single particle / : Park YJ, Lacourse KD, Cambillau C, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

EMDB-9343: 
Structure of the type VI secretion system TssK-TssF-TssG baseplate subcomplex revealed by cryo-electron microscopy - TssK focused map
Method: single particle / : Park YJ, Lacourse KD, Cambillau C, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

PDB-6n38: 
Structure of the type VI secretion system TssK-TssF-TssG baseplate subcomplex revealed by cryo-electron microscopy - full map sharpened
Method: single particle / : Park YJ, Lacourse KD, Cambillau C, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

EMDB-3641: 
Unraveling the self-assembly of Pseudomonas aeruginosa XcpQ secretin periplasmic domain provides new molecular insights into T2SS secreton architecture & dynamics
Method: single particle / : Douzi B, Trinh N, Ball G, Desmyter A, Michel-Souzy S, Barbier P, Kosta A, Durand E, Cambillau C, Roussel A, Voulhoux R

EMDB-3649: 
Unraveling the self-assembly of Pseudomonas aeruginosa XcpQ secretin periplasmic domain provides new molecular insights into T2SS secreton architecture & dynamics
Method: single particle / : Douzi B, Trinh N, Ball G, Desmyter A, Michel-Souzy S, Barbier P, Kosta A, Durand E, Cambillau C, Roussel A, Vouloux R

EMDB-4150: 
Structure of the baseplate of Siphophage J-1
Method: single particle / : Cambillau C, Spinelli S, Dieterle ME, Piuri M

EMDB-3282: 
Negative stain electron microscopy structure of TssA from EAEC type VI secretion system
Method: single particle / : Durand E, Fronzes R, Cambillau C, Cascales E

EMDB-2816: 
Electron cryoEM structure of lactococcal siphophage 1358 virion
Method: single particle / : Spinelli S, Bebeacua C, Orlov I, Tremblay D, Klaholz B, Moineau S, Cambillau C

EMDB-2817: 
Electron cryoEM structure of lactococcal siphophage 1358 virion
Method: single particle / : Spinelli S, Bebeacua C, Orlov I, Tremblay D, Klaholz B, Moineau S, Cambillau C

EMDB-2819: 
Electron cryoEM structure of lactococcal siphophage 1358 virion
Method: single particle / : Spinelli S, Bebeacua C, Orlov I, Tremblay D, Blangy S, Klaholz B, Moineau S, Cambillau C

EMDB-2820: 
Electron cryoEM structure of lactococcal siphophage 1358 virion
Method: single particle / : Spinelli S, Bebeacua C, Orlov I, Tremblay D, Klaholz B, Moineau S, Cambillau C

EMDB-2928: 
Negative stain structure of a type 6 secretion system membrane core complex
Method: single particle / : Durand E, Fronzes R

EMDB-2927: 
Negative stain structure of a type 6 secretion system membrane core complex
Method: single particle / : Durand E, Fronzes R

EMDB-2698: 
The cryoEM structure of Monalysin Toxin
Method: single particle / : Leone P, Bebeacua C, Opota O, Kellenberger C, Klaholz B, Cambillau C, Lemaitre B, Roussel A

EMDB-2647: 
electron cryo-microscopy of 1358 Lactococcus phage mature empty capsid
Method: single particle / : Spinelli S, Bebeacua C, Orlov I, Tremblay D, Klaholz B, Moineau S, Cambillau C

EMDB-2459: 
Structure and Host Adhesion Mechanism of Virulent Lactococcal Phage p2
Method: single particle / : Bebeacua C, Tremblay D, Farenc C, Chapot MP, Sadovskaya I, van Heel M, Veesler D, Moineau S, Cambillau C

EMDB-2462: 
Structure and Host Adhesion Mechanism of Virulent Lactococcal Phage p2
Method: single particle / : Bebeacua C, Tremblay D, Farenc C, Chapot MP, Sadovskaya I, van Heel M, Veesler D, Moineau S, Cambillau C

EMDB-2463: 
The electron cryo-miscroscopy reconstruction of the connector of lactococcal phage p2
Method: single particle / : Bebeacua C, Tremblay D, Farenc C, Chapot MP, Sadovskaya I, van Heel M, Veesler D, Moineau S, Cambillau C
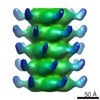
EMDB-2464: 
The electron miscroscopy reconstruction of the tail of lactococcal phage p2
Method: single particle / : Bebeacua C, Tremblay D, Farenc C, Chapot MP, Sadovskaya I, van Heel M, Veesler D, Moineau S, Cambillau C

EMDB-5739: 
Electron microscopy map of the T6SS TssK component
Method: single particle / : Zoued A, Durand E, Bebeacua C, Brunet YR, Douzi B, Cambillau C, Cascales E, Journet L

EMDB-2335: 
The capsid of mycobacteriophage Araucaria
Method: single particle / : Sassi M, Bebeacua C, Drancourt M, Cambillau C

EMDB-2336: 
The connector of mycobacteriophage Araucaria
Method: single particle / : Sassi M, Bebeacua C, Drancourt M, Cambillau C

EMDB-2337: 
The tail of mycobacteriophage Araucaria
Method: single particle / : Sassi M, Bebeacua C, Drancourt M, Cambillau C

EMDB-2338: 
The baseplate of mycobacteriophage Araucaria
Method: single particle / : Sassi M, Bebeacua C, Drancourt M, Cambillau C

EMDB-2340: 
The electron microscopy reconstruction of the baseplate of lactococcal phage Tuc2009.
Method: single particle / : Collins B, Bebeacua C, Mahony J, Douillard F, Veesler D, Blangy B, Cambillau C, van Sinderen D

EMDB-2343: 
The electron microscopy reconstruction of the BppU-CtAL tripod of lactococcal phage Tuc2009.
Method: single particle / : Collins B, Bebeacua C, Mahony J, Douillard F, Veesler D, Blangy B, Cambillau C, van Sinderen D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
